-Search query
-Search result
Showing all 49 items for (author: perry & k)

EMDB-15525: 
Cryo-EM structure of the RecA postsynaptic filament from S. pneumoniae
Method: helical / : Perry TN, Fronzes R, Polard P, Hertzog M

PDB-8amf: 
Cryo-EM structure of the RecA postsynaptic filament from S. pneumoniae
Method: helical / : Perry TN, Fronzes R, Polard P, Hertzog M

EMDB-41075: 
SARS-CoV-2 spike in complex with Fab 71281-33
Method: single particle / : Binshtein E, Crowe JE

EMDB-41076: 
SARS-CoV-2 spike in complex with Fab 71281-33 (2)
Method: single particle / : Binshtein E, Crowe JE

EMDB-15524: 
Cryo-EM structure of the RecA presynaptic filament from S.pneumoniae
Method: helical / : Perry TN, Fronzes R, Polard P, Hertzog M

PDB-8amd: 
Cryo-EM structure of the RecA presynaptic filament from S.pneumoniae
Method: helical / : Perry TN, Fronzes R, Polard P, Hertzog M

EMDB-26639: 
SARS-CoV-2 replication-transcription complex bound to Remdesivir triphosphate, in a pre-catalytic state
Method: single particle / : Malone BF, Perry JK, Appleby TC, Feng JY, Campbell EA, Darst SA

EMDB-26641: 
SARS-CoV-2 replication-transcription complex bound to ATP, in a pre-catalytic state
Method: single particle / : Malone BF, Perry JK, Appleby TC, Feng JY, Campbell EA, Darst SA

EMDB-26642: 
SARS-CoV-2 replication-transcription complex bound to UTP, in a pre-catalytic state
Method: single particle / : Malone BF, Perry JK, Appleby TC, Feng JY, Campbell EA, Darst SA

EMDB-26645: 
SARS-CoV-2 replication-transcription complex bound to GTP, in a pre-catalytic state
Method: single particle / : Malone BF, Perry JK, Appleby TC, Feng JY, Campbell EA, Darst SA

EMDB-26646: 
SARS-CoV-2 replication-transcription complex bound to CTP, in a pre-catalytic state
Method: single particle / : Malone BF, Perry JK, Appleby TC, Feng JY, Campbell EA, Darst SA

PDB-7uo4: 
SARS-CoV-2 replication-transcription complex bound to Remdesivir triphosphate, in a pre-catalytic state
Method: single particle / : Malone BF, Perry JK, Appleby TC, Feng JY, Campbell EA, Darst SA

PDB-7uo7: 
SARS-CoV-2 replication-transcription complex bound to ATP, in a pre-catalytic state
Method: single particle / : Malone BF, Perry JK, Appleby TC, Feng JY, Campbell EA, Darst SA

PDB-7uo9: 
SARS-CoV-2 replication-transcription complex bound to UTP, in a pre-catalytic state
Method: single particle / : Malone BF, Perry JK, Appleby TC, Feng JY, Campbell EA, Darst SA

PDB-7uob: 
SARS-CoV-2 replication-transcription complex bound to GTP, in a pre-catalytic state
Method: single particle / : Malone BF, Perry JK, Appleby TC, Feng JY, Campbell EA, Darst SA

PDB-7uoe: 
SARS-CoV-2 replication-transcription complex bound to CTP, in a pre-catalytic state
Method: single particle / : Malone BF, Perry JK, Appleby TC, Feng JY, Campbell EA, Darst SA
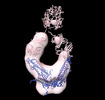
EMDB-26217: 
Negative stain EM map of COVA1-07 mAb bound to the S2 domain of SARS-CoV-2 S
Method: single particle / : Han J, Ward AB
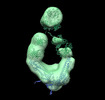
EMDB-26218: 
Negative stain EM map of COVA2-14 mAb bound to the S2 domain of SARS-CoV-2 S
Method: single particle / : Han J, Ward AB

EMDB-26219: 
Negative stain EM map of COVA2-18 mAb bound to the S2 domain of SARS-CoV-2 S
Method: single particle / : Han J, Ward AB
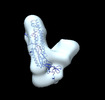
EMDB-26220: 
Negative stain EM map of the S2 domain of SARS-CoV-2 S
Method: single particle / : Han J, Ward AB

EMDB-26667: 
Cryo-EM structure of SHOC2-PP1c-MRAS holophosphatase complex
Method: single particle / : Fuller JR, Hajian B, Lemke C, Kwon J, Bian Y, Aguirre A

PDB-7upi: 
Cryo-EM structure of SHOC2-PP1c-MRAS holophosphatase complex
Method: single particle / : Fuller JR, Hajian B, Lemke C, Kwon J, Bian Y, Aguirre A

EMDB-24427: 
SARS-CoV-2 replication-transcription complex bound to nsp13 helicase - nsp13(2)-RTC - engaged class
Method: single particle / : Chen J, Malone B, Campbell EA, Darst SA

EMDB-24429: 
SARS-CoV-2 replication-transcription complex bound to nsp13 helicase - nsp13(2)-RTC - swiveled class
Method: single particle / : Chen J, Malone B, Campbell EA, Darst SA

