-Search query
-Search result
Showing 1 - 50 of 94 items for (author: makino & f)

EMDB-36190: 
Cryo-EM Structure of Na-dithionite Reduced Membrane-bound Fructose Dehydrogenase from Gluconobacter japonicus
Method: single particle / : Suzuki Y, Miyata T, Makino F, Tanaka H, Namba K, Sowa K, Kitazumi Y, Shirai O

EMDB-36191: 
Cryo-EM Structure of K-ferricyanide Oxidized Membrane-bound Fructose Dehydrogenase from Gluconobacter japonicus
Method: single particle / : Suzuki Y, Miyata T, Makino F, Tanaka H, Namba K, Sowa K, Kitazumi Y, Shirai O

PDB-8jej: 
Cryo-EM Structure of Na-dithionite Reduced Membrane-bound Fructose Dehydrogenase from Gluconobacter japonicus
Method: single particle / : Suzuki Y, Miyata T, Makino F, Tanaka H, Namba K, Sowa K, Kitazumi Y, Shirai O

PDB-8jek: 
Cryo-EM Structure of K-ferricyanide Oxidized Membrane-bound Fructose Dehydrogenase from Gluconobacter japonicus
Method: single particle / : Suzuki Y, Miyata T, Makino F, Tanaka H, Namba K, Sowa K, Kitazumi Y, Shirai O
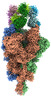
EMDB-33972: 
Cryo-EM structure of the SARS-CoV-2 spike protein (3-up RBD) bound to neutralizing antibody CSW1-1805
Method: single particle / : Anzai I, Fujita J, Ono C, Kosaka Y, Miyamoto Y, Shichinohe S, Takada K, Torii S, Taguwa S, Suzuki K, Makino F, Kajita T, Inoue T, Namba K, Watanabe T, Matsuura Y
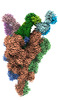
EMDB-33973: 
Cryo-EM structure of the SARS-CoV-2 spike protein (1-up RBD) bound to neutralizing antibody CSW2-1353
Method: single particle / : Anzai I, Fujita J, Ono C, Kosaka Y, Miyamoto Y, Shichinohe S, Takada K, Torii S, Taguwa S, Suzuki K, Makino F, Kajita T, Inoue T, Namba K, Watanabe T, Matsuura Y
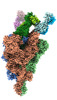
EMDB-33974: 
Cryo-EM structure of the SARS-CoV-2 spike protein (2-up RBD) bound to neutralizing antibody CSW2-1353
Method: single particle / : Anzai I, Fujita J, Ono C, Kosaka Y, Miyamoto Y, Shichinohe S, Takada K, Torii S, Taguwa S, Suzuki K, Makino F, Kajita T, Inoue T, Namba K, Watanabe T, Matsuura Y
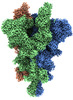
EMDB-33975: 
Cryo-EM structure of the SARS-CoV-2 spike protein (2-up RBD) with C480A mutation
Method: single particle / : Anzai I, Fujita J, Ono C, Kosaka Y, Miyamoto Y, Shichinohe S, Takada K, Torii S, Taguwa S, Suzuki K, Makino F, Kajita T, Inoue T, Namba K, Watanabe T, Matsuura Y

EMDB-34368: 
Cryo-EM Structure of Membrane-Bound Alcohol Dehydrogenase from Gluconobacter oxydans
Method: single particle / : Adachi T, Miyata T, Makino F, Tanaka H, Namba K, Sowa K, Kitazumi Y, Shirai O

EMDB-34369: 
Cryo-EM Structure of Membrane-Bound Aldehyde Dehydrogenase from Gluconobacter oxydans
Method: single particle / : Adachi T, Miyata T, Makino F, Tanaka H, Namba K, Sowa K, Kitazumi Y, Shirai O

PDB-8gy2: 
Cryo-EM Structure of Membrane-Bound Alcohol Dehydrogenase from Gluconobacter oxydans
Method: single particle / : Adachi T, Miyata T, Makino F, Tanaka H, Namba K, Sowa K, Kitazumi Y, Shirai O

PDB-8gy3: 
Cryo-EM Structure of Membrane-Bound Aldehyde Dehydrogenase from Gluconobacter oxydans
Method: single particle / : Adachi T, Miyata T, Makino F, Tanaka H, Namba K, Sowa K, Kitazumi Y, Shirai O

