-Search query
-Search result
Showing 1 - 50 of 229 items for (author: liu & hy)

EMDB-35931: 
cryo-EM structures of Ufd4 in complex with Ubc4-Ub
Method: single particle / : Ai HS, Mao JX, Wu XW, Cai HY, Pan M, Liu L
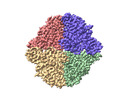
EMDB-42478: 
Trehalose Synthase (TreS) of Mycobacterium tuberculosis in complex with 6-TreAz compound
Method: single particle / : Pathirage R, Ronning DR

PDB-8uqv: 
Trehalose Synthase (TreS) of Mycobacterium tuberculosis in complex with 6-TreAz compound
Method: single particle / : Pathirage R, Ronning DR

EMDB-41907: 
Computationally Designed, Expandable O4 Octahedral Handshake Nanocage
Method: single particle / : Weidle C, Borst A
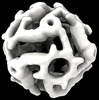
EMDB-42031: 
Computational Designed Nanocage O43_129_+8
Method: single particle / : Weidle C, Kibler RD

EMDB-43318: 
Twistless helix 12 repeat ring design R12B
Method: single particle / : Calise SJ, Kollman JM

EMDB-41423: 
Cryo-EM structure of DDB1dB:CRBN:Pomalidomide:SD40
Method: single particle / : Roy Burman SS, Hunkeler M, Fischer ES

EMDB-41424: 
Cryo-EM structure of DDB1dB:CRBN:PT-179:SD40, conformation 1
Method: single particle / : Roy Burman SS, Hunkeler M, Fischer ES

EMDB-41425: 
Cryo-EM structure of DDB1dB:CRBN:PT-179:SD40, conformation 2
Method: single particle / : Roy Burman SS, Hunkeler M, Fischer ES

EMDB-41777: 
Map from local refinement (focused on CRBN) of DDB1dB:CRBN:Pomalidomide:SD40
Method: single particle / : Roy Burman SS, Hunkeler M, Fischer ES

EMDB-41778: 
Map from local refinement (focused on CRBN) of DDB1dB:CRBN:PT-179:SD40, conformation 1
Method: single particle / : Roy Burman SS, Hunkeler M, Fischer ES

EMDB-41779: 
Map from local refinement (focused on CRBN) of DDB1dB:CRBN:PT-179:SD40, conformation 2
Method: single particle / : Roy Burman SS, Hunkeler M, Fischer ES

PDB-8tnp: 
Cryo-EM structure of DDB1dB:CRBN:Pomalidomide:SD40
Method: single particle / : Roy Burman SS, Hunkeler M, Fischer ES

PDB-8tnq: 
Cryo-EM structure of DDB1dB:CRBN:PT-179:SD40, conformation 1
Method: single particle / : Roy Burman SS, Hunkeler M, Fischer ES

PDB-8tnr: 
Cryo-EM structure of DDB1dB:CRBN:PT-179:SD40, conformation 2
Method: single particle / : Roy Burman SS, Hunkeler M, Fischer ES

EMDB-29974: 
Cryo-EM structure of synthetic tetrameric building block sC4
Method: single particle / : Redler RL, Huddy TF, Hsia Y, Baker D, Ekiert D, Bhabha G

EMDB-41364: 
CryoEM Structure of a Computationally Designed T3 Tetrahedral Nanocage
Method: single particle / : Weidle C, Borst AJ

EMDB-42906: 
Computational Designed Nanocage O43_129
Method: single particle / : Weidle C, Kibler RD

EMDB-42944: 
Computational Designed Nanocage O43_129_+4
Method: single particle / : Carr KD, Weidle C, Borst AJ

PDB-8gel: 
Cryo-EM structure of synthetic tetrameric building block sC4
Method: single particle / : Redler RL, Huddy TF, Hsia Y, Baker D, Ekiert D, Bhabha G

PDB-8tl7: 
CryoEM Structure of a Computationally Designed T3 Tetrahedral Nanocage
Method: single particle / : Weidle C, Borst AJ

PDB-8v3b: 
Computational Designed Nanocage O43_129_+4
Method: single particle / : Carr KD, Weidle C, Borst AJ

EMDB-34981: 
Cryo-EM structure of human NTCP-myr-preS1-YN9048Fab complex
Method: single particle / : Asami J, Shimizu T, Ohto U

EMDB-34982: 
Cryo-EM structure of human NTCP-myr-preS1-YN9016Fab complex
Method: single particle / : Asami J, Shimizu T, Ohto U

PDB-8hrx: 
Cryo-EM structure of human NTCP-myr-preS1-YN9048Fab complex
Method: single particle / : Asami J, Shimizu T, Ohto U

