-Search query
-Search result
Showing 1 - 50 of 2,362 items for (author: mei & z)
EMDB-43779: 
Rat GluN1-GluN2B NMDA receptor channel in complex with glycine, glutamate, and EU-1622-A, in open-channel conformation
Method: single particle / : Chou TH, Furukawa H
EMDB-43780: 
Rat GluN1-GluN2B NMDA receptor channel in complex with glycine, glutamate, and EU-1622-A, in nonactive1 conformation
Method: single particle / : Chou TH, Furukawa H
EMDB-43781: 
Rat GluN1-GluN2B NMDA receptor channel in apo conformation
Method: single particle / : Chou TH, Furukawa H
EMDB-43782: 
Rat GluN1-GluN2B NMDA receptor channel in complex with glycine
Method: single particle / : Chou TH, Furukawa H

EMDB-43783: 
Rat GluN1-GluN2B NMDA receptor channel in complex with glutamate
Method: single particle / : Chou TH, Furukawa H
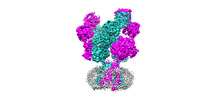
EMDB-44586: 
Rat GluN1-GluN2B NMDA receptor channel in complex with glycine, glutamate, and EU-1622-A, in open-channel conformation, C1 symmetry
Method: single particle / : Chou TH, Furukawa H

PDB-9are: 
Rat GluN1-GluN2B NMDA receptor channel in complex with glycine, glutamate, and EU-1622-A, in open-channel conformation
Method: single particle / : Chou TH, Furukawa H

PDB-9arf: 
Rat GluN1-GluN2B NMDA receptor channel in complex with glycine, glutamate, and EU-1622-A, in nonactive1 conformation
Method: single particle / : Chou TH, Furukawa H

PDB-9arg: 
Rat GluN1-GluN2B NMDA receptor channel in apo conformation
Method: single particle / : Chou TH, Furukawa H

PDB-9arh: 
Rat GluN1-GluN2B NMDA receptor channel in complex with glycine
Method: single particle / : Chou TH, Furukawa H

PDB-9ari: 
Rat GluN1-GluN2B NMDA receptor channel in complex with glutamate
Method: single particle / : Chou TH, Furukawa H

PDB-9bib: 
Rat GluN1-GluN2B NMDA receptor channel in complex with glycine, glutamate, and EU-1622-A, in open-channel conformation, C1 symmetry
Method: single particle / : Chou TH, Furukawa H

EMDB-37441: 
FCP tetramer in Chaetoceros gracilis
Method: single particle / : Feng Y, Li Z, Zhou C, Shen JR, Liu C, Wang W

EMDB-37442: 
FCP pentamer in Chaetoceros gracilis
Method: single particle / : Feng Y, Li Z, Zhou C, Liu C, Shen JR, Wang W

PDB-8wck: 
FCP tetramer in Chaetoceros gracilis
Method: single particle / : Feng Y, Li Z, Zhou C, Shen JR, Liu C, Wang W

PDB-8wcl: 
FCP pentamer in Chaetoceros gracilis
Method: single particle / : Feng Y, Li Z, Zhou C, Liu C, Shen JR, Wang W

EMDB-38460: 
Cryo-EM structure of SARS-CoV-2 Omicron BA.2.86 spike protein(6P), 1-RBD-up state
Method: single particle / : Li LJ, Gu YH, Shi KY, Qi JX, Gao GF

EMDB-38463: 
Cryo-EM structure of SARS-CoV-2 Omicron EG.5 spike protein(6P), RBD-closed state
Method: single particle / : Li LJ, Gu YH, Shi KY, Qi JX, Gao GF

EMDB-38476: 
Cryo-EM structure of SARS-CoV-2 Omicron HV.1 spike protein(6P), RBD-closed state
Method: single particle / : Li LJ, Gu YH, Shi KY, Qi JX, Gao GF

EMDB-38488: 
Cryo-EM structure of SARS-CoV-2 Omicron EG.5.1 spike protein(6P), RBD-closed state
Method: single particle / : Li LJ, Gu YH, Shi KY, Qi JX, Gao GF

