-Search query
-Search result
Showing 1 - 50 of 96 items for (author: pedersen & m)

EMDB-18350: 
S. cerevisia Niemann-Pick type C protein NCR1 in GDN at pH 7.5
Method: single particle / : Frain KM, Dedic E, Nel L, Olesen E, Stokes D, Panyella Pedersen B

EMDB-18351: 
S. cerevisia Niemann-Pick type C protein NCR1 in GDN at pH 5.5
Method: single particle / : Frain KM, Dedic E, Nel L, Olesen E, Stokes D, Panyella Pedersen B

EMDB-18352: 
S. cerevisia Niemann-Pick type C protein NCR1 in LMNG at pH 5.5
Method: single particle / : Frain KM, Nel L, Dedic E, Olesen E, Stokes D, Panyella Pedersen B

EMDB-18353: 
S. cerevisia Niemann-Pick type C protein NCR1 in Peptidisc at pH 7.5
Method: single particle / : Frain KM, Dedic E, Nel L, Olesen E, Stokes D, Panyella Pedersen B

PDB-8qeb: 
S. cerevisia Niemann-Pick type C protein NCR1 in GDN at pH 7.5
Method: single particle / : Frain KM, Dedic E, Nel L, Olesen E, Stokes D, Panyella Pedersen B

PDB-8qec: 
S. cerevisia Niemann-Pick type C protein NCR1 in GDN at pH 5.5
Method: single particle / : Frain KM, Dedic E, Nel L, Olesen E, Stokes D, Panyella Pedersen B

PDB-8qed: 
S. cerevisia Niemann-Pick type C protein NCR1 in LMNG at pH 5.5
Method: single particle / : Frain KM, Nel L, Dedic E, Olesen E, Stokes D, Panyella Pedersen B

PDB-8qee: 
S. cerevisia Niemann-Pick type C protein NCR1 in Peptidisc at pH 7.5
Method: single particle / : Frain KM, Dedic E, Nel L, Olesen E, Stokes D, Panyella Pedersen B

EMDB-28669: 
Single-Molecule 3D Image of 16 Helix RNA Origami Satellite by Individual Particle Electron Tomography (No. 01)
Method: electron tomography / : Liu J, Ren G
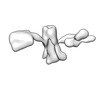
EMDB-28670: 
Single-Molecule 3D Image of 16 Helix RNA Origami Satellite by Individual Particle Electron Tomography (No. 02)
Method: electron tomography / : Liu J, Ren G
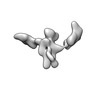
EMDB-28671: 
Single-Molecule 3D Image of 16 Helix RNA Origami Satellite by Individual Particle Electron Tomography (No. 03)
Method: electron tomography / : Liu J, Ren G
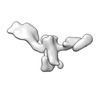
EMDB-28672: 
Single-Molecule 3D Image of 16 Helix RNA Origami Satellite by Individual Particle Electron Tomography (No. 04)
Method: electron tomography / : Liu J, Ren G

EMDB-28673: 
Single-Molecule 3D Image of 16 Helix RNA Origami Satellite by Individual Particle Electron Tomography (No. 05)
Method: electron tomography / : Liu J, Ren G
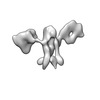
EMDB-28674: 
Single-Molecule 3D Image of 16 Helix RNA Origami Satellite by Individual Particle Electron Tomography (No. 06)
Method: electron tomography / : Liu J, Ren G
EMDB-28675: 
Single-Molecule 3D Image of 16 Helix RNA Origami Satellite by Individual Particle Electron Tomography (No. 07)
Method: electron tomography / : Liu J, Ren G
EMDB-28676: 
Single-Molecule 3D Image of 16 Helix RNA Origami Satellite by Individual Particle Electron Tomography (No. 08)
Method: electron tomography / : Liu J, Ren G

EMDB-28677: 
Single-Molecule 3D Image of 16 Helix RNA Origami Satellite by Individual Particle Electron Tomography (No. 09)
Method: electron tomography / : Liu J, Ren G

EMDB-28678: 
Single-Molecule 3D Image of 16 Helix RNA Origami Satellite by Individual Particle Electron Tomography (No. 10)
Method: electron tomography / : Liu J, Ren G

EMDB-28679: 
Single-Molecule 3D Image of 16 Helix RNA Origami Satellite by Individual Particle Electron Tomography (No. 11)
Method: electron tomography / : Liu J, Ren G

EMDB-28680: 
Single-Molecule 3D Image of 16 Helix RNA Origami Satellite by Individual Particle Electron Tomography (No. 12)
Method: electron tomography / : Liu J, Ren G

EMDB-28681: 
Single-Molecule 3D Image of 16 Helix RNA Origami Satellite by Individual Particle Electron Tomography (No. 13)
Method: electron tomography / : Liu J, Ren G

