-Search query
-Search result
Showing 1 - 50 of 184 items for (author: takayuki & kato)

EMDB-39212: 
Cryo-EM structure of Dragon Grouper nervous necrosis virus-like particle at pH8.0 (3.23A)
Method: single particle / : Wang CH, Chang WH

EMDB-39213: 
Cryo-EM structure of Dragon Grouper nervous necrosis virus-like particle at pH6.5 (2.82A)
Method: single particle / : Wang CH, Chang WH

EMDB-39214: 
Cryo-EM structure of Dragon Grouper nervous necrosis virus-like particle at pH5.0 (3.52A)
Method: single particle / : Wang CH, Chang WH

EMDB-39215: 
Cryo-EM structure of Dragon Grouper nervous necrosis virion at pH6.5 (3.12A)
Method: single particle / : Wang CH, Chang WH

EMDB-39217: 
Cryo-EM structure of Dragon Grouper nervous necrosis virion at pH5.0 (4.36A)
Method: single particle / : Wang CH, Chang WH

PDB-8yf6: 
Cryo-EM structure of Dragon Grouper nervous necrosis virus-like particle at pH8.0 (3.23A)
Method: single particle / : Wang CH, Chang WH

PDB-8yf7: 
Cryo-EM structure of Dragon Grouper nervous necrosis virus-like particle at pH6.5 (2.82A)
Method: single particle / : Wang CH, Chang WH

PDB-8yf8: 
Cryo-EM structure of Dragon Grouper nervous necrosis virus-like particle at pH5.0 (3.52A)
Method: single particle / : Wang CH, Chang WH

PDB-8yf9: 
Cryo-EM structure of Dragon Grouper nervous necrosis virion at pH6.5 (3.12A)
Method: single particle / : Wang CH, Chang WH

EMDB-35029: 
SARS-CoV2 spike protein with ACE2, no ACE2 binding.
Method: single particle / : Kishikawa J, Hirose M, Kato T, Okamoto T

EMDB-35030: 
SARS-CoV2 spike protein with ACE2. 1 ACE2 bound form.
Method: single particle / : Kishikawa J, Hirose M, Kato T, Okamoto T

EMDB-35031: 
SARS-CoV2 spike protein with ACE2. 2 ACE2 bound form.
Method: single particle / : Kishikawa J, Hirose M, Kato T, Okamoto T

EMDB-35032: 
SARS-CoV2 spike protein with ACE2. 3 ACE2 bound form.
Method: single particle / : Kishikawa J, Hirose M, Kato T, Okamoto T

EMDB-35036: 
SARS-CoV2 spike protein with ACE2 decoy.no ACE2 decoy binding
Method: single particle / : Kishikawa J, Hirose M, Kato T, Okamoto T

EMDB-35037: 
SARS-CoV2 spike protein with ACE2 decoy. 1 ACE2 decoy bound form.
Method: single particle / : Kishikawa J, Hirose M, Kato T, Okamoto T

EMDB-35038: 
SARS-CoV2 spike protein with ACE2 decoy. 1 ACE2 decoy bound and 2 RBD up form.
Method: single particle / : Kishikawa J, Hirose M, Kato T, Okamoto T

EMDB-35039: 
SARS-CoV2 spike protein with ACE2 decoy. 2 ACE2 decoy bound form.
Method: single particle / : Kishikawa J, Hirose M, Kato T, Okamoto T

EMDB-35040: 
SARS-CoV2 spike protein with ACE2 decoy. 3 ACE2 decoy bound form.
Method: single particle / : Kishikawa J, Hirose M, Kato T, Okamoto T

EMDB-36345: 
RBD of SARS-CoV2 spike protein with ACE2 decoy
Method: single particle / : Kishikawa J, Hirose M, Kato T, Okamoto T

PDB-8jje: 
RBD of SARS-CoV2 spike protein with ACE2 decoy
Method: single particle / : Kishikawa J, Hirose M, Kato T, Okamoto T

EMDB-33785: 
Cryo-EM structure of the histamine-bound histamine H4 receptor and Gq complex
Method: single particle / : Im D, Iwata S, Asada H

EMDB-33786: 
Cryo-EM structure of the imetit-bound histamine H4 receptor and Gq complex
Method: single particle / : Im D, Iwata S, Asada H

PDB-7yfc: 
Cryo-EM structure of the histamine-bound histamine H4 receptor and Gq complex
Method: single particle / : Im D, Iwata S, Asada H

PDB-7yfd: 
Cryo-EM structure of the imetit-bound histamine H4 receptor and Gq complex
Method: single particle / : Im D, Iwata S, Asada H

EMDB-34468: 
Human ATAD2 Walker B mutant, ATP state
Method: single particle / : Cho C, Song J

EMDB-36665: 
Human ATAD2 Walker B mutant-H3/H4K5Q complex, ATP state
Method: single particle / : Cho C, Song J

EMDB-36666: 
Human ATAD2 Walker B mutant-H3/H4K5Q complex, ATP state (Class II)
Method: single particle / : Cho C, Song J

