-Search query
-Search result
Showing 1 - 50 of 2,409 items for (author: tao & h)
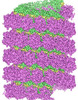

EMDB-31367: 
Structure of mumps virus nucleoprotein without C-arm
Method: helical / : Shen Q, Shan H, Zhang N, Qin Y
EMDB-37197: 
The cryo-EM structure of type3 amyloid beta 42 fibril.
Method: helical / : Zhao QY, Tao YQ, Liu C, Li D

EMDB-37200: 
The cryo-EM structure of AV-45 bound type3 amyloid beta 42 fibril.
Method: helical / : Zhao QY, Tao YQ, Liu C, Li D

EMDB-37199: 
The cryo-EM structure of type1 amyloid beta 42 fibril in AD3.
Method: helical / : Zhao QY, Tao YQ, Liu C, Li D

EMDB-37195: 
The cryo-EM structure of AV-45 bound type1 amyloid beta 42 fibril.
Method: helical / : Zhao QY, Tao YQ, Liu C, Li D

EMDB-37198: 
The cryo-EM structure of type1 amyloid beta 42 fibril in AD2 patient.
Method: helical / : Zhao QY, Tao YQ, Liu C, Li D

EMDB-38300: 
Cryo-EM structure of partial dimeric WDR11-FAM91A1 complex
Method: single particle / : Jia GW, Deng QH, Su ZM, Jia D

EMDB-39863: 
Cryo-EM structure of dimeric WDR11-FAM91A1 complex
Method: single particle / : Jia GW, Deng QH, Su ZM, Jia D

EMDB-39943: 
Cryo-EM structure of dimeric WDR11-FAM91A1 complex Body2
Method: single particle / : Jia GW, Deng QH, Su ZM, Jia D

EMDB-39947: 
Cryo-EM structure of dimeric WDR11-FAM91A1 complex Body1
Method: single particle / : Jia GW, Deng QH, Su ZM, Jia D

EMDB-39949: 
Cryo-EM structure of WDR11-dm-FAM91A1 complex
Method: single particle / : Jia GW, Deng QH, Su ZM, Jia D

PDB-8xfb: 
Cryo-EM structure of partial dimeric WDR11-FAM91A1 complex
Method: single particle / : Jia GW, Deng QH, Su ZM, Jia D

PDB-8z9m: 
Cryo-EM structure of dimeric WDR11-FAM91A1 complex
Method: single particle / : Jia GW, Deng QH, Su ZM, Jia D

EMDB-37170: 
The cryo-EM structure of type1 amyloid beta 42 fibril.
Method: helical / : Zhao QY, Tao YQ, Liu C, Li D
PDB-8kew: 
The cryo-EM structure of type1 amyloid beta 42 fibril.
Method: helical / : Zhao QY, Tao YQ, Liu C, Li D
EMDB-38099: 
Cryo-EM structures of RNF168/UbcH5c-Ub in complex with H2AK13Ub nucleosomes determined by intein-based E2-Ub-NCP conjugation strategy
Method: single particle / : Ai HS, Tong ZB, Deng ZH, Pan M, Liu L
EMDB-38100: 
Cryo-EM structures of RNF168/UbcH5c-Ub/nucleosomes complex determined by activity-based chemical trapping strategy
Method: single particle / : Ai HS, Tong ZB, Deng ZH, Pan M, Liu L
EMDB-38101: 
Cryo-EM structures of RNF168/UbcH5c-Ub in complex with H2AK13Ub nucleosomes determined by activity-based chemical trapping strategy (adjacent H2AK13/15 dual-monoubiquitination)
Method: single particle / : Ai HS, Tong ZB, Deng ZH, Pan M, Liu L
EMDB-38102: 
Cryo-EM map of RNF168/UbcH5c-Ub/nucleosome determined by E2-Ub-NCP conjugation strategy
Method: single particle / : Ai H, Zebin T, Deng Z, Pan M, Liu L

EMDB-37499: 
Cryo-EM structure of CRISPR-Csm effector complex from Mycobacterium canettii
Method: single particle / : Huo Y, Ma X, Jiang T

PDB-8wfx: 
Cryo-EM structure of CRISPR-Csm effector complex from Mycobacterium canettii
Method: single particle / : Huo Y, Ma X, Jiang T

EMDB-60607: 
A local Cryo-EM structure of Bitter taste receptor TAS2R14
Method: single particle / : Yuan Q, Duan J, Tao L, Xu EH

EMDB-60608: 
A Cryo-EM structure of Bitter taste receptor TAS2R14 with Ggust
Method: single particle / : Yuan Q, Duan J, Tao L, Xu EH

EMDB-60626: 
A Cryo-EM structure of Bitter taste receptor TAS2R14 with Gi complex
Method: single particle / : Yuan Q, Duan J, Tao L, Xu EH
EMDB-60627: 
A local Cryo-EM structure of Bitter taste receptor TAS2R14 with Gi complex
Method: single particle / : Yuan Q, Duan J, Tao L, Xu EH

PDB-9iiw: 
A local Cryo-EM structure of Bitter taste receptor TAS2R14
Method: single particle / : Yuan Q, Duan J, Tao L, Xu EH

