-Search query
-Search result
Showing all 50 items for (author: moreno & m)

EMDB-18520: 
Conserved Structures and Dynamics in 5-Proximal Regions of Betacoronavirus RNA Genomes
Method: single particle / : Moura TR, Purta E, Bernat A, Baulin E, Mukherjee S, Bujnicki JM

EMDB-18521: 
Conserved Structures and Dynamics in 5-Proximal Regions of Betacoronavirus RNA Genomes
Method: single particle / : Moura TR, Purta E, Bernat A, Baulin E, Mukherjee S, Bujnicki JM

EMDB-18522: 
Conserved Structures and Dynamics in 5-Proximal Regions of Betacoronavirus RNA Genomes
Method: single particle / : Moura TR, Purta E, Bernat A, Baulin E, Mukherjee S, Bujnicki JM

EMDB-18523: 
Conserved Structures and Dynamics in 5-Proximal Regions of Betacoronavirus RNA Genomes
Method: single particle / : Moura TR, Purta E, Bernat A, Baulin E, Mukherjee S, Bujnicki JM

PDB-8qo2: 
Conserved Structures and Dynamics in 5-Proximal Regions of Betacoronavirus RNA Genomes
Method: single particle / : Moura TR, Purta E, Bernat A, Baulin E, Mukherjee S, Bujnicki JM
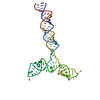
PDB-8qo3: 
Conserved Structures and Dynamics in 5-Proximal Regions of Betacoronavirus RNA Genomes
Method: single particle / : Moura TR, Purta E, Bernat A, Baulin E, Mukherjee S, Bujnicki JM

PDB-8qo4: 
Conserved Structures and Dynamics in 5-Proximal Regions of Betacoronavirus RNA Genomes
Method: single particle / : Moura TR, Purta E, Bernat A, Baulin E, Mukherjee S, Bujnicki JM

PDB-8qo5: 
Conserved Structures and Dynamics in 5-Proximal Regions of Betacoronavirus RNA Genomes
Method: single particle / : Moura TR, Purta E, Bernat A, Baulin E, Mukherjee S, Bujnicki JM

EMDB-40983: 
Negative stain EM assembly of MYC, JAZ, and NINJA complex
Method: single particle / : Zhou XE, Zhang Y, Zhou Y, He Q, Cao X, Kariapper L, Suino-Powell K, Zhu Y, Zhang F, Karsten M

PDB-8t2i: 
Negative stain EM assembly of MYC, JAZ, and NINJA complex
Method: single particle / : Zhou XE, Zhang Y, Zhou Y, He Q, Cao X, Kariapper L, Suino-Powell K, Zhu Y, Zhang F, Karsten M

EMDB-26604: 
H1 Solomon Islands 2006 hemagglutinin in complex with Ab111
Method: single particle / : Windsor IW, Caradonna TM, Schmidt AG
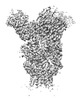
EMDB-26605: 
H1 Solomon Islands 2006 hemagglutinin in complex with Ab109
Method: single particle / : Windsor IW, Caradonna TM, Schmidt AG

PDB-7umm: 
H1 Solomon Islands 2006 hemagglutinin in complex with Ab109
Method: single particle / : Windsor IW, Caradonna TM, Schmidt AG

EMDB-25039: 
Structure of H1 NC99 influenza hemagglutinin bound to Fab 310-63E6
Method: single particle / : Ward A, Torrents de la Pena A

EMDB-25040: 
Structure of H1 influenza hemagglutinin bound to Fab 310-39G10
Method: single particle / : Torrents de la Pena A, Ward AB

PDB-7scn: 
Structure of H1 NC99 influenza hemagglutinin bound to Fab 310-63E6
Method: single particle / : Ward A, Torrents de la Pena A

PDB-7sco: 
Structure of H1 influenza hemagglutinin bound to Fab 310-39G10
Method: single particle / : Torrents de la Pena A, Ward AB

EMDB-13149: 
Mammalian orthoreovirus T3SA- in complex with the neural receptor NgR1
Method: single particle / : Strebl M, Sutherland DM, Yu X, Hu L, Wang Z, Prasad BVV, Stehle T, Dermody TS

EMDB-13150: 
Mammalian orthoreovirus T3SA-
Method: single particle / : Strebl M, Sutherland DM, Yu X, Hu L, Wang Z, Prasad BVV, Stehle T, Dermody TS

EMDB-11953: 
SARS-CoV-2 S 2P trimer in complex with monovalent DARPin R2 (State 1) - Composite Map
Method: single particle / : Hurdiss DL, Drulyte I

EMDB-11954: 
SARS-CoV-2 S 2P trimer in complex with monovalent DARPin R2 (State 2)
Method: single particle / : Hurdiss DL, Drulyte I

EMDB-14810: 
SARS-CoV-2 S 2P trimer in complex with monovalent DARPin R2 (State 1) - Consensus Map
Method: single particle / : Hurdiss DL, Drulyte I

EMDB-14811: 
SARS-CoV-2 S 2P trimer in complex with monovalent DARPin R2 (State 1) - Focused Refinement
Method: single particle / : Hurdiss DL, Drulyte I

EMDB-11860: 
Subtomogram average structure of anammoxosomal nitrite oxidoreductase
Method: subtomogram averaging / : Dietrich L

EMDB-11861: 
Helical reconstruction of nitrite oxidoreductase from anammox bacteria
Method: helical / : Parey K

EMDB-24279: 
Cryo-EM map of SARS-CoV-2 HexaPro Spike in complex with Ab090 Fab
Method: single particle / : Altomare CG, Feldman J, Schmidt AG, Bajic G

