-Search query
-Search result
Showing 1 - 50 of 141 items for (author: m & w & parker)

EMDB-43033: 
Structure of 80alpha portal protein expressed in E. coli
Method: single particle / : Kizziah JL, Mukherjee A, Dokland T

EMDB-43142: 
SaPI1 mature capsid structure containing DNA
Method: single particle / : Mukherjee A, Kizziah JL, Dokland T

EMDB-43143: 
SaPI1 mature capsid structure without DNA
Method: single particle / : Mukherjee A, Kizziah JL, Dokland T

EMDB-43145: 
SaPI1 portal structure in mature capsids containing DNA
Method: single particle / : Kizziah JL, Mukherjee A, Dokland T

EMDB-43146: 
SaPI1 portal structure in mature capsids without DNA
Method: single particle / : Mukherjee A, Kizziah JL, Dokland T

EMDB-43147: 
SaPI1 portal-capsid interface in mature capsids with DNA
Method: single particle / : Kizziah JL, Mukherjee A, Dokland T

PDB-8v8b: 
Structure of 80alpha portal protein expressed in E. coli
Method: single particle / : Kizziah JL, Mukherjee A, Dokland T

PDB-8vd4: 
SaPI1 mature capsid structure containing DNA
Method: single particle / : Mukherjee A, Kizziah JL, Dokland T

PDB-8vd5: 
SaPI1 mature capsid structure without DNA
Method: single particle / : Mukherjee A, Kizziah JL, Dokland T

PDB-8vd8: 
SaPI1 portal structure in mature capsids containing DNA
Method: single particle / : Kizziah JL, Mukherjee A, Dokland T

PDB-8vdc: 
SaPI1 portal structure in mature capsids without DNA
Method: single particle / : Mukherjee A, Kizziah JL, Dokland T

PDB-8vde: 
SaPI1 portal-capsid interface in mature capsids with DNA
Method: single particle / : Kizziah JL, Mukherjee A, Dokland T

EMDB-27641: 
The structure of the interleukin 11 signalling complex, truncated gp130
Method: single particle / : Metcalfe RD, Hanssen E, Griffin MDW

EMDB-27642: 
The structure of the IL-11 signalling complex, with full-length extracellular gp130
Method: single particle / : Metcalfe RD, Hanssen E, Griffin MDW

EMDB-29753: 
Tomogram of an apical end of a Toxoplasma Gondii tachyzoite displaying an extruded conoid
Method: electron tomography / : Martinez M, Chang YW, Mageswaran SK

EMDB-29754: 
Cryptosporidium parvum sporozoite apical end
Method: electron tomography / : Martinez M, Chang YW, Mageswaran SK
EMDB-29755: 
Cryptosporidium parvum sporozoite basal end
Method: electron tomography / : Martinez M, Chang YW, Mageswaran SK

EMDB-29784: 
Subtomogram average of the preconoidal rings from Cryptosporidium parvum sporozoites
Method: subtomogram averaging / : Martinez M, Mageswaran SK, Chang YW

EMDB-29791: 
Refined subtomogram average of the top preconoidal ring from Cryptosporidium parvum sporozoites
Method: subtomogram averaging / : Martinez M, Mageswaran SK, Chang YW

EMDB-29801: 
Refined subtomogram average of the bottom preconoidal ring from Cryptosporidium parvum sporozoites
Method: subtomogram averaging / : Martinez M, Mageswaran SK, Chang YW
EMDB-29808: 
Subtomogram average of the IMC surface filaments, top view, from Cryptosporidium parvum sporozoites
Method: subtomogram averaging / : Martinez M, Mageswaran SK, Chang YW

EMDB-29809: 
IMC surface filament, sawtooth conformation, from Cryptosporidium parvum sporozoites
Method: subtomogram averaging / : Martinez M, Mageswaran SK, Chang YW

EMDB-29810: 
IMC surface filament, side view, from Cryptosporidium parvum sporozoites
Method: subtomogram averaging / : Martinez M, Mageswaran SK, Chang YW

EMDB-29826: 
Upper preconoidal ring from Toxoplasma gondii tachyzoites
Method: subtomogram averaging / : Martinez M, Mageswaran SK, Chang YW

EMDB-29827: 
Lower preconoidal ring of Toxoplasma gondii tachyzoites
Method: subtomogram averaging / : Martinez M, Mageswaran SK, Chang YW

EMDB-29832: 
Preconoidal rings of Toxoplasma gondii tachyzoites
Method: subtomogram averaging / : Martinez M, Mageswaran SK, Chang YW

