-Search query
-Search result
Showing 1 - 50 of 53 items for (author: marlovits & tc)
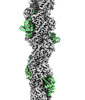
EMDB-16424: 
F-actin decorated by SipA497-669
Method: helical / : Yuan B, Wald J, Marlovits TC
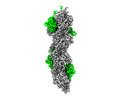
EMDB-16425: 
F-actin decorated by SipA426-685
Method: helical / : Yuan B, Wald J, Marlovits TC

EMDB-16805: 
Cryo-EM structure of PcrV/Fab(30-B8)
Method: single particle / : Yuan B, Simonis A, Marlovits TC

EMDB-16807: 
Cryo-EM structure of PcrV/Fab(11-E5)
Method: single particle / : Yuan B, Simonis A, Marlovits TC

EMDB-15362: 
Cryo-EM structure of Darobactin 22 bound BAM complex
Method: single particle / : Yuan B, Marlovits TC

EMDB-15363: 
Cryo-EM structure of Darobactin 9 bound BAM complex
Method: single particle / : Yuan B, Marlovits TC

EMDB-15085: 
RuvAB branch migration motor complexed to the Holliday junction - composite map 1 and 2
Method: single particle / : Wald J, Marlovits TC

EMDB-13294: 
RuvAB branch migration motor complexed to the Holliday junction - RuvB AAA+ state s1 [t2 dataset]
Method: single particle / : Wald J, Marlovits TC

EMDB-13295: 
RuvAB branch migration motor complexed to the Holliday junction - RuvB AAA+ state s2 [t2 dataset]
Method: single particle / : Wald J, Marlovits TC

EMDB-13296: 
RuvAB branch migration motor complexed to the Holliday junction - RuvB AAA+ state s3 [t2 dataset]
Method: single particle / : Wald J, Marlovits TC

EMDB-13297: 
RuvAB branch migration motor complexed to the Holliday junction - RuvB AAA+ state s4 [t2 dataset]
Method: single particle / : Wald J, Marlovits TC

EMDB-13298: 
RuvAB branch migration motor complexed to the Holliday junction - RuvB AAA+ state s5 [t2 dataset]
Method: single particle / : Wald J, Marlovits TC

EMDB-13299: 
RuvAB branch migration motor complexed to the Holliday junction - RuvB AAA+ state s0+A [t2 dataset]
Method: single particle / : Wald J, Marlovits TC

EMDB-13300: 
RuvAB branch migration motor complexed to the Holliday junction - RuvB AAA+ state s0-A [t2 dataset]
Method: single particle / : Wald J, Marlovits TC

EMDB-13301: 
RuvAB branch migration motor complexed to the Holliday junction - RuvB AAA+ state s0+A [t1 dataset]
Method: single particle / : Wald J, Marlovits TC

EMDB-13302: 
RuvAB branch migration motor complexed to the Holliday junction - RuvB AAA+ state s1 [t1 dataset]
Method: single particle / : Wald J, Marlovits TC

EMDB-13303: 
RuvAB branch migration motor complexed to the Holliday junction - RuvA-HJ core [t2 dataset]
Method: single particle / : Wald J, Marlovits TC

EMDB-13304: 
RuvAB branch migration motor complexed to the Holliday junction - RuvAB-HJ half [dataset t2]
Method: single particle / : Wald J, Marlovits TC

EMDB-13305: 
RuvAB branch migration motor complexed to the Holliday junction - RuvAB-HJ pseudo half [dataset t2]
Method: single particle / : Wald J, Marlovits TC

EMDB-15126: 
RuvAB branch migration motor complexed to the Holliday junction - full tripartite particle [dataset t2]
Method: single particle / : Wald J, Marlovits TC

EMDB-13542: 
HsPepT1 bound to Ala-Phe in the outward facing occluded conformation
Method: single particle / : Killer M, Wald J, Pieprzyk J, Marlovits TC, Loew C

EMDB-13543: 
HsPepT1 bound to Ala-Phe in the outward facing open conformation
Method: single particle / : Killer M, Wald J, Pieprzyk J, Marlovits TC, Loew C

EMDB-13544: 
HsPepT2 bound to Ala-Phe in the inward facing partially occluded conformation
Method: single particle / : Killer M, Wald J, Pieprzyk J, Marlovits TC, Loew C

EMDB-13545: 
Apo HsPepT1 in the outward facing open conformation
Method: single particle / : Killer M, Wald J, Pieprzyk J, Marlovits TC, Loew C

EMDB-13054: 
Cryo-EM structure of nonameric EPEC SctV-C
Method: single particle / : Yuan B, Wald J, Fahrenkamp D, Marlovits TC

EMDB-11311: 
Structure of the Salmonella PrgI needle filament attached to the basal body
Method: helical / : Kotov V, Lunelli M, Wald J, Kolbe M, Marlovits TC