EMDB-23502: 
Cryo-EM structure of the Pre3-1 20S proteasome core particle
Method: single particle / : Schnell HM, Walsh Jr RM

EMDB-23503: 
Cryo-EM structure of Pre-15S proteasome core particle assembly intermediate purified from Pre3-1 proteasome mutant (G34D)
Method: single particle / : Schnell HM, Walsh Jr RM

EMDB-23508: 
Cryo-EM structure of 13S proteasome core particle assembly intermediate purified from Pre3-1 proteasome mutant (G34D)
Method: single particle / : Schnell HM, Walsh Jr RM

PDB-7ls5: 
Cryo-EM structure of the Pre3-1 20S proteasome core particle
Method: single particle / : Schnell HM, Walsh Jr RM, Rawson S, Hanna JW

PDB-7ls6: 
Cryo-EM structure of Pre-15S proteasome core particle assembly intermediate purified from Pre3-1 proteasome mutant (G34D)
Method: single particle / : Schnell HM, Walsh Jr RM, Rawson S, Hanna JW

PDB-7lsx: 
Cryo-EM structure of 13S proteasome core particle assembly intermediate purified from Pre3-1 proteasome mutant (G34D)
Method: single particle / : Schnell HM, Walsh Jr RM, Rawson S, Hanna JW

EMDB-22829: 
Human Tom70 in complex with SARS CoV2 Orf9b
Method: single particle / : QCRG Structural Biology Consortium

PDB-7kdt: 
Human Tom70 in complex with SARS CoV2 Orf9b
Method: single particle / : QCRG Structural Biology Consortium

EMDB-11002: 
CryoEM structure of bovine cytochrome bc1 in complex with a tetrahydroquinolone inhibitor
Method: single particle / : Muench SP, Johnson R, Amporndanai K, Atonyuk S

EMDB-20159: 
Rotavirus A-VP3-8mer ssRNA complex (RVA-VP3-RNA)
Method: single particle / : Kumar D, Yu X

EMDB-0146: 
Yeast RNA polymerase I elongation complex stalled by cyclobutane pyrimidine dimer (CPD)
Method: single particle / : Sanz-Murillo M, Xu J, Gil-Carton D, Wang D, Fernandez-Tornero C

EMDB-0147: 
Yeast RNA polymerase I elongation complex stalled by cyclobutane pyrimidine dimer (CPD) with fully-ordered A49
Method: single particle / : Sanz-Murillo M, Xu J, Gil-Carton D, Wang D, Fernandez-Tornero C

PDB-6h67: 
Yeast RNA polymerase I elongation complex stalled by cyclobutane pyrimidine dimer (CPD)
Method: single particle / : Sanz-Murillo M, Xu J, Gil-Carton D, Wang D, Fernandez-Tornero C

PDB-6h68: 
Yeast RNA polymerase I elongation complex stalled by cyclobutane pyrimidine dimer (CPD) with fully-ordered A49
Method: single particle / : Sanz-Murillo M, Xu J, Gil-Carton D, Wang D, Fernandez-Tornero C

EMDB-3178: 
Cryo-EM structure of yeast RNA polymerase III elongation complex at 3.9 A
Method: single particle / : Hoffmann NA, Jakobi AJ, Moreno-Morcillo M, Glatt S, Kosinski J, Hagen WJ, Sachse C, Muller CW

EMDB-3179: 
Cryo-EM structure of yeast apo RNA polymerase III at 4.6 A
Method: single particle / : Hoffmann NA, Jakobi AJ, Moreno-Morcillo M, Glatt S, Kosinski J, Hagen WJ, Sachse C, Muller CW

EMDB-3180: 
Cryo-EM structure of yeast apo RNA polymerase III at 4.7 A
Method: single particle / : Hoffmann NA, Jakobi AJ, Moreno-Morcillo M, Glatt S, Kosinski J, Hagen WJ, Sachse C, Muller CW

PDB-5fj8: 
Cryo-EM structure of yeast RNA polymerase III elongation complex at 3. 9 A
Method: single particle / : Hoffmann NA, Jakobi AJ, Moreno-Morcillo M, Glatt S, Kosinski J, Hagen WJ, Sachse C, Muller CW

PDB-5fj9: 
Cryo-EM structure of yeast apo RNA polymerase III at 4.6 A
Method: single particle / : Hoffmann NA, Jakobi AJ, Moreno-Morcillo M, Glatt S, Kosinski J, Hagen WJ, Sachse C, Muller CW

PDB-5fja: 
Cryo-EM structure of yeast RNA polymerase III at 4.7 A
Method: single particle / : Hoffmann NA, Jakobi AJ, Moreno-Morcillo M, Glatt S, Kosinski J, Hagen WJ, Sachse C, Muller CW

EMDB-5996: 
Electron cryo-microscopy of chimeric EGFP-VP2 capsid
Method: single particle / : Pascual E, Mata CP, Gomez-Blanco J, Carrascosa JL, Caston JR

EMDB-5997: 
Electron cryo-microscopy of chimeric HT-VP2-466 capsid
Method: single particle / : Pascual E, Mata CP, Gomez-Blanco J, Carrascosa JL, Caston JR

EMDB-6182: 
3D structure of RepA-WH1 double filaments
Method: helical / : Torreira E, Moreno M, Fuentes-Perez ME, Fernandez C, Martin-Benito J, Moreno-Herrero F, Giraldo R, Llorca O
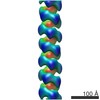
EMDB-6183: 
3D structure of RepA-WH1 single filaments
Method: helical / : Torreira E, Moreno M, Fuentes-Perez ME, Fernandez C, Martin-Benito J, Moreno-Herrero F, Giraldo R, Llorca O
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
