-Search query
-Search result
Showing all 25 items for (author: obarska-kosinska & a)
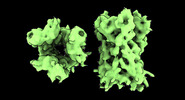
EMDB-17704: 
Subtomogram average of Vaccinia A10 trimer with open center from in vitro cores
Method: subtomogram averaging / : Turonova B, Liu J
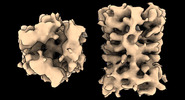
EMDB-17708: 
Subtomogram average of Vaccinia A10 trimer with tight center from in vitro cores
Method: subtomogram averaging / : Turonova B, Liu J

EMDB-17753: 
Subtomogram average of Vaccinia A10 trimer from in situ cores
Method: subtomogram averaging / : Turonova B, Liu J

EMDB-14325: 
Cytoplasmic ring of the human nuclear pore complex from isolated HeLa nuclear envelopes
Method: subtomogram averaging / : Mosalaganti S, Obarska-Kosinska A, Siggel M, Taniguchi R, Turonova B, Zimmerli CE, Buczak K, Schmidt FH, Margiotta E, Mackmull MT, Hagen WJH, Hummer G, Kosinski J, Beck M

EMDB-14326: 
Structure of nuclear pore complex from HEK cells with GP210 knockout
Method: subtomogram averaging / : Mosalaganti S, Obarska-Kosinska A, Siggel M, Taniguchi R, Turonova B, Zimmerli CE, Buczak K, Schmidt FH, Margiotta E, Mackmull MT, Hagen WJH, Hummer G, Kosinski J, Beck M

EMDB-14327: 
Nuclear pore complex from intact HeLa cells
Method: subtomogram averaging / : Mosalaganti S, Obarska-Kosinska A, Siggel M, Taniguchi R, Turonova B, Zimmerli CE, Buczak K, Schmidt FH, Margiotta E, Mackmull MT, Hagen WJH, Hummer G, Kosinski J, Beck M

EMDB-14328: 
Inner/spoke ring of the human nuclear pore complex from isolated HeLa nuclear envelopes.
Method: subtomogram averaging / : Mosalaganti S, Obarska-Kosinska A, Siggel M, Taniguchi R, Turonova B, Zimmerli CE, Buczak K, Schmidt FH, Margiotta E, Mackmull MT, Hagen WJH, Hummer G, Kosinski J, Beck M

EMDB-14330: 
Nuclear ring of the human nuclear pore complex from isolated HeLa nuclear envelopes
Method: subtomogram averaging / : Mosalaganti S, Obarska-Kosinska A, Siggel M, Taniguchi R, Turonova B, Zimmerli CE, Buczak K, Schmidt FH, Margiotta E, Mackmull MT, Hagen WJH, Hummer G, Kosinski J, Beck M

EMDB-14321: 
Human nuclear pore complex (dilated)
Method: subtomogram averaging / : Mosalaganti S, Obarska-Kosinska A, Siggel M, Taniguchi R, Turonova B, Zimmerli CE, Buczak K, Schmidt FH, Margiotta E, Mackmull MT, Hagen WJH, Hummer G, Kosinski J, Beck M

PDB-7r5j: 
Human nuclear pore complex (dilated)
Method: subtomogram averaging / : Mosalaganti S, Obarska-Kosinska A, Siggel M, Taniguchi R, Turonova B, Zimmerli CE, Buczak K, Schmidt FH, Margiotta E, Mackmull MT, Hagen WJH, Hummer G, Kosinski J, Beck M

EMDB-14322: 
Human nuclear pore complex (constricted)
Method: subtomogram averaging / : Mosalaganti S, Obarska-Kosinska A, Siggel M, Taniguchi R, Turonova B, Zimmerli CE, Buczak K, Schmidt FH, Margiotta E, Mackmull MT, Hagen WJH, Hummer G, Kosinski J, Beck M

PDB-7r5k: 
Human nuclear pore complex (constricted)
Method: subtomogram averaging / : Mosalaganti S, Obarska-Kosinska A, Siggel M, Taniguchi R, Turonova B, Zimmerli CE, Buczak K, Schmidt FH, Margiotta E, Mackmull MT, Hagen WJH, Hummer G, Kosinski J, Beck M

EMDB-14243: 
cryoEM structure of human Nup155 (residues 19-981)
Method: single particle / : Taniguchi R, Beck M

