-Search query
-Search result
Showing 1 - 50 of 1,134 items for (author: thomas & ja)

EMDB-16903: 
60S ribosomal subunit bound to the E3-UFM1 complex (native, UFM1 pulldown)
Method: single particle / : Penchev I, DaRosa PA, Becker T, Beckmann R, Kopito R

EMDB-16880: 
60S ribosomal subunit bound to the E3-UFM1 complex - state 3 (native)
Method: single particle / : Penchev I, DaRosa PA, Becker T, Beckmann R, Kopito R

EMDB-16902: 
60S ribosomal subunit bound to the E3-UFM1 complex - state 2 (native)
Method: single particle / : Penchev I, DaRosa PA, Becker T, Beckmann R, Kopito R
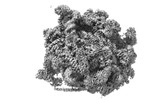
EMDB-16905: 
60S ribosomal subunit bound to the E3-UFM1 complex - state 3 (in-vitro reconstitution)
Method: single particle / : Penchev I, DaRosa PA, Peter JJ, Kulathu Y, Becker T, Beckmann R, Kopito R

EMDB-16908: 
60S ribosomal subunit bound to the E3-UFM1 complex - state 1 (native)
Method: single particle / : Penchev I, DaRosa PA, Becker T, Beckmann R, Kopito R

EMDB-15525: 
Cryo-EM structure of the RecA postsynaptic filament from S. pneumoniae
Method: helical / : Perry TN, Fronzes R, Polard P, Hertzog M

PDB-8amf: 
Cryo-EM structure of the RecA postsynaptic filament from S. pneumoniae
Method: helical / : Perry TN, Fronzes R, Polard P, Hertzog M
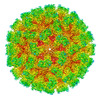

EMDB-43222: 
Cryo-EM structure of Tulane virus 9-6-17 variant capsid protein VP1 5-12-18
Method: single particle / : Sun C, Jiang W

EMDB-43292: 
Cryo-EM structure of Tulane virus 9-6-17 variant capsid protein VP1 9-14-18, DTT-treated
Method: single particle / : Sun C, Jiang W

EMDB-43293: 
Cryo-EM structure of Tulane virus 9-6-17 variant capsid protein VP1 9-14-18 without DTT treatment
Method: single particle / : Sun C, Jiang W

PDB-8vgr: 
Cryo-EM structure of Tulane virus 9-6-17 variant capsid protein VP1 5-12-18
Method: single particle / : Sun C, Jiang W

PDB-8vjr: 
Cryo-EM structure of Tulane virus 9-6-17 variant capsid protein VP1 9-14-18, DTT-treated
Method: single particle / : Sun C, Jiang W

PDB-8vjs: 
Cryo-EM structure of Tulane virus 9-6-17 variant capsid protein VP1 9-14-18 without DTT treatment
Method: single particle / : Sun C, Jiang W

EMDB-43542: 
ELIC5 with cysteamine in 2:1:1 POPC:POPE:POPG nanodisc in open conformation
Method: single particle / : Petroff II JT, Deng Z, Rau MJ, Fitzpatrick JAJ, Yuan P, Cheng WWL

PDB-8vuw: 
ELIC5 with cysteamine in 2:1:1 POPC:POPE:POPG nanodisc in open conformation
Method: single particle / : Petroff II JT, Deng Z, Rau MJ, Fitzpatrick JAJ, Yuan P, Cheng WWL

EMDB-41374: 
Antibody N3-1 bound to RBDs in the up and down conformations
Method: single particle / : Hsieh CL, McLellan JS
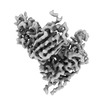
EMDB-41382: 
Antibody N3-1 bound to RBD in the up conformation
Method: single particle / : Hsieh CL, McLellan JS
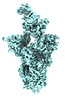
EMDB-41399: 
Antibody N3-1 bound to SARS-CoV-2 spike
Method: single particle / : Hsieh CL, McLellan JS

PDB-8tm1: 
Antibody N3-1 bound to RBDs in the up and down conformations
Method: single particle / : Hsieh CL, McLellan JS

PDB-8tma: 
Antibody N3-1 bound to RBD in the up conformation
Method: single particle / : Hsieh CL, McLellan JS
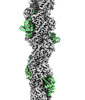
EMDB-16424: 
F-actin decorated by SipA497-669
Method: helical / : Yuan B, Wald J, Marlovits TC
EMDB-16425: 
F-actin decorated by SipA426-685
Method: helical / : Yuan B, Wald J, Marlovits TC

PDB-8c4c: 
F-actin decorated by SipA497-669
Method: helical / : Yuan B, Wald J, Marlovits TC