EMDB-29835: 
Basal IMC pore of Cryptosporidium parvum sporozoites
Method: subtomogram averaging / : Martinez M, Mageswaran SK, Chang YW

EMDB-29838: 
Preconoidal rings of Toxoplasma gondii tachyzoites with Formin1 conditionally depleted
Method: subtomogram averaging / : Martinez M, Mageswaran SK, Chang YW

EMDB-29839: 
Upper preconoidal ring of Toxoplasma gondii tachyzoites with Formin1 conditionally depleted
Method: subtomogram averaging / : Martinez M, Mageswaran SK, Chang YW

EMDB-29840: 
Lower preconoidal ring from Toxoplasma gondii tachyzoites with Formin1 conditionally depleted
Method: subtomogram averaging / : Martinez M, Mageswaran SK, Chang YW
EMDB-16269: 
Cryo-EM structure of rat SLC22A6 in the apo state
Method: single particle / : Parker JL, Kato T, Newstead S

EMDB-16270: 
Cryo-EM structure of rat SLC22A6 bound to tenofovir
Method: single particle / : Parker JL, Kato T, Newstead S
EMDB-16271: 
Cryo-EM structure of rat SLC22A6 bound to probenecid
Method: single particle / : Parker JL, Kato T, Newstead S
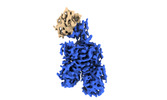
EMDB-16280: 
Cryo-EM structure of rat SLC22A6 bound to alpha-ketoglutaric acid
Method: single particle / : Parker JL, Kato T, Newstead S

EMDB-16977: 
Cryo-EM structure of rat SLC22A6 bound to alpha-ketoglutaric acid in a low occupancy state
Method: single particle / : Parker JL, Kato T, Newstead S

PDB-8bvr: 
Cryo-EM structure of rat SLC22A6 in the apo state
Method: single particle / : Parker JL, Kato T, Newstead S

PDB-8bvs: 
Cryo-EM structure of rat SLC22A6 bound to tenofovir
Method: single particle / : Parker JL, Kato T, Newstead S

PDB-8bvt: 
Cryo-EM structure of rat SLC22A6 bound to probenecid
Method: single particle / : Parker JL, Kato T, Newstead S

PDB-8bw7: 
Cryo-EM structure of rat SLC22A6 bound to alpha-ketoglutaric acid
Method: single particle / : Parker JL, Kato T, Newstead S

PDB-8omu: 
Cryo-EM structure of rat SLC22A6 bound to alpha-ketoglutaric acid in a low occupancy state
Method: single particle / : Parker JL, Kato T, Newstead S

EMDB-28120: 
DH272.2 Fab in Complex with CH848 TF DS SOSIP Env
Method: single particle / : Fera D

EMDB-33233: 
Cryo-EM structure of EDS1 and SAG101 with ATP-APDR
Method: single particle / : Huang SJ, Jia AL, Han ZF, Chai JJ

PDB-7xjp: 
Cryo-EM structure of EDS1 and SAG101 with ATP-APDR
Method: single particle / : Huang SJ, Jia AL, Han ZF, Chai JJ

EMDB-33144: 
Cryo-EM structure of EDS1 and PAD4
Method: single particle / : Huang SJ, Jia AL, Sun Y, Han ZF, Chai JJ

PDB-7xdd: 
Cryo-EM structure of EDS1 and PAD4
Method: single particle / : Huang SJ, Jia AL, Sun Y, Han ZF, Chai JJ

EMDB-24408: 
The Capsid Structure of the ChAdOx1 viral vector/chimpanzee adenovirus Y25
Method: single particle / : Baker AT, Boyd RJ, Sarkar D, Vermaas JV, Williams D, Singharoy A

PDB-7rd1: 
The Capsid Structure of the ChAdOx1 viral vector/chimpanzee adenovirus Y25
Method: single particle / : Baker AT, Boyd RJ, Sarkar D, Vermaas JV, Williams D, Singharoy A

EMDB-13266: 
Cryo EM structure of System XC- in complex with glutamate
Method: single particle / : Parker JL, Deme JC, Lea SM, Newstead S

EMDB-13267: 
Cryo EM structure of System XC-
Method: single particle / : Parker JL, Deme JC, Lea SM, Newstead S

PDB-7p9u: 
Cryo EM structure of System XC- in complex with glutamate
Method: single particle / : Parker JL, Deme JC, Lea SM, Newstead S
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
