-Search query
-Search result
Showing 1 - 50 of 157 items for (author: mascola & jr)
EMDB-41048: 
Lassa GPC Trimer in complex with Fab 8.11G and nanobody D5
Method: single particle / : Gorman J, Kwong PD
EMDB-28617: 
Cryo-EM structure of HIV-1 BG505 DS-SOSIP ENV trimer bound to VRC34.01 FAB
Method: single particle / : Pletnev S, Kwong P

EMDB-28618: 
Cryo-EM structure of HIV-1 BG505 DS-SOSIP ENV trimer bound to VRC34.01-COMBO1 FAB
Method: single particle / : Pletnev S, Kwong P

EMDB-28619: 
Cryo-EM structure of HIV-1 BG505 DS-SOSIP ENV trimer bound to VRC34.01-MM28 FAB
Method: single particle / : Pletnev S, Kwong P

EMDB-28910: 
Glycan-Base ConC Env Trimer
Method: single particle / : Olia AS, Kwong PD

EMDB-41089: 
Cryo-EM structure of RSV preF in complex with Fab 2.4K
Method: single particle / : McCool RS, McLellan JS

EMDB-41302: 
Lassa GPC trimer in complex with Fab GP23
Method: single particle / : Gorman J, Kwong PD

EMDB-26859: 
Ligand-free Lassa GPC Trimer with C3 Symmetry
Method: single particle / : Gorman J, Kwong PD
EMDB-26740: 
Ligand-free Lassa GPC Trimer with C1 Symmetry
Method: single particle / : Gorman J, Kwong PD

EMDB-29396: 
Antibody vFP53.02 in complex with HIV-1 envelope trimer BG505 DS-SOSIP
Method: single particle / : Wang S, Kwong PD
EMDB-29836: 
vFP52.02 Fab in complex with BG505 DS-SOSIP Env trimer
Method: single particle / : Gorman J, Kwong PD

EMDB-29880: 
Cryo-EM structure of vFP49.02 Fab in complex with HIV-1 Env BG505 DS-SOSIP.664 (conformation 1)
Method: single particle / : Changela A, Gorman J, Kwong PD

EMDB-29881: 
Cryo-EM structure of vFP49.02 Fab in complex with HIV-1 Env BG505 DS-SOSIP.664 (conformation 2)
Method: single particle / : Changela A, Gorman J, Kwong PD

EMDB-29882: 
Cryo-EM structure of vFP49.02 Fab in complex with HIV-1 Env BG505 DS-SOSIP.664 (conformation 3)
Method: single particle / : Changela A, Gorman J, Kwong PD

EMDB-29905: 
vFP48.02 Fab in complex with BG505 DS-SOSIP Env trimer
Method: single particle / : Gorman J, Kwong PD
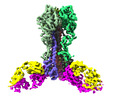
EMDB-27139: 
Cryo-EM structure of the VRC321 clinical trial, vaccine-elicited, human antibody 1B06 in complex with a stabilized NC99 HA trimer
Method: single particle / : Gorman J, Kwong PD
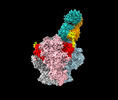
EMDB-29209: 
Structure of Bispecific CAP256V2LS-J3 Fab in complex with BG505 DS-SOSIP.664
Method: single particle / : Gorman J, Kwong PD

EMDB-26044: 
H10ssF: ferritin-based nanoparticle displaying H10 hemagglutinin stabilized stem epitopes
Method: single particle / : Gallagher JR, Harris AK

EMDB-24128: 
Cryo-EM structure of broadly neutralizing V2-apex-targeting antibody J033 in complex with HIV-1 Env
Method: single particle / : Zhou T, Gao F

EMDB-25794: 
Cryo-EM structure of SARS-CoV-2 spike in complex with antibodies B1-182.1 and A19-61.1
Method: single particle / : Zhou T, Tsybovsky T, Kwong PD

EMDB-25797: 
Locally refined region of SARS-CoV-2 spike in complex with antibodies B1-182.1 and A19-61.1
Method: single particle / : Zhou T, Tsybovsky T, Kwong PD

EMDB-25806: 
Locally refined region of SARS-CoV-2 spike in complex with antibody A19-46.1
Method: single particle / : Zhou T, Kwong PD

EMDB-25807: 
Cryo-EM structure of SARS-CoV-2 Omicron spike in complex with antibody A19-46.1
Method: single particle / : Zhou T, Kwong PD

EMDB-25808: 
Cryo-EM structure of SARS-CoV-2 Omicron spike in complex with antibodies A19-46.1 and B1-182.1
Method: single particle / : Zhou T, Kwong PD

EMDB-26256: 
Local refinement of cryo-EM structure of the interface of the Omicron RBD in complex with antibodies B-182.1 and A19-46.1
Method: single particle / : Zhou T, kwong PD

EMDB-24071: 
Cryo-EM structure of broadly neutralizing V2-apex-targeting antibody J038 in complex with HIV-1 Env
Method: single particle / : Zhou T, Gao F

