-Search query
-Search result
Showing 1 - 50 of 331 items for (author: ozorowski & g)

EMDB-42464: 
chEnv TTT protein in complex with 43A2 Fab
Method: single particle / : Ozorowski G, Lee WH, Ward AB

EMDB-42468: 
chEnv TTT protein in complex with CM01A Fab
Method: single particle / : Ozorowski G, Lee WH, Ward AB

EMDB-43664: 
Triple tandem trimer immunogens for HIV-1 and influenza nucleic acid-based vaccines
Method: single particle / : Ferguson JA, Leon AN, Ward AB

EMDB-43665: 
Triple tandem trimer immunogens for HIV-1 and influenza nucleic acid-based vaccines (cH125 TTT)
Method: single particle / : Ferguson JA, Leon AN, Ward AB

EMDB-43666: 
Triple tandem trimer immunogens for HIV-1 and influenza nucleic acid-based vaccines (H2/1 GCN4)
Method: single particle / : Ferguson JA, Leon AN, Ward AB

EMDB-43668: 
Triple tandem trimer immunogens for HIV-1 and influenza nucleic acid-based vaccines (H5/1 GCN4)
Method: single particle / : Ferguson JA, Leon AN, Ward AB

EMDB-43669: 
Triple tandem trimer immunogens for HIV-1 and influenza nucleic acid-based vaccines. H5 GCN4
Method: single particle / : Ferguson JA, Leon AN, Ward AB
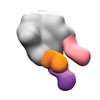
EMDB-40242: 
BG505 MD39 SOSIP in complex with Rh.NJ85 wk12 gp120GH, N611/FP, and base epitope polyclonal antibodies
Method: single particle / : Lee WH, Ozorowski G, Torres JL, Ward AB

EMDB-40243: 
BG505 MD39 SOSIP in complex with Rh.NJ86 wk12 V1V3, C3V5, N611/FP, and base epitope polyclonal antibodies
Method: single particle / : Lee WH, Ozorowski G, Torres JL, Ward AB

EMDB-40244: 
BG505 MD39 SOSIP in complex with Rh.NJ76 wk12 C3V5, N611/FP, and base epitope polyclonal antibodies
Method: single particle / : Lee WH, Ozorowski G, Torres JL, Ward AB
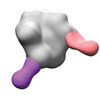
EMDB-40252: 
BG505 MD39 SOSIP in complex with Rh.NK04 wk12 gp120 glycan hole and base epitope polyclonal antibodies
Method: single particle / : Lee WS, Ozorowski G, Torres JL, Ward AB

EMDB-40254: 
BG505 MD39 SOSIP in complex with Rh.NJ75 wk12 gp120 glycan hole and base epitope polyclonal antibodies
Method: single particle / : Lee WS, Ozorowski G, Torres JL, Ward AB

EMDB-40255: 
BG505 MD39 SOSIP in complex with Rh.NJ87 wk12 C3V5 and base epitope polyclonal antibodies
Method: single particle / : Lee WS, Ozorowski G, Torres JL, Ward AB

EMDB-40256: 
BG505 MD39 SOSIP in complex with Rh.NJ84 wk12 V1V3, gp120 glycan hole and base epitope polyclonal antibodies
Method: single particle / : Lee WS, Ozorowski G, Torres JL, Ward AB

EMDB-40257: 
BG505 MD39 SOSIP in complex with Rh.NJ77 wk12 V1V3, C3V5, N611/FP and base epitope polyclonal
Method: single particle / : Lee WS, Ozorowski G, Torres JL, Ward AB

EMDB-28850: 
SARS-CoV-2 Gamma 6P Mut7 S + COVA309-3 Fab
Method: single particle / : Torres JL, Ward AB

EMDB-28851: 
SARS-CoV-2 Gamma 6P Mut7 S + COVA309-10 Fab
Method: single particle / : Torres JL, Ward AB

EMDB-28852: 
SARS-CoV-2 Omicron 6P S + COVA309-35 Fab
Method: single particle / : Torres JL, Ward AB
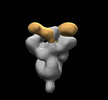
EMDB-28853: 
SARS-CoV-2 Gamma 6P Mut7 + S COVA309-38 Fab
Method: single particle / : Torres JL, Ward AB

EMDB-40088: 
HIV-1 Env subtype C CZA97.12 SOSIP.664 in complex with 3BNC117 Fab
Method: single particle / : Ozorowski G, Lee JH, Ward AB

PDB-8gje: 
HIV-1 Env subtype C CZA97.12 SOSIP.664 in complex with 3BNC117 Fab
Method: single particle / : Ozorowski G, Lee JH, Ward AB

EMDB-27920: 
3H03 Fab in complex with influenza virus neuraminidase from A/Brevig Mission/1/1918 (H1N1)
Method: single particle / : Turner HL, Ozorowski G, Ward AB

