-Search query
-Search result
Showing 1 - 50 of 235 items for (author: schulman & ba)

EMDB-18214: 
Structure of CUL9-RBX1 ubiquitin E3 ligase complex - hexameric assembly
Method: single particle / : Hopf LVM, Horn-Ghetko D, Schulman BA

EMDB-18216: 
Structure of CUL9-RBX1 ubiquitin E3 ligase complex in unneddylated and neddylated conformation - focused cullin dimer
Method: single particle / : Hopf LVM, Horn-Ghetko D, Schulman BA

EMDB-18217: 
Structure of CUL9-RBX1 ubiquitin E3 ligase complex in unneddylated and neddylated conformation - focused on E2-like density
Method: single particle / : Hopf LVM, Horn-Ghetko D, Schulman BA

EMDB-18218: 
Structure of CUL9-RBX1 ubiquitin E3 ligase complex in unneddylated and neddylated conformation - focused dimeric core
Method: single particle / : Hopf LVM, Horn-Ghetko D, Schulman BA

EMDB-18220: 
Structure of the hexameric CUL9-RBX1 complex with deletion of CUL9 CPH domain
Method: single particle / : Hopf LVM, Horn-Ghetko D, Schulman BA

EMDB-18221: 
Structure of the hexameric CUL9-RBX1 complex with deletion of CUL9 DOC domain
Method: single particle / : Hopf LVM, Horn-Ghetko D, Schulman BA

EMDB-18222: 
Structure of the hexameric CUL9-RBX1 complex with deletion of CUL9 ARM9 domain
Method: single particle / : Hopf LVM, Horn-Ghetko D, Schulman BA

EMDB-18223: 
Structure of the hexameric CUL9-RBX1 complex with deletion of CUL9 ARIH-RBR element
Method: single particle / : Hopf LVM, Horn-Ghetko D, Schulman BA

EMDB-19179: 
Structure of CUL9-RBX1 ubiquitin E3 ligase complex in unneddylated conformation - symmetry expanded unneddylated dimer
Method: single particle / : Hopf LVM, Horn-Ghetko D, Prabu JR, Schulman BA

PDB-8q7e: 
Structure of CUL9-RBX1 ubiquitin E3 ligase complex - hexameric assembly
Method: single particle / : Hopf LVM, Horn-Ghetko D, Schulman BA

PDB-8q7h: 
Structure of CUL9-RBX1 ubiquitin E3 ligase complex in unneddylated and neddylated conformation - focused cullin dimer
Method: single particle / : Hopf LVM, Horn-Ghetko D, Schulman BA

PDB-8rhz: 
Structure of CUL9-RBX1 ubiquitin E3 ligase complex in unneddylated conformation - symmetry expanded unneddylated dimer
Method: single particle / : Hopf LVM, Horn-Ghetko D, Prabu JR, Schulman BA

EMDB-19039: 
Map of YPEL5-bound WDR26 dimer obtained by focused refinement of the WDR26-CTLH subcomplex
Method: single particle / : Chrustowicz J, Schulman BA

EMDB-19856: 
Focused map 1- K48-linked ubiquitin chain formation with a cullin-RING E3 ligase & Cdc34: NEDD8-CUL2-RBX1-ELOB/C-FEM1C with trapped UBE2R2~donor UB~acceptor UB-SIL1 peptide
Method: single particle / : Liwocha J, Prabu JR, Kleiger G, Schulman BA

EMDB-19857: 
Focused map 2 - K48-linked ubiquitin chain formation with a cullin-RING E3 ligase & Cdc34: NEDD8-CUL2-RBX1-ELOB/C-FEM1C with trapped UBE2R2~donor UB~acceptor UB-SIL1 peptide
Method: single particle / : Liwocha J, Prabu JR, Kleiger G, Schulman BA

EMDB-19858: 
Focused map 3 - K48-linked ubiquitin chain formation with a cullin-RING E3 ligase & Cdc34: NEDD8-CUL2-RBX1-ELOB/C-FEM1C with trapped UBE2R2~donor UB~acceptor UB-SIL1 peptide
Method: single particle / : Liwocha J, Prabu JR, Kleiger G, Schulman BA

EMDB-19859: 
Focused map 4 - K48-linked ubiquitin chain formation with a cullin-RING E3 ligase & Cdc34: NEDD8-CUL2-RBX1-ELOB/C-FEM1C with trapped UBE2R2~donor UB~acceptor UB-SIL1 peptide
Method: single particle / : Liwocha J, Prabu JR, Kleiger G, Schulman BA