EMDB-33643: 
The SARS-CoV-2 receptor binding domain bound with the Fab fragment of a human neutralizing antibody Ab816
Method: single particle / : Uchikubo T, Shirouzu M

EMDB-33644: 
The SARS-CoV-2 receptor binding domain bound with the Fab fragment of a human neutralizing antibody Ab803
Method: single particle / : Uchikubo T, Shirouzu M

PDB-7y6l: 
The SARS-CoV-2 receptor binding domain bound with the Fab fragment of a human neutralizing antibody Ab816
Method: single particle / : Uchikubo T, Shirouzu M

PDB-7y6n: 
The SARS-CoV-2 receptor binding domain bound with the Fab fragment of a human neutralizing antibody Ab803
Method: single particle / : Uchikubo T, Shirouzu M

EMDB-33065: 
The SARS-CoV-2 receptor binding domain bound with the Fab fragment of a human neutralizing antibody Ab765
Method: single particle / : Kamada K, Shirouzu M
EMDB-33066: 
The SARS-CoV-2 receptor binding domain bound with the Fab fragment of a human neutralizing antibody Ab712
Method: single particle / : Kamada K, Shirouzu M

EMDB-33067: 
The SARS-CoV-2 receptor binding domain bound with the Fab fragment of a human neutralizing antibody Ab709
Method: single particle / : Kamada K, Shirouzu M

EMDB-33068: 
The SARS-CoV-2 receptor binding domain bound with the Fab fragment of a human neutralizing antibody Ab847
Method: single particle / : Kamada K, Shirouzu M

PDB-7x93: 
The SARS-CoV-2 receptor binding domain bound with the Fab fragment of a human neutralizing antibody Ab765
Method: single particle / : Kamada K, Shirouzu M

PDB-7x94: 
The SARS-CoV-2 receptor binding domain bound with the Fab fragment of a human neutralizing antibody Ab712
Method: single particle / : Kamada K, Shirouzu M

PDB-7x95: 
The SARS-CoV-2 receptor binding domain bound with the Fab fragment of a human neutralizing antibody Ab709
Method: single particle / : Kamada K, Shirouzu M

PDB-7x96: 
The SARS-CoV-2 receptor binding domain bound with the Fab fragment of a human neutralizing antibody Ab847
Method: single particle / : Kamada K, Shirouzu M

EMDB-34106: 
GroEL on EG-grid stored for 3 months after graphene oxidation
Method: single particle / : Fujita J, Makino F, Asahara H, Moriguchi M, Kumano S, Anzai I, Kishikawa J, Matsuura Y, Kato T, Namba K, Inoue T
EMDB-34743: 
GroEL on Quantifoil grid
Method: single particle / : Fujita J, Makino F, Asahara H, Moriguchi M, Kumano S, Anzai I, Kishikawa J, Matsuura Y, Kato T, Namba K, Inoue T

EMDB-32764: 
Cryo-EM Structure of Membrane-bound Fructose Dehydrogenase from Gluconobacter japonicus
Method: single particle / : Suzuki Y, Makino F

PDB-7wsq: 
Cryo-EM Structure of Membrane-bound Fructose Dehydrogenase from Gluconobacter japonicus
Method: single particle / : Suzuki Y, Makino F, Miyata T, Tanaka H, Namba K, Sowa K, Kitazumi Y, Shirai O

EMDB-33059: 
The SARS-CoV-2 receptor binding domain bound with the Fab fragment of a human neutralizing antibody Ab354
Method: single particle / : Kamada K, Shirouzu M

EMDB-33060: 
The SARS-CoV-2 receptor binding domain bound with the Fab fragment of a human neutralizing antibody Ab159
Method: single particle / : Kamada K, Shirouzu M

EMDB-33061: 
The SARS-CoV-2 receptor binding domain bound with the Fab fragment of a human neutralizing antibody Ab188
Method: single particle / : Kamada K, Shirouzu M

EMDB-33062: 
The SARS-CoV-2 receptor binding domain bound with the Fab fragment of a human neutralizing antibody Ab326
Method: single particle / : Kamada K, Shirouzu M

EMDB-33063: 
The SARS-CoV-2 receptor binding domain bound with an Fv-clasp form of a human neutralizing antibody Ab496
Method: single particle / : Kamada K, Shirouzu M

EMDB-33064: 
The SARS-CoV-2 receptor binding domain bound with the Fab fragment of a human neutralizing antibody Ab445
Method: single particle / : Kamada K, Shirouzu M

