-Search query
-Search result
Showing 1 - 50 of 4,034 items for (author: n & cheng)

EMDB-38845: 
Icosahedrally averaged cryo-EM reconstruction of PhiKZ capsid before applying the "block-based" reconstruction method
Method: single particle / : Yang Y, Shao Q, Guo M, Han L, Zhao X, Wang A, Li X, Wang B, Pan J, Chen Z, Fokine A, Sun L, Fang Q

EMDB-38846: 
Block 1 of PhiKZ capsid
Method: single particle / : Yang Y, Shao Q, Guo M, Han L, Zhao X, Wang A, Li X, Wang B, Pan J, Chen Z, Fokine A, Sun L, Fang Q

EMDB-38848: 
Block 2 of PhiKZ capsid
Method: single particle / : Yang Y, Shao Q, Guo M, Han L, Zhao X, Wang A, Li X, Wang B, Pan J, Chen Z, Fokine A, Sun L, Fang Q

EMDB-39002: 
Composite cryo-EM map of PhiKZ capsid after applying the "block-based" reconstruction method
Method: single particle / : Yang Y, Shao Q, Guo M, Han L, Zhao X, Wang A, Li X, Wang B, Pan J, Chen Z, Fokine A, Sun L, Fang Q

PDB-8y6v: 
Near-atomic structure of icosahedrally averaged jumbo bacteriophage PhiKZ capsid
Method: single particle / : Yang Y, Shao Q, Guo M, Han L, Zhao X, Wang A, Li X, Wang B, Pan J, Chen Z, Fokine A, Sun L, Fang Q
EMDB-41485: 
Subtomogram averaged consensus structure of the malarial 80S ribosome in Plasmodium falciparum-infected human erythrocytes
Method: subtomogram averaging / : Anton L, Cheng W, Zhu X, Ho CM
EMDB-41486: 
Subtomogram averaged decoding-1 state of the malarial 80S ribosome in Plasmodium falciparum-infected human erythrocytes
Method: subtomogram averaging / : Anton L, Cheng W, Zhu X, Ho CM

EMDB-41487: 
Subtomogram averaged decoding-2 state of the malarial 80S ribosome in Plasmodium falciparum-infected human erythrocytes
Method: subtomogram averaging / : Anton L, Cheng W, Zhu X, Ho CM

EMDB-41488: 
Subtomogram averaged classical iPRE state of the malarial 80S ribosome in Plasmodium falciparum-infected human erythrocytes
Method: subtomogram averaging / : Anton L, Cheng W, Zhu X, Ho CM
EMDB-41489: 
Subtomogram averaged rotated-1 PRE state of the malarial 80S ribosome in Plasmodium falciparum-infected human erythrocytes
Method: subtomogram averaging / : Anton L, Cheng W, Zhu X, Ho CM

EMDB-41490: 
Subtomogram averaged rotated-2 PRE state of the malarial 80S ribosome in Plasmodium falciparum-infected human erythrocytes
Method: subtomogram averaging / : Anton L, Cheng W, Zhu X, Ho CM

EMDB-41491: 
Subtomogram averaged translocation state of the malarial 80S ribosome in Plasmodium falciparum-infected human erythrocytes
Method: subtomogram averaging / : Anton L, Cheng W, Zhu X, Ho CM
EMDB-41492: 
Subtomogram averaged POST state of the malarial 80S ribosome in Plasmodium falciparum-infected human erythrocytes
Method: subtomogram averaging / : Anton L, Cheng W, Zhu X, Ho CM

EMDB-41493: 
Subtomogram averaged unloaded state of the malarial 80S ribosome in Plasmodium falciparum-infected human erythrocytes
Method: subtomogram averaging / : Anton L, Cheng W, Zhu X, Ho CM
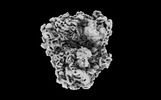
EMDB-41494: 
Subtomogram averaged non-rotated 80S ribosome of the malarial 80S ribosome in Plasmodium falciparum-infected human erythrocytes
Method: subtomogram averaging / : Anton L, Cheng W, Zhu X, Ho CM

PDB-8tpu: 
Subtomogram averaged consensus structure of the malarial 80S ribosome in Plasmodium falciparum-infected human erythrocytes
Method: subtomogram averaging / : Anton L, Cheng W, Zhu X, Ho CM

EMDB-37441: 
FCP tetramer in Chaetoceros gracilis
Method: single particle / : Feng Y, Li Z, Zhou C, Shen JR, Liu C, Wang W

EMDB-37442: 
FCP pentamer in Chaetoceros gracilis
Method: single particle / : Feng Y, Li Z, Zhou C, Liu C, Shen JR, Wang W

PDB-8wck: 
FCP tetramer in Chaetoceros gracilis
Method: single particle / : Feng Y, Li Z, Zhou C, Shen JR, Liu C, Wang W

PDB-8wcl: 
FCP pentamer in Chaetoceros gracilis
Method: single particle / : Feng Y, Li Z, Zhou C, Liu C, Shen JR, Wang W

EMDB-42400: 
RORC mRNA 3'UTR riboswitch A97G/G98A mutant class C
Method: single particle / : Asarnow D, Khoroshkin M, Goodarzi H, Cheng Y

EMDB-42401: 
RORC mRNA 3'UTR riboswitch 77-GA mutant class A
Method: single particle / : Asarnow D, Khoroshkin M, Goodarzi H, Cheng Y