EMDB-19860: 
Focused map 5 - K48-linked ubiquitin chain formation with a cullin-RING E3 ligase & Cdc34: NEDD8-CUL2-RBX1-ELOB/C-FEM1C with trapped UBE2R2~donor UB~acceptor UB-SIL1 peptide
Method: single particle / : Liwocha J, Prabu JR, Kleiger G, Schulman BA

EMDB-19342: 
Human 20S proteasome assembly intermediate structure 5 beta7 tagged
Method: single particle / : Schulman BA, Hanna JW, Harper JW, Adolf F, Du J, Rawson SD, Walsh Jr RM, Goodall EA

EMDB-19343: 
Human 20S core particle beta7 tagged
Method: single particle / : Schulman BA, Hanna JW, Harper JW, Adolf F, Du J, Rawson SD, Walsh Jr RM, Goodall EA

EMDB-18755: 
Human 20S proteasome assembly structure 1
Method: single particle / : Schulman BA, Hanna JW, Harper JW, Adolf F, Du J, Rawson SD, Walsh Jr RM, Goodall EA

EMDB-18757: 
Human 20S proteasome assembly intermediate structure 2
Method: single particle / : Schulman BA, Hanna JW, Harper JW, Adolf F, Du J, Rawson SD, Walsh Jr RM, Goodall EA

EMDB-18758: 
Human 20S proteasome assembly intermediate structure 3
Method: single particle / : Schulman BA, Hanna JW, Harper JW, Adolf F, Du J, Rawson SD, Walsh Jr RM, Goodall EA

EMDB-18759: 
Human 20S proteasome assembly intermediate structure 5
Method: single particle / : Schulman BA, Hanna JW, Harper JW, Adolf F, Du J, Rawson SD, Walsh Jr RM, Goodall EA
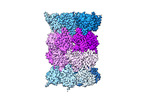
EMDB-18760: 
Human proteasome 20S core particle
Method: single particle / : Schulman BA, Hanna JW, Harper JW, Adolf F, Du J, Rawson SD, Walsh Jr RM, Goodall EA

EMDB-18761: 
Human preholo proteasome 20S core particle
Method: single particle / : Schulman BA, Hanna JW, Harper JW, Adolf F, Du J, Rawson SD, Walsh Jr RM, Goodall EA

EMDB-18762: 
Human premature proteasome 20S core particle
Method: single particle / : Schulman BA, Hanna JW, Harper JW, Adolf F, Du J, Rawson SD, Walsh Jr RM, Goodall EA

EMDB-18773: 
Human 20S proteasome assembly intermediate structure 4
Method: single particle / : Schulman BA, Hanna JW, Harper JW, Adolf F, Du J, Rawson SD, Walsh Jr RM, Goodall EA

EMDB-18207: 
Ubiquitin ligation to substrate by a cullin-RING E3 ligase & Cdc34: NEDD8-CUL2-RBX1-ELOB/C-FEM1C with trapped UBE2R2~donor UB-Sil1 peptide, Glacios map
Method: single particle / : Liwocha J, Prabu JR, Kleiger G, Schulman BA

EMDB-18230: 
Ubiquitin ligation to substrate by a cullin-RING E3 ligase & Cdc34: NEDD8-CUL2-RBX1-ELOB/C-FEM1C with trapped UBE2R2~donor UB-Sil1 peptide
Method: single particle / : Liwocha J, Prabu JR, Kleiger G, Schulman BA
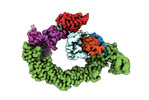
EMDB-18915: 
Ubiquitin ligation to neosubstrate by a cullin-RING E3 ligase & Cdc34: NEDD8-CUL2-RBX1-ELOB/C-VHL-MZ1 with trapped UBE2R2~donor UB-BRD4 BD2
Method: single particle / : Liwocha J, Prabu JR, Kleiger G, Schulman BA

PDB-8q7r: 
Ubiquitin ligation to substrate by a cullin-RING E3 ligase & Cdc34: NEDD8-CUL2-RBX1-ELOB/C-FEM1C with trapped UBE2R2~donor UB-Sil1 peptide
Method: single particle / : Liwocha J, Prabu JR, Kleiger G, Schulman BA

