-Search query
-Search result
Showing all 24 items for (author: pierson & ee)
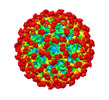

EMDB-29020: 
Structure of dengue virus (DENV2) in complex with prM12, an anti-PrM monoclonal antibody
Method: single particle / : Dowd AD, Sirohi D, Speer S, Mukherjee S, Govero J, Aleshnick M, Larman B, Sukupolvi-Petty S, Sevvana M, Miller AS, Klose T, Zheng A, Kielian M, Kuhn RJ, Diamond MS, Pierson TC

EMDB-29021: 
Structure of dengue virus (DENV2) in complex with prM13, an anti-PrM monoclonal antibody
Method: single particle / : Dowd AD, Sirohi D, Speer S, Mukherjee S, Govero J, Aleshnick M, Larman B, Sukupolvi-Petty S, Sevvana M, Miller AS, Klose T, Zheng A, Kielian M, Kuhn RJ, Diamond MS, Pierson TC

PDB-8fe3: 
Structure of dengue virus (DENV2) in complex with prM12, an anti-PrM monoclonal antibody
Method: single particle / : Dowd AD, Sirohi D, Speer S, Mukherjee S, Govero J, Aleshnick M, Larman B, Sukupolvi-Petty S, Sevvana M, Miller AS, Klose T, Zheng A, Kielian M, Kuhn RJ, Diamond MS, Pierson TC

PDB-8fe4: 
Structure of dengue virus (DENV2) in complex with prM13, an anti-PrM monoclonal antibody
Method: single particle / : Dowd AD, Sirohi D, Speer S, Mukherjee S, Govero J, Aleshnick M, Larman B, Sukupolvi-Petty S, Sevvana M, Miller AS, Klose T, Zheng A, Kielian M, Kuhn RJ, Diamond MS, Pierson TC
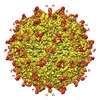
EMDB-25606: 
Zika Virus particle bound with IgM antibody DH1017 Fab fragment
Method: single particle / : Miller AS, Kuhn RJ

EMDB-3266: 
Importin-beta can bind Hepatitis B Virus core protein and empty core-like particles and induce structural changes
Method: single particle / : Chen C, Wang JC-Y, Pierson EE, Kiefer DZ, Delaleau M, Gallucci L, Cazenave C, Kann M, Jarrold MF, Zlotnick A

EMDB-3267: 
Importin-beta can bind Hepatitis B Virus core protein and empty core-like particles and induce structural changes
Method: single particle / : Chen C, Wang JC-Y, Pierson EE, Kiefer DZ, Delaleau M, Gallucci L, Cazenave C, Kann M, Jarrold MF, Zlotnick A

EMDB-3268: 
Importin-beta can bind Hepatitis B Virus core protein and empty core-like particles and induce structural changes
Method: single particle / : Chen C, Wang JC-Y, Pierson EE, Kiefer DZ, Delaleau M, Gallucci L, Cazenave C, Kann M, Jarrold MF, Zlotnick A

EMDB-3269: 
Importin-beta can bind Hepatitis B Virus core protein and empty core-like particles and induce structural changes
Method: single particle / : Chen C, Wang JC-Y, Pierson EE, Kiefer DZ, Delaleau M, Gallucci L, Cazenave C, Kann M, Jarrold MF, Zlotnick A

EMDB-3270: 
Importin-beta can bind Hepatitis B Virus core protein and empty core-like particles and induce structural changes
Method: single particle / : Chen C, Wang JC-Y, Pierson EE, Kiefer DZ, Delaleau M, Gallucci L, Cazenave C, Kann M, Jarrold MF, Zlotnick A

EMDB-3271: 
Importin-beta can bind Hepatitis B Virus core protein and empty core-like particles and induce structural changes
Method: single particle / : Chen C, Wang JC-Y, Pierson EE, Kiefer DZ, Delaleau M, Gallucci L, Cazenave C, Kann M, Jarrold MF, Zlotnick A