PDB-8c4e: 
F-actin decorated by SipA426-685
Method: helical / : Yuan B, Wald J, Marlovits TC

EMDB-40856: 
Single particle reconstruction of the human LINE-1 ORF2p without substrate (apo)
Method: single particle / : van Eeuwen T, Taylor MS, Rout MP
EMDB-40858: 
Structure of LINE-1 ORF2p with template:primer hybrid
Method: single particle / : van Eeuwen T, Taylor MS, Rout MP
EMDB-40859: 
Structure of LINE-1 ORF2p with an oligo(A) template
Method: single particle / : van Eeuwen T, Taylor MS, Rout MP

PDB-8sxt: 
Structure of LINE-1 ORF2p with template:primer hybrid
Method: single particle / : van Eeuwen T, Taylor MS, Rout MP

PDB-8sxu: 
Structure of LINE-1 ORF2p with an oligo(A) template
Method: single particle / : van Eeuwen T, Taylor MS, Rout MP

EMDB-16805: 
Cryo-EM structure of PcrV/Fab(30-B8)
Method: single particle / : Yuan B, Simonis A, Marlovits TC

EMDB-16807: 
Cryo-EM structure of PcrV/Fab(11-E5)
Method: single particle / : Yuan B, Simonis A, Marlovits TC

PDB-8cr9: 
Cryo-EM structure of PcrV/Fab(30-B8)
Method: single particle / : Yuan B, Simonis A, Marlovits TC

PDB-8crb: 
Cryo-EM structure of PcrV/Fab(11-E5)
Method: single particle / : Yuan B, Simonis A, Marlovits TC

EMDB-42124: 
Cryo-EM structure of human STEAP1 in complex with AMG 509 Fab
Method: single particle / : Li F, Bailis JM

PDB-8ucd: 
Cryo-EM structure of human STEAP1 in complex with AMG 509 Fab
Method: single particle / : Li F, Bailis JM, Zhang H

EMDB-41617: 
CryoEM structure of PI3Kalpha
Method: single particle / : Valverde R, Shi H, Holliday M, Sun M

PDB-8tu6: 
CryoEM structure of PI3Kalpha
Method: single particle / : Valverde R, Shi H, Holliday M

EMDB-41667: 
Cryo-EM structure of the PP2A:B55-FAM122A complex, B55 body
Method: single particle / : Fuller JR, Padi SKR, Peti W, Page R

EMDB-41668: 
Cryo-EM structure of the PP2A:B55-FAM122A complex, PP2Ac body
Method: single particle / : Fuller JR, Padi SKR, Peti W, Page R

PDB-8twe: 
Cryo-EM structure of the PP2A:B55-FAM122A complex, B55 body
Method: single particle / : Fuller JR, Padi SKR, Peti W, Page R

PDB-8twi: 
Cryo-EM structure of the PP2A:B55-FAM122A complex, PP2Ac body
Method: single particle / : Fuller JR, Padi SKR, Peti W, Page R

EMDB-29281: 
Cryo-EM structure of STING oligomer bound to cGAMP and NVS-STG2
Method: single particle / : Li J, Canham SM, Zhang X, Bai X, Feng Y

EMDB-29282: 
Cryo-EM structure of STING oligomer bound to cGAMP, NVS-STG2 and C53
Method: single particle / : Li J, Canham SM, Zhang X, Bai X, Feng Y

PDB-8flk: 
Cryo-EM structure of STING oligomer bound to cGAMP and NVS-STG2
Method: single particle / : Li J, Canham SM, Zhang X, Bai X, Feng Y

PDB-8flm: 
Cryo-EM structure of STING oligomer bound to cGAMP, NVS-STG2 and C53
Method: single particle / : Li J, Canham SM, Zhang X, Bai X, Feng Y

EMDB-40644: 
Cryo-EM structure of the PP2A:B55-FAM122A complex
Method: single particle / : Fuller JR, Padi SKR, Peti W, Page R

EMDB-41604: 
Cryo-EM structure of the PP2A:B55-ARPP19 complex
Method: single particle / : Fuller JR, Padi SKR, Peti W, Page R

PDB-8so0: 
Cryo-EM structure of the PP2A:B55-FAM122A complex
Method: single particle / : Fuller JR, Padi SKR, Peti W, Page R

PDB-8ttb: 
Cryo-EM structure of the PP2A:B55-ARPP19 complex
Method: single particle / : Fuller JR, Padi SKR, Peti W, Page R

EMDB-40789: 
BAP1/ASXL1 bound to the H2AK119Ub Nucleosome
Method: single particle / : Thomas JF, Valencia-Sanchez MI, Armache KJ
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
