-Search query
-Search result
Showing 1 - 50 of 3,343 items for (author: sun & r)

EMDB-45962: 
Cryo-EM structure of Tulane virus 9-6-17 variant capsid protein VP1 5-12-18
Method: single particle / : Sun C, Jiang W

EMDB-45963: 
Cryo-EM structure of Tulane virus 9-6-17 variant capsid protein VP1 9-14-18
Method: single particle / : Sun C, Jiang W

EMDB-45964: 
Cryo-EM structure of Tulane virus 9-6-17 variant capsid protein VP1 9-14-18, DTT-treated
Method: single particle / : Sun C, Jiang W

EMDB-38845: 
Icosahedrally averaged cryo-EM reconstruction of PhiKZ capsid before applying the "block-based" reconstruction method
Method: single particle / : Yang Y, Shao Q, Guo M, Han L, Zhao X, Wang A, Li X, Wang B, Pan J, Chen Z, Fokine A, Sun L, Fang Q

EMDB-38846: 
Block 1 of PhiKZ capsid
Method: single particle / : Yang Y, Shao Q, Guo M, Han L, Zhao X, Wang A, Li X, Wang B, Pan J, Chen Z, Fokine A, Sun L, Fang Q

EMDB-38848: 
Block 2 of PhiKZ capsid
Method: single particle / : Yang Y, Shao Q, Guo M, Han L, Zhao X, Wang A, Li X, Wang B, Pan J, Chen Z, Fokine A, Sun L, Fang Q

EMDB-39002: 
Composite cryo-EM map of PhiKZ capsid after applying the "block-based" reconstruction method
Method: single particle / : Yang Y, Shao Q, Guo M, Han L, Zhao X, Wang A, Li X, Wang B, Pan J, Chen Z, Fokine A, Sun L, Fang Q

PDB-8y6v: 
Near-atomic structure of icosahedrally averaged jumbo bacteriophage PhiKZ capsid
Method: single particle / : Yang Y, Shao Q, Guo M, Han L, Zhao X, Wang A, Li X, Wang B, Pan J, Chen Z, Fokine A, Sun L, Fang Q
EMDB-37139: 
Structure of SARS-CoV Spike protein complexed with antibody PW5-5
Method: single particle / : Sun L, Mao Q, Wang Y
EMDB-37143: 
The local refined map of SARS-CoV-2 XBB Variant Spike protein complexed with antibody PW5-535
Method: single particle / : Sun L, Mao Q, Wang Y
EMDB-37144: 
Trimer state of SARS-CoV Spike protein complexed with antibody PW5-535
Method: single particle / : Sun L, Mao Q, Wang Y

EMDB-37145: 
The local refined map of SARS-CoV Spike protein complexed with antibody PW5-5
Method: single particle / : Sun L, Mao Q, Wang Y
EMDB-37160: 
Monomer state of SARS-CoV-2 XBB Variant Spike protein trimer complexed with antibody PW5-5
Method: single particle / : Sun L, Mao Q, Wang Y
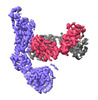
EMDB-37161: 
Monomer state of SARS-CoV Spike protein complexed with antibody PW5-535
Method: single particle / : Sun L, Mao Q, Wang Y
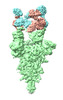
EMDB-37162: 
Structure of SARS-CoV-2 Omicron BA.1 Spike complexed with antibody PW5-570
Method: single particle / : Sun L, Mao Q, Wang Y
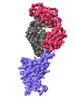
EMDB-37163: 
The local refined map of SARS-CoV-2 Omicron BA.1 Spike complexed with antibody PW5-570
Method: single particle / : Sun L, Mao Q, Wang Y
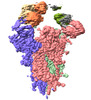
EMDB-37164: 
State 1 of SARS-CoV-2 XBB Variant Spike protein trimer complexed with antibody PW5-5
Method: single particle / : Sun L, Mao Q, Wang Y
EMDB-37165: 
Structure of SARS-CoV-2 XBB Variant Spike protein complexed with broadly neutralizing antibody PW5-535
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8kdm: 
Structure of SARS-CoV Spike protein complexed with antibody PW5-5
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8kdr: 
The local refined map of SARS-CoV-2 XBB Variant Spike protein complexed with antibody PW5-535
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8kds: 
Trimer state of SARS-CoV Spike protein complexed with antibody PW5-535
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8kdt: 
The local refined map of SARS-CoV Spike protein complexed with antibody PW5-5
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8kej: 
Monomer state of SARS-CoV-2 XBB Variant Spike protein trimer complexed with antibody PW5-5
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8kek: 
Monomer state of SARS-CoV Spike protein complexed with antibody PW5-535
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8keo: 
Structure of SARS-CoV-2 Omicron BA.1 Spike complexed with antibody PW5-570
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8kep: 
The local refined map of SARS-CoV-2 Omicron BA.1 Spike complexed with antibody PW5-570
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8keq: 
State 1 of SARS-CoV-2 XBB Variant Spike protein trimer complexed with antibody PW5-5
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8ker: 
Structure of SARS-CoV-2 XBB Variant Spike protein complexed with broadly neutralizing antibody PW5-535
Method: single particle / : Sun L, Mao Q, Wang Y