EMDB-3272: 
Importin-beta can bind Hepatitis B Virus core protein and empty core-like particles and induce structural changes
Method: single particle / : Chen C, Wang JC-Y, Pierson EE, Kiefer DZ, Delaleau M, Gallucci L, Cazenave C, Kann M, Jarrold MF, Zlotnick A

EMDB-6630: 
Glutamate dehydrogenase in the unliganded state
Method: single particle / : Borgnia MJ, Banerjee S, Merk A, Matthies D, Bartesaghi A, Rao P, Pierson J, Earl LA, Falconieri V, Subramaniam S, Milne JLS

EMDB-6631: 
Glutamate dehydrogenase in complex with GTP
Method: single particle / : Borgnia MJ, Banerjee S, Merk A, Matthies D, Bartesaghi A, Rao P, Pierson J, Earl LA, Falconieri V, Subramaniam S, Milne JLS

EMDB-6632: 
Glutamate dehydrogenase in complex with NADH and GTP, open conformation
Method: single particle / : Borgnia MJ, Banerjee S, Merk A, Matthies D, Bartesaghi A, Rao P, Pierson J, Earl LA, Falconieri V, Subramaniam S, Milne JLS

EMDB-6633: 
Glutamate dehydrogenase in complex with NADH and GTP, closed conformation
Method: single particle / : Borgnia MJ, Banerjee S, Merk A, Matthies D, Bartesaghi A, Rao P, Pierson J, Earl LA, Falconieri V, Subramaniam S, Milne JLS

EMDB-6634: 
Glutamate dehydrogenase in complex with NADH, closed conformation
Method: single particle / : Borgnia MJ, Banerjee S, Merk A, Matthies D, Bartesaghi A, Rao P, Pierson J, Earl LA, Falconieri V, Subramaniam S, Milne JLS

EMDB-6635: 
Glutamate dehydrogenase in complex with NADH, open conformation
Method: single particle / : Borgnia MJ, Banerjee S, Merk A, Matthies D, Bartesaghi A, Rao P, Pierson J, Earl LA, Falconieri V, Subramaniam S, Milne JLS

PDB-3jcz: 
Structure of bovine glutamate dehydrogenase in the unliganded state
Method: single particle / : Borgnia MJ, Banerjee S, Merk A, Matthies D, Bartesaghi A, Rao P, Pierson J, Earl LA, Falconieri V, Subramaniam S, Milne JLS

PDB-3jd0: 
Glutamate dehydrogenase in complex with GTP
Method: single particle / : Borgnia MJ, Banerjee S, Merk A, Matthies D, Bartesaghi A, Rao P, Pierson J, Earl LA, Falconieri V, Subramaniam S, Milne JLS

PDB-3jd1: 
Glutamate dehydrogenase in complex with NADH, closed conformation
Method: single particle / : Borgnia MJ, Banerjee S, Merk A, Matthies D, Bartesaghi A, Rao P, Pierson J, Earl LA, Falconieri V, Subramaniam S, Milne JLS

PDB-3jd2: 
Glutamate dehydrogenase in complex with NADH, open conformation
Method: single particle / : Borgnia MJ, Banerjee S, Merk A, Matthies D, Bartesaghi A, Rao P, Pierson J, Earl LA, Falconieri V, Subramaniam S, Milne JLS

PDB-3jd3: 
Glutamate dehydrogenase in complex with NADH and GTP, open conformation
Method: single particle / : Borgnia MJ, Banerjee S, Merk A, Matthies D, Bartesaghi A, Rao P, Pierson J, Earl LA, Falconieri V, Subramaniam S, Milne JLS

PDB-3jd4: 
Glutamate dehydrogenase in complex with NADH and GTP, closed conformation
Method: single particle / : Borgnia MJ, Banerjee S, Merk A, Matthies D, Bartesaghi A, Rao P, Pierson J, Earl LA, Falconieri V, Subramaniam S, Milne JLS
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
