-Search query
-Search result
Showing 1 - 50 of 184 items for (author: robert & b & sim)
EMDB-19184: 
Late alpha-Synuclein fibril structure from liquid-liquid phase separations.
Method: helical / : De Simone A, Barritt JD, Chen S, Cascella R, Cecchi C, Bigi A, Jarvis JA, Chiti F, Dobson CM, Fusco G

PDB-8ri9: 
Late alpha-Synuclein fibril structure from liquid-liquid phase separations.
Method: helical / : De Simone A, Barritt JD, Chen S, Cascella R, Cecchi C, Bigi A, Jarvis JA, Chiti F, Dobson CM, Fusco G

EMDB-16976: 
Human tRNA guanine transglycosylase (TGT) bound to tRNAAsp
Method: single particle / : Sievers K, Neumann P, Susac L, Trowitzsch S, Tampe R, Ficner R

PDB-8omr: 
Human tRNA guanine transglycosylase (TGT) bound to tRNAAsp
Method: single particle / : Sievers K, Neumann P, Susac L, Trowitzsch S, Tampe R, Ficner R

EMDB-36891: 
96nm repeat of human respiratory doublet microtubule, IDAf local refined
Method: single particle / : Gui M, Brown A

EMDB-36895: 
Consensus map of 96nm repeat of human respiratory doublet microtubule, RS3 region
Method: single particle / : Gui M, Brown A

EMDB-16328: 
Outer membrane attachment porin OmpM1 from Veillonella parvula
Method: single particle / : Silale A, van den Berg B

EMDB-16332: 
Outer membrane attachment porin OmpM1 from Veillonella parvula, native
Method: single particle / : Silale A, van den Berg B

EMDB-16333: 
Outer membrane attachment porin OmpM1 from Veillonella parvula, C3 symmetry
Method: single particle / : Silale A, van den Berg B

PDB-8bym: 
Outer membrane attachment porin OmpM1 from Veillonella parvula
Method: single particle / : Silale A, van den Berg B

PDB-8bys: 
Outer membrane attachment porin OmpM1 from Veillonella parvula, native
Method: single particle / : Silale A, van den Berg B

PDB-8byt: 
Outer membrane attachment porin OmpM1 from Veillonella parvula, C3 symmetry
Method: single particle / : Silale A, van den Berg B

EMDB-16569: 
PfRH5-PfCyRPA-PfRIPR complex from Plasmodium falciparum bound to antibody Cy.003
Method: single particle / : Farrell B, Higgins MK
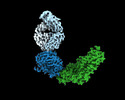
EMDB-16570: 
PfCyRPA-PfRIPR complex from Plasmodium falciparum bound to antibody Cy.003
Method: single particle / : Farrell B, Higgins MK

EMDB-16635: 
PfCyRPA-PfRIPR complex from Plasmodium falciparum bound to antibody Cy.003 - maps for local refinement on PfRIPR
Method: single particle / : Farrell B, Higgins MK

EMDB-16636: 
PfCyRPA-PfRIPR complex from Plasmodium falciparum bound to antibody Cy.003 consensus maps
Method: single particle / : Farrell B, Higgins MK

EMDB-16637: 
PfRH5-PfCyRPA-PfRIPR complex from Plasmodium falciparum bound to antibody Cy.003 - consensus map
Method: single particle / : Farrell B, Higgins MK

EMDB-16638: 
PfRH5-PfCyRPA-PfRIPR complex from Plasmodium falciparum bound to antibody Cy.003 - local refinement on PfRH5
Method: single particle / : Farrell B, Higgins MK

EMDB-16639: 
PfRH5-PfCyRPA-PfRIPR complex from Plasmodium falciparum bound to antibody Cy.003 - local refinement on PfRIPR
Method: single particle / : Farrell B, Higgins MK

EMDB-16640: 
PfRH5-PfCyRPA-PfRIPR complex from Plasmodium falciparum bound to antibody Cy.003 - map with additional PfRIPR tail density
Method: single particle / : Farrell B, Higgins MK

PDB-8cdd: 
PfRH5-PfCyRPA-PfRIPR complex from Plasmodium falciparum bound to antibody Cy.003
Method: single particle / : Farrell B, Higgins MK

PDB-8cde: 
PfCyRPA-PfRIPR complex from Plasmodium falciparum bound to antibody Cy.003
Method: single particle / : Farrell B, Higgins MK
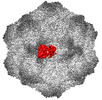
EMDB-26786: 
CPV Affinity Purified Polyclonal Fab A Site Fab
Method: single particle / : Hartmann SR, Hafenstein SL, Charnesky AJ
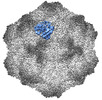
EMDB-26787: 
CPV Affinity Purified Polyclonal Fab B Site Fab
Method: single particle / : Hartmann SR, Hafenstein SL, Charnesky AJ
EMDB-26788: 
CPV Total-Fab Polyclonal A Site Fab
Method: single particle / : Hartmann SR, Hafenstein SL, Charnesky AJ
EMDB-26789: 
CPV Total-Fab Polyclonal B Site Fab (1 of 2)
Method: single particle / : Hartmann SR, Hafenstein SL, Charnesky AJ
EMDB-26790: 
CPV Total-Fab Polyclonal B Site Fab (2 of 2)
Method: single particle / : Hartmann SR, Hafenstein SL, Charnesky AJ