EMDB-38495: 
SARS-CoV-2 Omicron EG.5.1 RBD in complex with human ACE2 (local refined from the spike protein)
Method: single particle / : Li LJ, Gu YH, Shi KY, Qi JX, Gao GF

EMDB-38496: 
SARS-CoV-2 Omicron HV.1 RBD in complex with human ACE2 (local refinement from the spike protein)
Method: single particle / : Li LJ, Gu YH, Shi KY, Qi JX, Gao GF

EMDB-38498: 
Cryo-EM structure of SARS-CoV-2 Omicron EG.5.1 spike protein(6P) in complex with human ACE2
Method: single particle / : Li LJ, Gu YH, Shi KY, Qi JX, Gao GF

EMDB-38502: 
Cryo-EM structure of SARS-CoV-2 Omicron BA.2.86 spike protein(6P) in complex with human ACE2
Method: single particle / : Li LJ, Gu YH, Shi KY, Qi JX, Gao GF

EMDB-38505: 
Cryo-EM structure of SARS-CoV-2 Omicron HV.1 spike protein(6P) in complex with human ACE2
Method: single particle / : Li LJ, Gu YH, Shi KY, Qi JX, Gao GF

EMDB-38826: 
Cryo-EM structure of SARS-CoV-2 Omicron JN.1 spike protein in complex with human ACE2
Method: single particle / : Li LJ, Gu YH, Qi JX, Gao GF

EMDB-38827: 
Cryo-EM structure of SARS-CoV-2 Omicron JN.1 RBD in complex with human ACE2 (local refinement from the spike protein)
Method: single particle / : Li LJ, Gu YH, Qi JX, Gao GF

EMDB-38937: 
Cryo-EM structure of SARS-CoV-2 Omicron JN.1 spike protein
Method: single particle / : Li LJ, Gu YH, Qi JX, Gao GF

EMDB-38983: 
Cryo-EM structure of SARS-CoV-2 Omicron BA.2.86 RBD in complex with human ACE2 and S309 Fab
Method: single particle / : Li LJ, Gu YH, Qi JX, Gao GF

PDB-8xlv: 
Cryo-EM structure of SARS-CoV-2 Omicron BA.2.86 spike protein(6P), 1-RBD-up state
Method: single particle / : Li LJ, Gu YH, Shi KY, Qi JX, Gao GF

PDB-8xm5: 
Cryo-EM structure of SARS-CoV-2 Omicron EG.5 spike protein(6P), RBD-closed state
Method: single particle / : Li LJ, Gu YH, Shi KY, Qi JX, Gao GF

PDB-8xmg: 
Cryo-EM structure of SARS-CoV-2 Omicron HV.1 spike protein(6P), RBD-closed state
Method: single particle / : Li LJ, Gu YH, Shi KY, Qi JX, Gao GF

PDB-8xmt: 
Cryo-EM structure of SARS-CoV-2 Omicron EG.5.1 spike protein(6P), RBD-closed state
Method: single particle / : Li LJ, Gu YH, Shi KY, Qi JX, Gao GF

PDB-8xn2: 
SARS-CoV-2 Omicron EG.5.1 RBD in complex with human ACE2 (local refined from the spike protein)
Method: single particle / : Li LJ, Gu YH, Shi KY, Qi JX, Gao GF

PDB-8xn3: 
SARS-CoV-2 Omicron HV.1 RBD in complex with human ACE2 (local refinement from the spike protein)
Method: single particle / : Li LJ, Gu YH, Shi KY, Qi JX, Gao GF

PDB-8xn5: 
Cryo-EM structure of SARS-CoV-2 Omicron EG.5.1 spike protein(6P) in complex with human ACE2
Method: single particle / : Li LJ, Gu YH, Shi KY, Qi JX, Gao GF