EMDB-42403: 
RORC mRNA 3'UTR riboswitch 117-AC mutant class C
Method: single particle / : Asarnow D, Khoroshkin M, Goodarzi H, Cheng Y

EMDB-42404: 
RORC mRNA 3'UTR riboswitch 117-AC mutant class B
Method: single particle / : Asarnow D, Khoroshkin M, Goodarzi H, Cheng Y

EMDB-32979: 
Cryo-EM structure of Coxsackievirus B1 A-particle in complex with nAb 8A10 (CVB1-A:8A10)
Method: single particle / : Zheng Q, Zhu R, Sun H, Cheng T, Li S, Xia N

PDB-7x35: 
Cryo-EM structure of Coxsackievirus B1 A-particle in complex with nAb 8A10 (CVB1-A:8A10)
Method: single particle / : Zheng Q, Zhu R, Sun H, Cheng T, Li S, Xia N

EMDB-38617: 
SARS-CoV-2 RBD + IMCAS-123 + IMCAS-72 Fab
Method: single particle / : Tong Z, Cui Y, Xie Y, Tong J, Gao GF, Qi J

EMDB-38618: 
SARS-CoV-2 RBD + IMCAS-364 + hACE2
Method: single particle / : Tong Z, Cui Y, Xie Y, Tong J, Gao GF, Qi J

EMDB-38619: 
SARS-CoV-2 RBD + IMCAS-364 (Local Refinement)
Method: single particle / : Tong Z, Cui Y, Xie Y, Tong J, Gao GF, Qi J

EMDB-38620: 
SARS-CoV-2 Omicron BA.4 RBD + IMCAS-316 + ACE2
Method: single particle / : Tong Z, Cui Y, Xie Y, Tong J, Gao GF, Qi J

EMDB-38621: 
SARS-CoV-2 spike + IMCAS-123
Method: single particle / : Tong Z, Cui Y, Xie Y, Tong J, Gao GF, Qi J

EMDB-38823: 
Cryo-EM structure of the 123-316 scDb/PT-RBD complex
Method: single particle / : Jia GW, Tong Z, Tong JY, Su ZM

PDB-8xse: 
SARS-CoV-2 RBD + IMCAS-123 + IMCAS-72 Fab
Method: single particle / : Tong Z, Cui Y, Xie Y, Tong J, Gao GF, Qi J

PDB-8xsf: 
SARS-CoV-2 RBD + IMCAS-364 + hACE2
Method: single particle / : Tong Z, Cui Y, Xie Y, Tong J, Gao GF, Qi J

PDB-8xsi: 
SARS-CoV-2 RBD + IMCAS-364 (Local Refinement)
Method: single particle / : Tong Z, Cui Y, Xie Y, Tong J, Gao GF, Qi J

PDB-8xsj: 
SARS-CoV-2 Omicron BA.4 RBD + IMCAS-316 + ACE2
Method: single particle / : Tong Z, Cui Y, Xie Y, Tong J, Gao GF, Qi J

PDB-8xsl: 
SARS-CoV-2 spike + IMCAS-123
Method: single particle / : Tong Z, Cui Y, Xie Y, Tong J, Gao GF, Qi J

PDB-8y0y: 
Cryo-EM structure of the 123-316 scDb/PT-RBD complex
Method: single particle / : Jia GW, Tong Z, Tong JY, Su ZM

EMDB-36756: 
CryoEM structure of the transketolase ANIP from Streptomyces hygrospinosus
Method: single particle / : Jiang WX, Ma LX, Xing Q

EMDB-36836: 
Cryo-EM structure of the human 55S mitoribosome with Tigecycline
Method: single particle / : Li X, Wang M, Cheng J

EMDB-36837: 
Cryo-EM structure of the human 39S mitoribosome with Tigecycline
Method: single particle / : Li X, Wang M, Cheng J

EMDB-36838: 
Cryo-EM structure of the human 80S ribosome with Tigecycline
Method: single particle / : Li X, Wang M, Cheng J

EMDB-36839: 
Cryo-EM structure of the yeast 80S ribosome with tigecycline, eEF2, Stm1 and eIF5A
Method: single particle / : Buschauer R, Beckmann R, Cheng J

EMDB-36945: 
Cryo-EM structure of the yeast 80S ribosome with tigecycline, Not5 and P-site tRNA
Method: single particle / : Buschauer R, Beckmann R, Cheng J

EMDB-38629: 
Cryo-EM structure of the human 80S ribosome with Tigecycline, E-tRNA, SERBP1 and eEF2
Method: single particle / : Li X, Wang M, Cheng J

EMDB-38630: 
Cryo-EM structure of the human 80S ribosome with Tigecycline, e-tRNA and CCDC124 (40S head Swivelled)
Method: single particle / : Li X, Wang M, Cheng J

EMDB-38631: 
Cryo-EM structure of the human 80S ribosome with Tigecycline, E-tRNA and P-tRNA
Method: single particle / : Li X, Wang M, Cheng J

EMDB-38632: 
Cryo-EM structure of the human 55S mitoribosome with 5um Tigecycline
Method: single particle / : Li X, Wang M, Cheng J

EMDB-38633: 
Cryo-EM structure of the human 39S mitoribosome with 5uM Tigecycline
Method: single particle / : Li X, Wang M, Cheng J

EMDB-38634: 
Cryo-EM structure of the human 55S mitoribosome with 10uM Tigecycline
Method: single particle / : Li X, Wang M, Cheng J
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