PDB-7r1y: 
cryoEM structure of human Nup155 (residues 19-981)
Method: single particle / : Taniguchi R, Beck M

EMDB-13081: 
In cellulo nuclear pore complex from S. pombe cells exposed to a hypertonic osmotic shock
Method: subtomogram averaging / : Zimmerli CE, Allegretti M, Rantos V, Goetz S, Obarska-Kosinska A, Zagoriy I, Mahamid J, Kosinski J, Beck M

EMDB-11373: 
In cellulo nuclear pore complex from S. pombe cells
Method: subtomogram averaging / : Zimmerli CE, Allegretti M, Rantos V, Goetz S, Obarska-Kosinska A, Zagoriy I, Hummer G, Mahamid J, Kosinski J, Beck M

EMDB-11374: 
In cellulo nuclear pore complex with intermediate diameter from energy depleted S. pombe cells
Method: subtomogram averaging / : Zimmerli CE, Allegretti M, Rantos V, Goetz S, Obarska-Kosinska A, Zagoriy I, Hummer G, Mahamid J, Kosinski J, Beck M

EMDB-11375: 
In cellulo nuclear pore complex in constricted conformation from energy depleted S. pombe cells
Method: subtomogram averaging / : Zimmerli CE, Allegretti M, Rantos V, Goetz S, Obarska-Kosinska A, Zagoriy I, Hummer G, Mahamid J, Kosinski J, Beck M
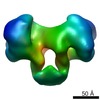
EMDB-1890: 
EcoR124 Type I DNA restriction-modification enzyme complex in closed state with bound 30bp cognate DNA fragment. 3D reconstruction by single particle analysis from negative stain EM.
Method: single particle / : Kennaway CK, Taylor JE, Song CF, Potrzebowski W, White JH, Swiderska A, Obarska-Kosinska A, Callow P, Cooper LP, Roberts GA, Bujnicki JM, Trinick J, Kneale GG, Dryden DTF

EMDB-1891: 
EcoR124 Type I DNA restriction-modification enzyme complex without DNA (open state). Low resolution 3D reconstruction by single particle analysis from negative stain EM.
Method: single particle / : Kennaway CK, Taylor JE, Song CF, Potrzebowski W, White JH, Swiderska A, Obarska-Kosinska A, Callow P, Cooper LP, Roberts GA, Bujnicki JM, Trinick J, Kneale GG, Dryden DTF
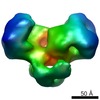
EMDB-1892: 
EcoR124 Type I DNA restriction-modification enzyme complex (in closed state) with bound DNA mimic protein Ocr from phage T7. 3D reconstruction by single particle analysis from negative stain EM.
Method: single particle / : Kennaway CK, Taylor JE, Song CF, Potrzebowski W, White JH, Swiderska A, Obarska-Kosinska A, Callow P, Cooper LP, Roberts GA, Bujnicki JM, Trinick J, Kneale GG, Dryden DTF

EMDB-1893: 
EcoKI Type I DNA restriction-modification enzyme complex in closed state with bound 75bp cognate DNA fragment. 3D reconstruction by single particle analysis from negative stain EM.
Method: single particle / : Kennaway CK, Taylor JE, Song CF, Potrzebowski W, White JH, Swiderska A, Obarska-Kosinska A, Callow P, Cooper LP, Roberts GA, Bujnicki JM, Trinick J, Kneale GG, Dryden DTF

PDB-2y7c: 
Atomic model of the Ocr-bound methylase complex from the Type I restriction-modification enzyme EcoKI (M2S1). Based on fitting into EM map 1534.
Method: single particle / : Kennaway CK, Obarska-Kosinska A, White JH, Tuszynska I, Cooper LP, Bujnicki JM, Trinick J, Dryden DTF

PDB-2y7h: 
Atomic model of the DNA-bound methylase complex from the Type I restriction-modification enzyme EcoKI (M2S1). Based on fitting into EM map 1534.
Method: single particle / : Kennaway CK, Obarska-Kosinska A, White JH, Tuszynska I, Cooper LP, Bujnicki JM, Trinick J, Dryden DTF

EMDB-1534: 
EcoKI type I RM methyltransferase with DNA mimic Ocr. Negative stain 3D.
Method: single particle / : Kennaway CK, Obarska-Kosinska A, White JH, Tuszynska I, Cooper LP, Bujnicki JM, Trinick J, Dryden DTF
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