PDB-8r5h: 
Ubiquitin ligation to neosubstrate by a cullin-RING E3 ligase & Cdc34: NEDD8-CUL2-RBX1-ELOB/C-VHL-MZ1 with trapped UBE2R2~donor UB-BRD4 BD2
Method: single particle / : Liwocha J, Prabu JR, Kleiger G, Schulman BA
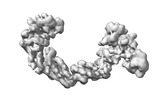
EMDB-17798: 
CUL2-RBX1-ELOB/C-FEM1C-SIL1 conformation 1
Method: single particle / : Liwocha J, Prabu JR, Kleiger G, Schulman BA

EMDB-17799: 
CUL2-RBX1-ELOB/C-FEM1C-SIL1 conformation 2
Method: single particle / : Liwocha J, Prabu JR, Kleiger G, Schulman BA
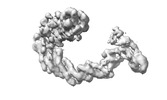
EMDB-17800: 
NEDD8-CUL2-RBX1-ELOB/C-FEM1C-SIL1 conformation 1
Method: single particle / : Liwocha J, Prabu JR, Kleiger G, Schulman BA

EMDB-17801: 
NEDD8-CUL2-RBX1-ELOB/C-FEM1C-SIL1 conformation 2
Method: single particle / : Liwocha J, Prabu JR, Kleiger G, Schulman BA

EMDB-17802: 
K48-linked ubiquitin chain formation with a cullin-RING E3 ligase and Cdc34: NEDD8-CUL1-RBX1-SKP1-FBXW7 with trapped UBE2R2-donor UB-acceptor UB-cyclin E peptide
Method: single particle / : Liwocha J, Prabu JR, Kleiger G, Schulman BA

EMDB-17803: 
Consensus map - K48-linked ubiquitin chain formation with a cullin-RING E3 ligase and Cdc34: NEDD8-CUL2-RBX1-ELOB/C-FEM1C with trapped UBE2R2-donor UB-acceptor UB-SIL1 peptide
Method: single particle / : Liwocha J, Prabu JR, Kleiger G, Schulman BA

EMDB-17822: 
K48-linked ubiquitin chain formation with a cullin-RING E3 ligase and Cdc34: NEDD8-CUL2-RBX1-ELOB/C-FEM1C with trapped UBE2R2-donor UB-acceptor UB-SIL1 peptide
Method: single particle / : Liwocha J, Prabu JR, Kleiger G, Schulman BA

EMDB-18767: 
K48-linked ubiquitin chain formation with a cullin-RING E3 ligase & Cdc34: NEDD8-CUL2-RBX1-ELOB/C-VHL-MZ1 with trapped UBE2R2~donor UB~acceptor UB-BRD4
Method: single particle / : Liwocha J, Prabu JR, Kleiger G, Schulman BA

PDB-8pql: 
K48-linked ubiquitin chain formation with a cullin-RING E3 ligase and Cdc34: NEDD8-CUL2-RBX1-ELOB/C-FEM1C with trapped UBE2R2-donor UB-acceptor UB-SIL1 peptide
Method: single particle / : Liwocha J, Prabu JR, Kleiger G, Schulman BA

EMDB-19194: 
In situ cryo-electron tomogram of an autophagosome in the projection of an iPSC-derived neuron #2
Method: electron tomography / : Hoyer MJ, Capitanio C, Smith IR, Paoli JC, Bieber A, Jiang Y, Paulo JA, Gonzalez-Lozano MA, Baumeister W, Wilfling F, Schulman BA, Harper WJ

EMDB-19346: 
In situ cryo-electron tomogram of an autophagosome in the projection of an iPSC-derived neuron #1
Method: electron tomography / : Hoyer MJ, Capitanio C, Smith IR, Paoli JC, Bieber A, Jiang Y, Paulo JA, Gonzalez-Lozano MA, Baumeister W, Wilfling F, Schulman BA, Harper WJ

EMDB-17597: 
cryo-EM structure of Doa10 in MSP1E3D1
Method: single particle / : Botsch JJ, Braeuning B, Schulman BA

EMDB-17608: 
cryo-EM structure of Doa10 with RING domain in MSP1E3D1
Method: single particle / : Botsch JJ, Braeuning B, Schulman BA

EMDB-17609: 
Low resolution map of Doa10 in MSP1E3D1
Method: single particle / : Botsch JJ, Schulman BA

EMDB-17610: 
Doa10 in MSP2N2
Method: single particle / : Botsch JJ, Schulman BA

PDB-8pd0: 
cryo-EM structure of Doa10 in MSP1E3D1
Method: single particle / : Botsch JJ, Braeuning B, Schulman BA

PDB-8pda: 
cryo-EM structure of Doa10 with RING domain in MSP1E3D1
Method: single particle / : Botsch JJ, Braeuning B, Schulman BA
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