EMDB-28682: 
Single-Molecule 3D Image of 16 Helix RNA Origami Satellite by Individual Particle Electron Tomography (No. 14)
Method: electron tomography / : Liu J, Ren G
EMDB-28683: 
Single-Molecule 3D Image of 16 Helix RNA Origami Satellite by Individual Particle Electron Tomography (No. 15)
Method: electron tomography / : Liu J, Ren G
EMDB-28684: 
Single-Molecule 3D Image of 16 Helix RNA Origami Satellite by Individual Particle Electron Tomography (No. 16)
Method: electron tomography / : Liu J, Ren G

EMDB-13926: 
Twist-corrected RNA origami 5-helix Tile A
Method: single particle / : McRae EKS, Bogglid A, Boesen T, Andersen ES

PDB-7qdu: 
Twist-corrected RNA origami 5-helix Tile A
Method: single particle / : McRae EKS, Andersen ES

EMDB-13625: 
Mature conformer 2 of a 6-helix bundle of RNA with a clasp
Method: single particle / : McRae EKS, Bogglid A, Boesen T, Andersen ES

EMDB-13626: 
Young conformer 2 of a 6-helix bundle of RNA with a clasp
Method: single particle / : McRae EKS, Bogglid A, Boesen T, Andersen ES

EMDB-13627: 
6-Helix bundle of RNA
Method: single particle / : McRae EKS, Bogglid A, Boesen T, Andersen ES

EMDB-13628: 
Young conformer of a 6-helix bundle of RNA with clasp
Method: single particle / : McRae EKS, Bogglid A, Boesen T, Andersen ES

EMDB-13630: 
Mature conformer of a 6-helix bundle of RNA with clasp
Method: single particle / : McRae EKS, Bogglid A, Boesen T, Andersen ES

EMDB-13633: 
RNA origami 5-helix tile
Method: single particle / : McRae EKS, Bogglid A, Boesen T, Andersen ES

EMDB-13636: 
RNA origami 5-helix tile
Method: single particle / : McRae EKS, Bogglid A, Boesen T, Andersen ES

PDB-7ptk: 
Young conformer of a 6-helix bundle of RNA with clasp
Method: single particle / : McRae EKS, Andersen ES

PDB-7ptl: 
Mature conformer of a 6-helix bundle of RNA with clasp
Method: single particle / : McRae EKS, Andersen ES

PDB-7ptq: 
RNA origami 5-helix tile
Method: single particle / : McRae EKS, Andersen ES

PDB-7pts: 
RNA origami 5-helix tile
Method: single particle / : McRae EKS, Andersen ES

EMDB-13592: 
Cryo-EM SPA reconstruction of an extended RNA origami 5 helix tile (5HT-B-3X)
Method: single particle / : McRae EKS, Bogglid A, Boesen T, Andersen ES

EMDB-14115: 
Outward-facing apo-form of auxin transporter PIN8
Method: single particle / : Ung KL, Winkler MBL, Dedic E, Stokes DL, Pedersen BP

EMDB-14116: 
Outward-facing auxin bound form of auxin transporter PIN8
Method: single particle / : Ung KL, Winkler MBL, Dedic E, Stokes DL, Pedersen BP

EMDB-14117: 
Inward-facing NPA bound form of auxin transporter PIN8
Method: single particle / : Ung KL, Winkler MBL, Dedic E, Stokes DL, Pedersen BP

EMDB-14118: 
Outward-facing apo-form of auxin transporter PIN8 in detergent
Method: single particle / : Ung KL, Winkler MBL, Dedic E, Stokes DL, Pedersen BP

PDB-7qp9: 
Outward-facing apo-form of auxin transporter PIN8
Method: single particle / : Ung KL, Winkler MBL, Dedic E, Stokes DL, Pedersen BP

PDB-7qpa: 
Outward-facing auxin bound form of auxin transporter PIN8
Method: single particle / : Ung KL, Winkler MBL, Dedic E, Stokes DL, Pedersen BP

PDB-7qpc: 
Inward-facing NPA bound form of auxin transporter PIN8
Method: single particle / : Ung KL, Winkler MBL, Dedic E, Stokes DL, Pedersen BP

EMDB-13326: 
T. maritima CorA in DDM micelles without Mg2+ bound in D2O
Method: single particle / : Larsen AH, Johansen NT, Arleth L

EMDB-13327: 
T. maritima CorA in DDM micelles with Mg2+ bound in D2O
Method: single particle / : Larsen AH, Johansen NT, Arleth L

EMDB-12465: 
SARS-CoV-2 Spike RBD (dimer) in complex with two Fu2 nanobodies
Method: single particle / : Das H, Hallberg BM

EMDB-12561: 
SARS-CoV-2 Spike (dimers) in complex with six Fu2 nanobodies
Method: single particle / : Das H, Hallberg BM

PDB-7nll: 
SARS-CoV-2 Spike RBD (dimer) in complex with two Fu2 nanobodies
Method: single particle / : Das H, Hallberg BM
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