EMDB-50981: 
Endophilin B1 dimers bound to nanodiscs
Method: single particle / : Thorlacius A, Sundborger-Lunna A

EMDB-50984: 
Endophilin B1 dimer bound to nanodisc center
Method: single particle / : Thorlacius A, Sundborger-Lunna A

PDB-9g2r: 
Endophilin B1 dimers bound to nanodiscs
Method: single particle / : Thorlacius A, Sundborger-Lunna A

PDB-9g2u: 
Endophilin B1 dimer bound to nanodisc center
Method: single particle / : Thorlacius A, Sundborger-Lunna A

EMDB-50986: 
Endophilin B1 dimer bound to nanodisc edge
Method: single particle / : Thorlacius A, Sundborger-Lunna A

PDB-9g2w: 
Endophilin B1 dimer bound to nanodisc edge
Method: single particle / : Thorlacius A, Sundborger-Lunna A
EMDB-43779: 
Rat GluN1-GluN2B NMDA receptor channel in complex with glycine, glutamate, and EU-1622-A, in open-channel conformation
Method: single particle / : Chou TH, Furukawa H
EMDB-43780: 
Rat GluN1-GluN2B NMDA receptor channel in complex with glycine, glutamate, and EU-1622-A, in nonactive1 conformation
Method: single particle / : Chou TH, Furukawa H
EMDB-43781: 
Rat GluN1-GluN2B NMDA receptor channel in apo conformation
Method: single particle / : Chou TH, Furukawa H
EMDB-43782: 
Rat GluN1-GluN2B NMDA receptor channel in complex with glycine
Method: single particle / : Chou TH, Furukawa H

EMDB-43783: 
Rat GluN1-GluN2B NMDA receptor channel in complex with glutamate
Method: single particle / : Chou TH, Furukawa H
EMDB-44586: 
Rat GluN1-GluN2B NMDA receptor channel in complex with glycine, glutamate, and EU-1622-A, in open-channel conformation, C1 symmetry
Method: single particle / : Chou TH, Furukawa H

PDB-9are: 
Rat GluN1-GluN2B NMDA receptor channel in complex with glycine, glutamate, and EU-1622-A, in open-channel conformation
Method: single particle / : Chou TH, Furukawa H

PDB-9arf: 
Rat GluN1-GluN2B NMDA receptor channel in complex with glycine, glutamate, and EU-1622-A, in nonactive1 conformation
Method: single particle / : Chou TH, Furukawa H

PDB-9arg: 
Rat GluN1-GluN2B NMDA receptor channel in apo conformation
Method: single particle / : Chou TH, Furukawa H

PDB-9arh: 
Rat GluN1-GluN2B NMDA receptor channel in complex with glycine
Method: single particle / : Chou TH, Furukawa H

PDB-9ari: 
Rat GluN1-GluN2B NMDA receptor channel in complex with glutamate
Method: single particle / : Chou TH, Furukawa H

PDB-9bib: 
Rat GluN1-GluN2B NMDA receptor channel in complex with glycine, glutamate, and EU-1622-A, in open-channel conformation, C1 symmetry
Method: single particle / : Chou TH, Furukawa H

EMDB-39646: 
the complex structure of the H4B6 Fab with the RBD of Omicron BA.5 S protein
Method: single particle / : Xia LY, Zhang YY, Zhou Q, Yan RH

EMDB-37467: 
SARS-CoV-2 Omicron BQ.1.1 RBD complexed with human ACE2
Method: single particle / : Li W, Xie Y

EMDB-37468: 
SARS-CoV-2 Omicron BQ.1 RBD complexed with human ACE2
Method: single particle / : Li W, Xie Y

EMDB-37469: 
SARS-CoV-2 Omicron XBB RBD complexed with human ACE2
Method: single particle / : Li W, Xie Y
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