PDB-7x8w: 
The SARS-CoV-2 receptor binding domain bound with the Fab fragment of a human neutralizing antibody Ab354
Method: single particle / : Kamada K, Shirouzu M

PDB-7x8y: 
The SARS-CoV-2 receptor binding domain bound with the Fab fragment of a human neutralizing antibody Ab159
Method: single particle / : Kamada K, Shirouzu M

PDB-7x8z: 
The SARS-CoV-2 receptor binding domain bound with the Fab fragment of a human neutralizing antibody Ab188
Method: single particle / : Kamada K, Shirouzu M

PDB-7x90: 
The SARS-CoV-2 receptor binding domain bound with the Fab fragment of a human neutralizing antibody Ab326
Method: single particle / : Kamada K, Shirouzu M

PDB-7x91: 
The SARS-CoV-2 receptor binding domain bound with an Fv-clasp form of a human neutralizing antibody Ab496
Method: single particle / : Kamada K, Shirouzu M

PDB-7x92: 
The SARS-CoV-2 receptor binding domain bound with the Fab fragment of a human neutralizing antibody Ab445
Method: single particle / : Kamada K, Shirouzu M

EMDB-32262: 
Cryo-EM Structure of Membrane-bound Fructose Dehydrogenase from Gluconobacter japonicus
Method: single particle / : Suzuki Y, Makino F, Miyata T, Tanaka H, Namba K, Sowa K, Kitazumi Y, Shirai O

PDB-7w2j: 
Cryo-EM Structure of Membrane-bound Fructose Dehydrogenase from Gluconobacter japonicus
Method: single particle / : Suzuki Y, Makino F, Miyata T, Tanaka H, Namba K, Sowa K, Kitazumi Y, Shirai O

EMDB-32159: 
GroEL on hydrophilized graphene grid (high particle density)
Method: single particle / : Fujita J, Makino F, Asahara H, Moriguchi M, Kumano S, Anzai I, Kishikawa J, Matsuura Y, Kato T, Namba K, Inoue T
EMDB-32160: 
SARS-CoV-2 spike protein (1-up RBD) on EG-grid
Method: single particle / : Fujita J, Makino F, Asahara H, Moriguchi M, Kumano S, Anzai I, Kishikawa J, Matsuura Y, Kato T, Namba K, Inoue T
EMDB-32161: 
SARS-CoV-2 spike protein (1-up RBD) on Quantifoil grid
Method: single particle / : Fujita J, Makino F, Asahara H, Moriguchi M, Kumano S, Anzai I, Kishikawa J, Matsuura Y, Kato T, Namba K, Inoue T
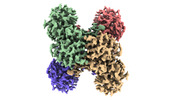
EMDB-32162: 
GAPDH on EG-grid
Method: single particle / : Fujita J, Makino F, Asahara H, Moriguchi M, Kumano S, Anzai I, Kishikawa J, Matsuura Y, Kato T, Namba K, Inoue T

EMDB-31605: 
Cryo-EM structure of cyanobacterial photosystem I in the presence of ferredoxin and cytochrome c6
Method: single particle / : Li J, Kurisu G

PDB-7fix: 
Cryo-EM structure of cyanobacterial photosystem I in the presence of ferredoxin and cytochrome c6
Method: single particle / : Li J, Kurisu G
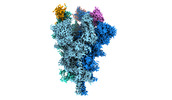
EMDB-32078: 
Cryo-EM structure of the SARS-CoV-2 spike protein (2-up RBD) bound to neutralizing nanobodies P86
Method: single particle / : Maeda R, Fujita J, Konishi Y, Kazuma Y, Yamazaki H, Anzai I, Yamaguchi K, Kasai K, Nagata K, Yamaoka Y, Miyakawa K, Ryo A, Shirakawa K, Makino F, Matsuura Y, Inoue T, Imura A, Namba K, Takaori-Kondo A
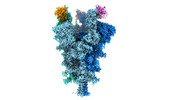
EMDB-32079: 
Cryo-EM structure of the SARS-CoV-2 spike protein (3-up RBD) bound to neutralizing nanobodies P86
Method: single particle / : Maeda R, Fujita J, Konishi Y, Kazuma Y, Yamazaki H, Anzai I, Yamaguchi K, Kasai K, Nagata K, Yamaoka Y, Miyakawa K, Ryo A, Shirakawa K, Makino F, Matsuura Y, Inoue T, Imura A, Namba K, Takaori-Kondo A
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