EMDB-36667: 
Human ATAD2 Walker B mutant-H3/H4K5Q complex, ATP state (Class III)
Method: single particle / : Cho C, Song J

PDB-8juw: 
Human ATAD2 Walker B mutant-H3/H4K5Q complex, ATP state
Method: single particle / : Cho C, Song J

PDB-8juy: 
Human ATAD2 Walker B mutant-H3/H4K5Q complex, ATP state (Class II)
Method: single particle / : Cho C, Song J

PDB-8juz: 
Human ATAD2 Walker B mutant-H3/H4K5Q complex, ATP state (Class III)
Method: single particle / : Cho C, Song J

EMDB-34871: 
Cryo-EM structure of biparatopic antibody Bp109-92 in complex with TNFR2
Method: single particle / : Akiba H, Fujita J, Ise T, Nishiyama K, Miyata T, Kato T, Namba K, Ohno H, Kamada H, Nagata S, Tsumoto K

PDB-8hlb: 
Cryo-EM structure of biparatopic antibody Bp109-92 in complex with TNFR2
Method: single particle / : Akiba H, Fujita J, Ise T, Nishiyama K, Miyata T, Kato T, Namba K, Ohno H, Kamada H, Nagata S, Tsumoto K

EMDB-34106: 
GroEL on EG-grid stored for 3 months after graphene oxidation
Method: single particle / : Fujita J, Makino F, Asahara H, Moriguchi M, Kumano S, Anzai I, Kishikawa J, Matsuura Y, Kato T, Namba K, Inoue T
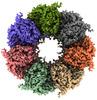
EMDB-34743: 
GroEL on Quantifoil grid
Method: single particle / : Fujita J, Makino F, Asahara H, Moriguchi M, Kumano S, Anzai I, Kishikawa J, Matsuura Y, Kato T, Namba K, Inoue T

EMDB-34805: 
cryoEM structure of glutamate dehydrogenase from Thermococcus profundus in complex with NADP
Method: single particle / : Wakabayashi T, Oide M, Kato T, Nakasako M

EMDB-34826: 
cryoEM structure of glutamate dehydrogenase from Thermococcus profundus in complex with NADP
Method: single particle / : Wakabayashi T, Oide M, Kato T, Nakasako M

EMDB-34830: 
cryoEM structure of glutamate dehydrogenase from Thermococcus profundus in complex with NADP
Method: single particle / : Wakabayashi T, Oide M, Kato T, Nakasako M

EMDB-34831: 
cryoEM structure of glutamate dehydrogenase from Thermococcus profundus in complex with NADP
Method: single particle / : Wakabayashi T, Oide M, Kato T, Nakasako M

EMDB-34835: 
cryoEM structure of glutamate dehydrogenase from Thermococcus profundus in complex with NADP
Method: single particle / : Wakabayashi T, Oide M, Kato T, Nakasako M

PDB-8hho: 
cryoEM structure of glutamate dehydrogenase from Thermococcus profundus in complex with NADP
Method: single particle / : Wakabayashi T, Oide M, Kato T, Nakasako M

PDB-8hiq: 
cryoEM structure of glutamate dehydrogenase from Thermococcus profundus in complex with NADP
Method: single particle / : Wakabayashi T, Oide M, Kato T, Nakasako M

PDB-8hiz: 
cryoEM structure of glutamate dehydrogenase from Thermococcus profundus in complex with NADP
Method: single particle / : Wakabayashi T, Oide M, Kato T, Nakasako M

PDB-8hj3: 
cryoEM structure of glutamate dehydrogenase from Thermococcus profundus in complex with NADP
Method: single particle / : Wakabayashi T, Oide M, Kato T, Nakasako M

PDB-8hj9: 
cryoEM structure of glutamate dehydrogenase from Thermococcus profundus in complex with NADP
Method: single particle / : Wakabayashi T, Oide M, Kato T, Nakasako M

EMDB-32159: 
GroEL on hydrophilized graphene grid (high particle density)
Method: single particle / : Fujita J, Makino F, Asahara H, Moriguchi M, Kumano S, Anzai I, Kishikawa J, Matsuura Y, Kato T, Namba K, Inoue T
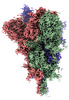
EMDB-32160: 
SARS-CoV-2 spike protein (1-up RBD) on EG-grid
Method: single particle / : Fujita J, Makino F, Asahara H, Moriguchi M, Kumano S, Anzai I, Kishikawa J, Matsuura Y, Kato T, Namba K, Inoue T
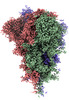
EMDB-32161: 
SARS-CoV-2 spike protein (1-up RBD) on Quantifoil grid
Method: single particle / : Fujita J, Makino F, Asahara H, Moriguchi M, Kumano S, Anzai I, Kishikawa J, Matsuura Y, Kato T, Namba K, Inoue T
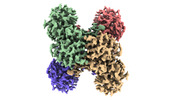
EMDB-32162: 
GAPDH on EG-grid
Method: single particle / : Fujita J, Makino F, Asahara H, Moriguchi M, Kumano S, Anzai I, Kishikawa J, Matsuura Y, Kato T, Namba K, Inoue T
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search




 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