EMDB-24430: 
SARS-CoV-2 replication-transcription complex bound to nsp13 helicase - nsp13(2)-RTC (composite)
Method: single particle / : Chen J, Malone B, Campbell EA, Darst SA

EMDB-24431: 
SARS-CoV-2 replication-transcription complex bound to nsp13 helicase - nsp13(1)-RTC
Method: single particle / : Chen J, Malone B, Campbell EA, Darst SA

PDB-7rdy: 
SARS-CoV-2 replication-transcription complex bound to nsp13 helicase - nsp13(2)-RTC - engaged class
Method: single particle / : Chen J, Malone B, Campbell EA, Darst SA

PDB-7re0: 
SARS-CoV-2 replication-transcription complex bound to nsp13 helicase - nsp13(2)-RTC - swiveled class
Method: single particle / : Chen J, Malone B, Campbell EA, Darst SA

PDB-7re1: 
SARS-CoV-2 replication-transcription complex bound to nsp13 helicase - nsp13(2)-RTC (composite)
Method: single particle / : Chen J, Malone B, Campbell EA, Darst SA

PDB-7re2: 
SARS-CoV-2 replication-transcription complex bound to nsp13 helicase - nsp13(1)-RTC
Method: single particle / : Chen J, Malone B, Campbell EA, Darst SA

EMDB-24426: 
SARS-CoV-2 replication-transcription complex bound to nsp13 helicase - nsp13(2)-RTC - open class
Method: single particle / : Chen J, Malone B, Campbell EA, Darst SA

EMDB-24428: 
SARS-CoV-2 replication-transcription complex bound to nsp13 helicase - nsp13(2)-RTC - apo class
Method: single particle / : Chen J, Malone B, Campbell EA, Darst SA

EMDB-24432: 
SARS-CoV-2 replication-transcription complex bound to nsp13 helicase - nsp13(2)-RTC dimer
Method: single particle / : Chen J, Malone B, Campbell EA, Darst SA

PDB-7rdx: 
SARS-CoV-2 replication-transcription complex bound to nsp13 helicase - nsp13(2)-RTC - open class
Method: single particle / : Chen J, Malone B, Campbell EA, Darst SA

PDB-7rdz: 
SARS-CoV-2 replication-transcription complex bound to nsp13 helicase - nsp13(2)-RTC - apo class
Method: single particle / : Chen J, Malone B, Campbell EA, Darst SA

PDB-7re3: 
SARS-CoV-2 replication-transcription complex bound to nsp13 helicase - nsp13(2)-RTC dimer
Method: single particle / : Chen J, Malone B, Campbell EA, Darst SA

EMDB-12693: 
Cryo-EM structure of an Escherichia coli TnaC-ribosome complex stalled in response to L-tryptophan
Method: single particle / : van der Stel AX, Gordon ER

EMDB-12694: 
Cryo-EM structure of an Escherichia coli TnaC(R23F)-ribosome complex stalled in response to L-tryptophan
Method: single particle / : van der Stel AX, Gordon ER

EMDB-12695: 
Cryo-EM structure of an Escherichia coli TnaC(R23F)-ribosome-RF2 complex stalled in response to L-tryptophan
Method: single particle / : van der Stel AX, Gordon ER

PDB-7o19: 
Cryo-EM structure of an Escherichia coli TnaC-ribosome complex stalled in response to L-tryptophan
Method: single particle / : van der Stel AX, Gordon ER, Sengupta A, Martinez AK, Klepacki D, Perry TN, Herrero del Valle A, Vazquez-Laslop N, Sachs MS, Cruz-Vera LR, Innis CA

PDB-7o1a: 
Cryo-EM structure of an Escherichia coli TnaC(R23F)-ribosome complex stalled in response to L-tryptophan
Method: single particle / : van der Stel AX, Gordon ER, Sengupta A, Martinez AK, Klepacki D, Perry TN, Herrero del Valle A, Vazquez-Laslop N, Sachs MS, Cruz-Vera LR, Innis CA

PDB-7o1c: 
Cryo-EM structure of an Escherichia coli TnaC(R23F)-ribosome-RF2 complex stalled in response to L-tryptophan
Method: single particle / : van der Stel AX, Gordon ER, Sengupta A, Martinez AK, Klepacki D, Perry TN, Herrero del Valle A, Vazquez-Laslop N, Sachs MS, Cruz-Vera LR, Innis CA

EMDB-7800: 
Cryo-EM map of F.graminearum Mitochondrial Calcium Uniporter in lipid nanodisc - overall
Method: single particle / : Orlando BJ, Liao M

EMDB-7801: 
Cryo-EM map of F.graminearum Mitochondrial Calcium Uniporter in lipid nanodisc - type 1
Method: single particle / : Orlando BJ, Liao M

EMDB-7802: 
Cryo-EM map of F.graminearum Mitochondrial Calcium Uniporter in lipid nanodisc - type 2
Method: single particle / : Orlando BJ, Liao M

EMDB-7803: 
Cryo-EM map of F.graminearum Mitochondrial Calcium Uniporter in PMAL-C8
Method: single particle / : Orlando BJ, Liao M

EMDB-7804: 
Cryo-EM map of M.acridum Mitochondrial Calcium Uniporter in A8-35 amphipol
Method: single particle / : Orlando BJ, Liao M
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search




 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
