-Search query
-Search result
Showing all 22 items for (author: oliveira & tm)

EMDB-16145: 
Cryo-EM structure of NHEJ supercomplex(trimer)
Method: single particle / : Hardwick SW, Chaplin AK

EMDB-14986: 
CryoEM structure of Ku heterodimer bound to DNA
Method: single particle / : Hardwick SW, Kefala-Stavridi A, Chirgadze DY, Blundell TL, Chaplin AK

EMDB-14955: 
Cryo-EM structure of Ku 70/80 bound to inositol hexakisphosphate
Method: single particle / : Kefala Stavridi A, Chaplin AK, Blundell TL

EMDB-15987: 
In situ sub-tomogram average of the Ca. L. ossiferum ribosome
Method: subtomogram averaging / : Rodrigues-Oliveira T, Wollweber F, Ponce-Toledo R, Xu J, Rittmann S, Klingl A, Pilhofer M, Schleper C

EMDB-15988: 
In situ sub-tomogram average of Ca. L. ossiferum actin filament
Method: subtomogram averaging / : Rodrigues-Oliveira T, Wollweber F, Ponce-Toledo R, Xu J, Rittmann S, Klingl A, Pilhofer M, Schleper C

EMDB-15989: 
cryo-tomogram of a Ca. L. ossiferum cell
Method: electron tomography / : Rodrigues-Oliveira T, Wollweber F, Ponce-Toledo R, Xu J, Rittmann S, Klingl A, Pilhofer M, Schleper C

EMDB-15990: 
cryo-tomogram of Ca. L. ossiferum protrusions
Method: electron tomography / : Rodrigues-Oliveira T, Wollweber F, Ponce-Toledo R, Xu J, Rittmann S, Klingl A, Pilhofer M, Schleper C

EMDB-15991: 
cryo-tomogram of a Ca.L. ossiferum cell
Method: electron tomography / : Rodrigues-Oliveira T, Wollweber F, Ponce-Toledo R, Xu J, Rittmann S, Klingl A, Pilhofer M, Schleper C

EMDB-15992: 
cryo-tomogram of a Ca. L. ossiferum cell
Method: electron tomography / : Rodrigues-Oliveira T, Wollweber F, Ponce-Toledo R, Xu J, Rittmann S, Klingl A, Pilhofer M, Schleper C

EMDB-15993: 
cryo-tomogram of a Ca. L. ossiferum cell
Method: electron tomography / : Rodrigues-Oliveira T, Wollweber F, Ponce-Toledo R, Xu J, Rittmann S, Klingl A, Pilhofer M, Schleper C

EMDB-12299: 
Cryo-EM structure of NHEJ super-complex (dimer)
Method: single particle / : Chaplin AK, Hardwick SW

EMDB-12301: 
Cryo-EM structure of NHEJ super-complex (monomer)
Method: single particle / : Chaplin AK, Hardwick SW

EMDB-11185: 
Cryo-EM structure of DNA-PKcs (State 2)
Method: single particle / : Chaplin AK, Hardwick SW

EMDB-11211: 
Cryo-EM structure of DNA-PKcs (State 1)
Method: single particle / : Chaplin AK, Hardwick SW

EMDB-11213: 
Cryo-EM structure of DNA-PKcs (State 3)
Method: single particle / : Chaplin AK, Hardwick SW

EMDB-11215: 
Cryo-EM structure of DNA-PKcs:Ku80ct194
Method: single particle / : Chaplin AK, Hardwick SW

EMDB-11216: 
Cryo-EM structure of DNA-PKcs:DNA
Method: single particle / : Chaplin AK, Hardwick SW

EMDB-11217: 
Cryo-EM structure of DNA-PK monomer
Method: single particle / : Chaplin AK, Hardwick SW

EMDB-11219: 
Cryo-EM structure of DNA-PK dimer
Method: single particle / : Chaplin AK, Hardwick SW

EMDB-10288: 
structural insights and activating mutations in diverse pathologies define mechanisms of deregulation for phospholipase C gamma enzymes
Method: single particle / : Liu Y, Bunney T, Phillips C, Katan M

EMDB-3896: 
Cryo-EM Structure of Saccharomyces cerevisiae Target of Rapamycin Complex 2
Method: single particle / : Karuppasamy M, Kusmider B
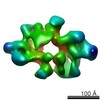
EMDB-2990: 
Structure of Target Of Rapapmycin Complex 2 (TORC2) from Saccharomyces cerevisiae
Method: single particle / : Gaubitz C, Oliveira TM, Prouteau M, Leitner A, Karuppasamy M, Konstantinidou G, Rispal D, Eltschinger S, Robinson GC, Thore S, Aebersold R, Schaffitzel C, Loewith R
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