EMDB-25792: 
Cryo-EM structure of the spike of SARS-CoV-2 Omicron variant of concern
Method: single particle / : Zhou T, Tsybovsky T, Kwong PD

EMDB-25105: 
Structure of SARS-CoV S protein in complex with Receptor Binding Domain antibody DH1047
Method: single particle / : Gobeil S, Acharya P

EMDB-23098: 
Cryo-EM structure of the VRC316 clinical trial, vaccine-elicited, human antibody 316-310-1B11 in complex with an H2 CAN05 HA trimer
Method: single particle / : Gorman J, Kwong PD

EMDB-23816: 
Cryo-EM structure of the VRC310 clinical trial, vaccine-elicited, human antibody 310-030-1D06 Fab in complex with an H1 NC99 HA trimer
Method: single particle / : Gorman J, Kwong PD

EMDB-23312: 
BG505 SOSIP.v5.2 in complex with VRC40.01 and RM19R Fabs
Method: single particle / : Cottrell CA, Torres JL, Wu NR, Ward AB

EMDB-23982: 
Structure of freshly purified SARS-CoV-2 S2P spike at pH 7.4
Method: single particle / : Tsybovsky Y, Olia AS, Kwong PD

EMDB-23983: 
Structure of aged SARS-CoV-2 S2P spike at pH 7.4
Method: single particle / : Tsybovsky Y, Olia AS, Kwong PD

EMDB-23984: 
Structure of SARS-CoV-2 S2P spike at pH 7.4 refolded by low-pH treatment
Method: single particle / : Tsybovsky Y, Olia AS, Kwong PD

EMDB-23605: 
Cryo-EM Structure of disulfide stabilized HMPV F v4-B
Method: single particle / : Gorman J, Kwong PD

EMDB-23625: 
HMPV F v3-B in complex with MPE33 Fab
Method: single particle / : Gorman J, Kwong PD

EMDB-23914: 
Cryo-EM structure of SARS-CoV-2 spike in complex with neutralizing antibody B1-182.1 that targets the receptor-binding domain
Method: single particle / : Zhou T, Tsybovsky T, Kwong PD

EMDB-23915: 
Cryo-EM structure of SARS-CoV-2 spike in complex with neutralizing antibody B1-182.1 that targets the receptor-binding domain
Method: single particle / : Zhou T, Tsybovsky T, Kwong PD

EMDB-22302: 
Cryo-EM Structure of Vaccine-Elicited Rhesus Antibody 789-203-3C12 in Complex with Stabilized SI06 (A/Solomon Islands/3/06) Influenza Hemagglutinin Trimer
Method: single particle / : Cerutti G, Gorman J, Kwong PD, Shapiro L

EMDB-23498: 
Cryo-EM structure of SARS-CoV-2 spike in complex with neutralizing antibody A23-58.1 that targets the receptor-binding domain
Method: single particle / : Zhou T, Tsybovsky Y

EMDB-23499: 
Cryo-EM structure of SARS-CoV-2 spike in complex with neutralizing antibody A23-58.1 that targets the receptor-binding domain
Method: single particle / : Zhou T, Tsybovsky T

EMDB-23521: 
Prefusion RSV F glycoprotein bound by neutralizing site V-directed antibody ADI-14442
Method: single particle / : Gilman MSA, McLellan JS

EMDB-23424: 
Cryo-EM map of Q23.17_DS-SOSIP in complex with Glycan276-Dependent Broadly Neutralizing Antibody 179NC75 Fab
Method: single particle / : Manne K, Acharya P

EMDB-22804: 
Cryo-EM structure of SRR2899884.46167H+MEDI8852L fab in complex with Victoria HA
Method: single particle / : Gorman J, Kwong PD

EMDB-23520: 
Cryo-EM structure of RSV preF bound by Fabs 32.4K and 01.4B
Method: single particle / : Wrapp D, McLellan JS

EMDB-22935: 
Elicitation of broadly protective immunity to influenza viruses by multivalent hemagglutinin nanoparticle vaccines
Method: single particle / : Park YJ, Veesler D

EMDB-22938: 
H5 hemagglutinin ectodomain bound to 1 polyclonal Fab fragment elicited by the qsMosaic-I53_dn5 immunogen
Method: single particle / : Park YJ, Veesler D

EMDB-22939: 
H5 hemagglutinin ectodomain bound to 2 polyclonal Fab fragments elicited by the qsMosaic-I53_dn5 immunogen
Method: single particle / : Park YJ, Veesler D

EMDB-22940: 
H5 hemagglutinin ectodomain bound to 3 polyclonal Fab fragments elicited by the qsMosaic-I53_dn5 immunogen
Method: single particle / : Park YJ, Veesler D

EMDB-22937: 
Localized reconstruction of the H1 A/Michigan/45/2015 ectodomain displayed at the surface of I53_dn5 nanoparticle
Method: single particle / : Acton OJ, Park YJ, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