EMDB-27921: 
2H08 Fab in complex with influenza virus neuraminidase from A/Brevig Mission/1/1918 (H1N1)
Method: single particle / : Turner HL, Ozorowski G, Ward AB

PDB-8e6j: 
3H03 Fab in complex with influenza virus neuraminidase from A/Brevig Mission/1/1918 (H1N1)
Method: single particle / : Turner HL, Ozorowski G, Ward AB

PDB-8e6k: 
2H08 Fab in complex with influenza virus neuraminidase from A/Brevig Mission/1/1918 (H1N1)
Method: single particle / : Turner HL, Ozorowski G, Ward AB

EMDB-40803: 
BG505 Boost2 SOSIP.664 in complex with NHP polyclonal antibody FP2
Method: single particle / : Pratap PP, Antansijevic A, Ward AB

EMDB-40804: 
BG505 Boost2 SOSIP.664 in complex with NHP Polyclonal Antibody FP4
Method: single particle / : Pratap PP, Antansijevic A, Ward AB

EMDB-40805: 
BG505 Boost 2 in complex with NHP antibody V1V2
Method: single particle / : Pratap PP, Antansijevic A, Ward AB

EMDB-40806: 
BG505 Boost 2 in complex with NHP Polyclonal Antibody N625
Method: single particle / : Pratap PP, Antansijevic A, Ward AB

EMDB-40807: 
BG505 Boost2 SOSIP.664 in complex with NHP polyclonal antibody Base1
Method: single particle / : Pratap PP, Antansijevic A, Ward AB

EMDB-40808: 
Boost 2 SOSIP.664 in complex with NHP polyclonal antibody Base2
Method: single particle / : Pratap PP, Antansijevic A, Ward AB

EMDB-40809: 
BG505 Boost2 SOSIP.664 in complex with NHP polyclonal antibody Base3
Method: single particle / : Pratap PP, Antansijevic A, Ward AB

EMDB-40810: 
BG505 Boost2 SOSIP.664 in complex with NHP polyclonal antibody FP1
Method: single particle / : Pratap PP, Antansijevic A, Ozorowski G, Ward AB

PDB-8sw7: 
BG505 Boost2 SOSIP.664 in complex with NHP polyclonal antibody FP1
Method: single particle / : Pratap PP, Antansijevic A, Ozorowski G, Ward AB

EMDB-27692: 
LM18/Nb136 bispecific tetra-nanobody immunoglobulin in complex with SARS-CoV-2-6P-Mut7 S protein (focused refinement)
Method: single particle / : Ozorowski G, Turner HL, Ward AB

EMDB-27693: 
LM18/Nb136 bispecific tetra-nanobody immunoglobulin in complex with SARS-CoV-2-6P-Mut7 S protein (global refinement)
Method: single particle / : Ozorowski G, Turner HL, Ward AB

PDB-8dt8: 
LM18/Nb136 bispecific tetra-nanobody immunoglobulin in complex with SARS-CoV-2-6P-Mut7 S protein (focused refinement)
Method: single particle / : Ozorowski G, Turner HL, Ward AB

EMDB-27112: 
S728-1157 IgG in complex with SARS-CoV-2-6P-Mut7 Spike protein (global refinement)
Method: single particle / : Ozorowski G, Torres JL, Turner HL, Ward AB

EMDB-27113: 
S728-1157 IgG in complex with SARS-CoV-2-6P-Mut7 Spike protein (focused refinement)
Method: single particle / : Ozorowski G, Torres JL, Ward AB

PDB-8d0z: 
S728-1157 IgG in complex with SARS-CoV-2-6P-Mut7 Spike protein (focused refinement)
Method: single particle / : Ozorowski G, Torres JL, Ward AB

EMDB-14888: 
CM01B in complex with ConM SOSIP
Method: single particle / : van Schooten J, Ward AB

EMDB-14889: 
CM05A1 in complex with conM SOSIP
Method: single particle / : van Schooten J, Ward AB

EMDB-14890: 
CM02A in complex with conM SOSIP
Method: single particle / : van Schooten J, Ward AB

EMDB-14891: 
CM04F in complex with conM SOSIP
Method: single particle / : van Schooten J, Ward AB

EMDB-14892: 
CM03B in complex with conM SOSIP
Method: single particle / : van Schooten J, Ward AB

EMDB-14893: 
CM05E in complex with conM SOSIP
Method: single particle / : van Schooten J, Ward AB

EMDB-14894: 
CM05L in complex with conM SOSIP
Method: single particle / : van Schooten J, Ward AB

EMDB-14895: 
NHP MCM02 wk26 N355/N289 pAb in complex with ConM SOSIP
Method: single particle / : van Schooten J, Ward AB

EMDB-14896: 
NHP MCM04 wk26 base pAb in complex with ConM SOSIP
Method: single particle / : van Schooten J, Ward AB

EMDB-14897: 
NHP MCM04 wk26 N611 pAb in complex with ConM SOSIP
Method: single particle / : van Schooten J, Ward AB
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