PDB-7utp: 
CPV Affinity Purified Polyclonal Fab A Site Fab
Method: single particle / : Hartmann SR, Hafenstein SL, Charnesky AJ

PDB-7utr: 
CPV Affinity Purified Polyclonal Fab B Site Fab
Method: single particle / : Hartmann SR, Hafenstein SL, Charnesky AJ

PDB-7uts: 
CPV Total-Fab Polyclonal A Site Fab
Method: single particle / : Hartmann SR, Hafenstein SL, Charnesky AJ

PDB-7utu: 
CPV Total-Fab Polyclonal B Site Fab (1 of 2)
Method: single particle / : Hartmann SR, Hafenstein SL, Charnesky AJ

PDB-7utv: 
CPV Total-Fab Polyclonal B Site Fab (2 of 2)
Method: single particle / : Hartmann SR, Hafenstein SL, Charnesky AJ
EMDB-28013: 
Cryo-EM structure of the Glutaminase C core filament (fGAC)
Method: single particle / : Ambrosio AL, Dias SM, Quesnay JE, Portugal RV, Cassago A, van Heel MG, Islam Z, Rodrigues CT

PDB-8ec6: 
Cryo-EM structure of the Glutaminase C core filament (fGAC)
Method: single particle / : Ambrosio AL, Dias SM, Quesnay JE, Portugal RV, Cassago A, van Heel MG, Islam Z, Rodrigues CT

EMDB-16963: 
Leishmania tarentolae proteasome 20S subunit in complex with 1-Benzyl-N-(3-(cyclopropylcarbamoyl)phenyl)-6-oxo-1,6-dihydropyridazine-3-carboxamide
Method: single particle / : Rowland P

PDB-8olu: 
Leishmania tarentolae proteasome 20S subunit in complex with 1-Benzyl-N-(3-(cyclopropylcarbamoyl)phenyl)-6-oxo-1,6-dihydropyridazine-3-carboxamide
Method: single particle / : Rowland P

EMDB-35888: 
96nm repeat of human respiratory doublet microtubule and associated axonemal complexes
Method: single particle / : Gui M, Brown A

PDB-8j07: 
96nm repeat of human respiratory doublet microtubule and associated axonemal complexes
Method: single particle / : Gui M, Brown A

EMDB-40220: 
96-nm repeat unit of doublet microtubules from Chlamydomonas reinhardtii flagella
Method: single particle / : Walton T, Brown A

PDB-8glv: 
96-nm repeat unit of doublet microtubules from Chlamydomonas reinhardtii flagella
Method: single particle / : Walton T, Brown A

EMDB-14923: 
Structure of the human tRNA splicing endonuclease defines substrate recognition
Method: single particle / : Sekulovski S, Trowitzsch S

EMDB-14925: 
Human TSEN with pre-tRNA-Tyr-GTA
Method: single particle / : Sekulovski S, Trowitzsch S

PDB-7zrz: 
Structure of the human tRNA splicing endonuclease defines substrate recognition
Method: single particle / : Sekulovski S, Trowitzsch S

EMDB-26809: 
KDM2A-nucleosome structure stabilized by H3K36C-UNC8015 covalent conjugate
Method: single particle / : Spangler CJ, Skrajna A, Foley CA, Budziszewski GR, Azzam DN, James LI, Frye SV, McGinty RK

EMDB-26810: 
KDM2B-nucleosome complex stabilized by H3K36C-UNC8015 covalent conjugate
Method: single particle / : Spangler CJ, Skrajna A, Foley CA, Budziszewski GR, Azzam DN, James LI, Frye SV, McGinty RK

PDB-7uv9: 
KDM2A-nucleosome structure stabilized by H3K36C-UNC8015 covalent conjugate
Method: single particle / : Spangler CJ, Skrajna A, Foley CA, Budziszewski GR, Azzam DN, James LI, Frye SV, McGinty RK

EMDB-27296: 
E. coli ATP synthase imaged in 10mM MgATP State1
Method: single particle / : Sobti M, Stewart AG

EMDB-27297: 
E. coli ATP synthase imaged in 10mM MgATP State1 "half-up
Method: single particle / : Sobti M, Stewart AG
EMDB-27298: 
E. coli ATP synthase imaged in 10mM MgATP State1 "half-up" Fo classified
Method: single particle / : Sobti M, Stewart AG

EMDB-27299: 
E. coli ATP synthase imaged in 10mM MgATP State 1 "half-up" Fo refine
Method: single particle / : Sobti M, Stewart AG
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