PDB-8xnf: 
Cryo-EM structure of SARS-CoV-2 Omicron BA.2.86 spike protein(6P) in complex with human ACE2
Method: single particle / : Li LJ, Gu YH, Shi KY, Qi JX, Gao GF

PDB-8xnk: 
Cryo-EM structure of SARS-CoV-2 Omicron HV.1 spike protein(6P) in complex with human ACE2
Method: single particle / : Li LJ, Gu YH, Shi KY, Qi JX, Gao GF

PDB-8y16: 
Cryo-EM structure of SARS-CoV-2 Omicron JN.1 spike protein in complex with human ACE2
Method: single particle / : Li LJ, Gu YH, Qi JX, Gao GF

PDB-8y18: 
Cryo-EM structure of SARS-CoV-2 Omicron JN.1 RBD in complex with human ACE2 (local refinement from the spike protein)
Method: single particle / : Li LJ, Gu YH, Qi JX, Gao GF

PDB-8y5j: 
Cryo-EM structure of SARS-CoV-2 Omicron JN.1 spike protein
Method: single particle / : Li LJ, Gu YH, Qi JX, Gao GF

PDB-8y6a: 
Cryo-EM structure of SARS-CoV-2 Omicron BA.2.86 RBD in complex with human ACE2 and S309 Fab
Method: single particle / : Li LJ, Gu YH, Qi JX, Gao GF

EMDB-50504: 
in situ 70S Myxococcus xanthus ribosome
Method: subtomogram averaging / : Xu P, Schumacher D, Liu C, Dickmanns M, Beck F, Plitzko J, Baumeister W, Sogaard-Andersen L

EMDB-17418: 
CryoEM structure of a C7-symmetrical GroEL7-GroES7 cage in presence of ADP-BeFx
Method: single particle / : Wagner J, Beck F, Bracher A, Caravajal AI, Wan W, Bohn S, Koerner R, Baumeister W, Fernandez-Busnadiego R, Hartl FU

EMDB-17420: 
CryoEM structure of a GroEL7-GroES7 cage with encapsulated disordered substrate MetK in the presence of ADP-BeFx
Method: single particle / : Wagner J, Beck F, Bracher A, Caravajal AI, Wan W, Bohn S, Koerner R, Baumeister W, Fernandez-Busnadiego R, Hartl FU

EMDB-17421: 
CryoEM structure of a GroEL7-GroES7 cage with encapsulated ordered substrate MetK in the presence of ADP-BeFx
Method: single particle / : Wagner J, Beck F, Bracher A, Caravajal AI, Wan W, Bohn S, Koerner R, Baumeister W, Fernandez-Busnadiego R, Hartl FU

EMDB-17422: 
Density for MetK encapsulated in the GroEL7-GroES7 cage
Method: single particle / : Wagner J, Caravajal AI, Beck F, Bracher A, Wan W, Bohn S, Koerner R, Fernandez-Busnadiego R, Baumeister W, Hartl FU

EMDB-17423: 
Symmetry-averaged GroEL7-GroES7 chamber with encapsulated disordered substrate MetK obtained by in vitro cryo electron tomography
Method: subtomogram averaging / : Bracher A, Wagner J, Caravajal AI, Beck F, Wan W, Bohn S, Koerner R, Baumeister W, Fernandez-Busnadiego R, Hartl FU
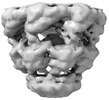
EMDB-17424: 
Symmetry-averaged GroEL7-GroES7 chamber with encapsulated ordered substrate MetK obtained by in vitro cryo electron tomography
Method: subtomogram averaging / : Wagner J, Caravajal AI, Beck F, Bracher A, Wan W, Bohn S, Koerner R, Baumeister W, Fernandez-Busnadiego R, Hartl FU

EMDB-17425: 
Structure average of GroEL14 complexes found in the cytosol of Escherichia coli overexpressing GroEL obtained by cryo electron tomography
Method: subtomogram averaging / : Wagner J, Caravajal AI, Beck F, Bracher A, Wan W, Bohn S, Koerner R, Baumeister W, Fernandez-Busnadiego R, Hartl FU
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