PDB-9iix: 
A Cryo-EM structure of Bitter taste receptor TAS2R14 with Ggust
Method: single particle / : Yuan Q, Duan J, Tao L, Xu EH

PDB-9ij9: 
A Cryo-EM structure of Bitter taste receptor TAS2R14 with Gi complex
Method: single particle / : Yuan Q, Duan J, Tao L, Xu EH

PDB-9ija: 
A local Cryo-EM structure of Bitter taste receptor TAS2R14 with Gi complex
Method: single particle / : Yuan Q, Duan J, Tao L, Xu EH

EMDB-39025: 
Structure of HCoV-HKU1A spike in the functionally anchored-3up conformation with 3TMPRSS2
Method: single particle / : Lu YC, Zhang X, Wang HF, Liu XC, Sun L, Yang HT

EMDB-39026: 
Local structure of HCoV-HKU1A spike in complex with TMPRSS2 and glycan
Method: single particle / : Wang HF, Zhang X, Lu Y, Liu X, Sun L, Yang HT

EMDB-39036: 
Structure of HCoV-HKU1C spike in the functionally anchored-1up conformation with 1TMPRSS2
Method: single particle / : Lu YC, Zhang X, Wang HF, Liu XC, Sun L, Yang HT

EMDB-39037: 
Structure of HCoV-HKU1C spike in the functionally anchored-2up conformation with 2TMPRSS2
Method: single particle / : Lu YC, Zhang X, Wang HF, Liu XC, Sun L, Yang HT

EMDB-39038: 
Structure of HCoV-HKU1C spike in the functionally anchored-3up conformation with 2TMPRSS2
Method: single particle / : Lu YC, Wang HF, Zhang X, Liu XC, Sun L, Yang HT

EMDB-39039: 
Structure of HCoV-HKU1C spike in the functionally anchored-3up conformation with 3TMPRSS2
Method: single particle / : Lu YC, Zhang X, Wang HF, Liu XC, Sun L, Yang HT

EMDB-39040: 
Local structure of HCoV-HKU1C spike in complex with TMPRSS2 and glycan
Method: single particle / : Wang HF, Zhang X, Lu YC, Liu XC, Sun L, Yang HT

EMDB-39041: 
Structure of HCoV-HKU1C spike in the inactive-closed conformation
Method: single particle / : Lu YC, Zhang X, Wang HF, Sun L, Yang HT

EMDB-39042: 
Structure of HCoV-HKU1C spike in the inactive-1up conformation
Method: single particle / : Lu YC, Zhang X, Wang HF, Sun L, Yang HT

EMDB-39043: 
Structure of HCoV-HKU1C spike in the inactive-2up conformation
Method: single particle / : Lu YC, Zhang X, Wang HF, Sun L, Yang HT

EMDB-39044: 
Structure of HCoV-HKU1C spike in the glycan-activated-closed conformation
Method: single particle / : Lu YC, Zhang X, Wang HF, Sun L, Yang HT

EMDB-39045: 
Structure of HCoV-HKU1C spike in the glycan-activated-1up conformation
Method: single particle / : Lu YC, Zhang X, Wang HF, Sun L, Yang HT

EMDB-39046: 
Structure of HCoV-HKU1C spike in the glycan-activated-2up conformation
Method: single particle / : Lu YC, Zhang X, Wang HF, Sun L, Yang HT

EMDB-39047: 
Structure of HCoV-HKU1C spike in the glycan-activated-3up conformation
Method: single particle / : Lu YC, Zhang X, Wang HF, Sun L, Yang HT

EMDB-39048: 
Local structure of HCoV-HKU1C spike in complex with glycan
Method: single particle / : Lu YC, Zhang X, Wang HF, Sun L, Yang HT

PDB-8y7x: 
Structure of HCoV-HKU1A spike in the functionally anchored-3up conformation with 3TMPRSS2
Method: single particle / : Lu YC, Zhang X, Wang HF, Liu XC, Sun L, Yang HT

PDB-8y7y: 
Local structure of HCoV-HKU1A spike in complex with TMPRSS2 and glycan
Method: single particle / : Wang HF, Zhang X, Lu Y, Liu X, Sun L, Yang HT

PDB-8y87: 
Structure of HCoV-HKU1C spike in the functionally anchored-1up conformation with 1TMPRSS2
Method: single particle / : Lu YC, Zhang X, Wang HF, Liu XC, Sun L, Yang HT

PDB-8y88: 
Structure of HCoV-HKU1C spike in the functionally anchored-2up conformation with 2TMPRSS2
Method: single particle / : Lu YC, Zhang X, Wang HF, Liu XC, Sun L, Yang HT

PDB-8y89: 
Structure of HCoV-HKU1C spike in the functionally anchored-3up conformation with 2TMPRSS2
Method: single particle / : Lu YC, Wang HF, Zhang X, Liu XC, Sun L, Yang HT

PDB-8y8a: 
Structure of HCoV-HKU1C spike in the functionally anchored-3up conformation with 3TMPRSS2
Method: single particle / : Lu YC, Zhang X, Wang HF, Liu XC, Sun L, Yang HT
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