EMDB-11312: 
Structure of the Shigella MxiH needle filament attached to the basal body
Method: helical / : Lunelli M, Kotov V, Marlovits TC, Kolbe M

EMDB-12514: 
Structure of an intact ESX-5 inner membrane complex, C1 symmetry
Method: single particle / : Bunduc CM, Fahrenkamp D, Wald J, Ummels R, Bitter W, Houben ENG, Marlovits TC

EMDB-12517: 
Structure of an intact ESX-5 inner membrane complex, C3 symmetry
Method: single particle / : Bunduc CM, Fahrenkamp D, Wald J, Ummels R, Bitter W, Houben ENG, Marlovits TC

EMDB-12521: 
MycP5-free ESX-5 inner membrane complex, state I
Method: single particle / : Bunduc CM, Fahrenkamp D, Wald J, Ummels R, Bitter W, Houben ENG, Marlovits TC

EMDB-12522: 
MycP5-free ESX-5 inner membrane complex, State II
Method: single particle / : Bunduc CM, Fahrenkamp D, Wald J, Ummels R, Bitter W, Houben ENG, Marlovits TC

EMDB-12518: 
Structure of the periplasmic assembly from the ESX-5 inner membrane complex, C1 symmetry
Method: single particle / : Bunduc CM, Fahrenkamp D, Wald J, Ummels R, Bitter W, Houben ENG, Marlovits TC

EMDB-12519: 
Periplasmic assembly of the intact ESX-5 inner membrane complex, C3 symmetry
Method: single particle / : Bunduc CM, Fahrenkamp D, Wald J, Ummels R, Bitter W, Houben ENG, Marlovits TC

EMDB-12520: 
Cytosolic bridge of an intact ESX-5 inner membrane complex
Method: single particle / : Bunduc CM, Fahrenkamp D, Wald J, Ummels R, Bitter W, Houben ENG, Marlovits TC

EMDB-12523: 
Structure of an intact ESX-5 inner membrane complex, EccC5 extended
Method: single particle / : Bunduc CM, Fahrenkamp D, Wald J, Ummels R, Bitter W, Houben ENG, Marlovits TC

EMDB-12525: 
Structure of an intact ESX-5 inner membrane complex, EccC contracted
Method: single particle / : Bunduc CM, Fahrenkamp D, Wald J, Ummels R, Bitter W, Houben ENG, Marlovits TC

EMDB-11780: 
Apo-state type 3 secretion system export apparatus complex from Salmonella enterica typhimurium
Method: single particle / : Goessweiner-Mohr N, Fahrenkamp D, Miletic S, Wald J, Marlovits T

EMDB-11781: 
Substrate-engaged type 3 secretion system needle complex from Salmonella enterica typhimurium - SpaR state 1
Method: single particle / : Fahrenkamp D, Goessweiner-Mohr N, Miletic S, Wald J, Marlovits T

EMDB-10220: 
Cryo-EM structure of rhinovirus-B5 complexed to antiviral OBR-5-340
Method: single particle / : Wald J, Goessweiner-Mohr N, Blaas D, Pasin M

EMDB-10221: 
Cryo-EM structure of rhinovirus-B5
Method: single particle / : Wald J, Goessweiner-Mohr N, Blaas D, Pasin M

EMDB-10222: 
Cryo-EM structure of rhinovirus-A89
Method: single particle / : Wald J, Goessweiner-Mohr N, Blaas D, Pasin M

EMDB-3596: 
The structure of the ESX-5 mycobacterial type VII secretion system membrane complex
Method: single particle / : Ciccarelli L, Marlovits TC

EMDB-2480: 
Tomogram of Injectisome (wild type) from Salmonella typhiumurium (substrate trapped) in situ
Method: electron tomography / : Radics J, Konigsmaier L, Marlovits TC

EMDB-2481: 
Injectisome (wild type) from Salmonella typhimurium (substrate free)
Method: single particle / : Radics J, Konigsmaier L, Marlovits TC

EMDB-2482: 
Injectisome (wild type) from Salmonella typhimurium (substrate trapped)
Method: single particle / : Radics J, Konigsmaier L, Marlovits TC

EMDB-2483: 
Tomogram of Injectisome (wild type) from Salmonella typhimurium (substrate free) in situ
Method: electron tomography / : Radics J, Konigsmaier L, Marlovits TC

EMDB-1871: 
Three-dimensional model of Salmonella's needle complex at subnanometer resolution
Method: single particle / : Schraidt O, Marlovits TC

EMDB-1874: 
Three-dimensional model of Salmonella's needle complex at subnanometer resolution
Method: single particle / : Schraidt O, Marlovits TC

EMDB-1875: 
Three dimensional structure of the injectisome
Method: single particle / : Schraidt O, Marlovits TC

EMDB-1214: 
Assembly of the inner rod determines needle length in the type III secretion injectisome.
Method: single particle / : Marlovits TC, Kubori T, Lara-Tejero M, Thomas D, Unger VM, Galan JE
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