PDB-8hry: 
Cryo-EM structure of human NTCP-myr-preS1-YN9016Fab complex
Method: single particle / : Asami J, Shimizu T, Ohto U

EMDB-40219: 
A subtomogram averaged structure of Helicobacter pylori flagellar motor from a flgV deletion mutant.
Method: subtomogram averaging / : Liu J, Tachiyama S

EMDB-41046: 
Subtomogram average structure of the injectisome
Method: subtomogram averaging / : Park D
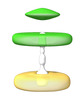
EMDB-41047: 
Intermembrane Tether between SCV and Mitochondria
Method: subtomogram averaging / : Park D, Liu J, Galan J

EMDB-29725: 
Vaccine-elicited human antibody 2C06 in complex with HIV-1 envelope trimer BG505 DS-SOSIP
Method: single particle / : Wang S, Morano NC, Shapiro L, Kwong PD

EMDB-29731: 
Vaccine-elicited human antibody 2C09 in complex with HIV-1 envelope trimer BG505 DS-SOSIP
Method: single particle / : Wang S, Kwong PD

PDB-8g4m: 
Vaccine-elicited human antibody 2C06 in complex with HIV-1 envelope trimer BG505 DS-SOSIP
Method: single particle / : Wang S, Morano NC, Shapiro L, Kwong PD

PDB-8g4t: 
Vaccine-elicited human antibody 2C09 in complex with HIV-1 envelope trimer BG505 DS-SOSIP
Method: single particle / : Wang S, Kwong PD

EMDB-29915: 
CryoEM map of a de novo designed octahedral nanocage with programmable volume; design cage_O4_34
Method: single particle / : Kibler RD, Borst AJ

EMDB-40070: 
Cryo-EM map of synthetic cage_O3_10 reconstructed without symmetry (C1)
Method: single particle / : Coudray N, Redler R, Hsia Y, Huddy TF, Baker D, Ekiert D, Bhabha G

EMDB-40071: 
Cryo-EM map of synthetic cage_O3_10 reconstructed with O symmetry
Method: single particle / : Coudray N, Redler R, Hsia Y, Huddy TF, Baker D, Ekiert D, Bhabha G

EMDB-40073: 
Cryo-EM map of synthetic cage_T3_5 reconstructed without symmetry (C1), with 1 monomer missing (class 3.0)
Method: single particle / : Coudray N, Redler R, Huddy TF, Hsia Y, Baker D, Ekiert D, Bhabha G

EMDB-40074: 
Cryo-EM map of synthetic cage_T3_5 reconstructed with T symmetry
Method: single particle / : Coudray N, Redler R, Huddy TF, Hsia Y, Baker D, Ekiert D, Bhabha G

EMDB-40075: 
Cryo-EM map of synthetic cage_T3_5 reconstructed without symmetry (C1)
Method: single particle / : Coudray N, Redler R, Huddy TF, Hsia Y, Baker D, Ekiert D, Bhabha G

EMDB-40076: 
Cryo-EM map of synthetic cage_T3_5+2 reconstructed without symmetry (C1)
Method: single particle / : Coudray N, Redler R, Huddy TF, Hsia Y, Baker D, Ekiert D, Bhabha G

EMDB-40072: 
Cryo-EM map of synthetic cage_T3_5 reconstructed without symmetry (C1), with 1 trimer missing (class 3.1)
Method: single particle / : Coudray N, Redler R, Huddy TF, Hsia Y, Baker D, Ekiert D, Bhabha G

EMDB-34276: 
Cryo-EM structure of CB2-G protein complex
Method: single particle / : Wu LJ, Hua T, Liu ZJ, Li XT, Chang H

EMDB-34277: 
Cryo-EM structure of CP-CB2-G protein complex
Method: single particle / : Wu LJ, Hua T, Liu ZJ, Li XT, Chang H

EMDB-34278: 
Cryo-EM structure of HU-CB2-G protein complex
Method: single particle / : Wu LJ, Hua T, Liu ZJ, Li XT, Chang H

EMDB-34279: 
Cryo-EM structure of LEI-CB2-Gi complex
Method: single particle / : Liu ZJ, Hua T, Li XT, Chang H, Wu LJ

PDB-8guq: 
Cryo-EM structure of CB2-G protein complex
Method: single particle / : Wu LJ, Hua T, Liu ZJ, Li XT, Chang H

PDB-8gur: 
Cryo-EM structure of CP-CB2-G protein complex
Method: single particle / : Wu LJ, Hua T, Liu ZJ, Li XT, Chang H

PDB-8gus: 
Cryo-EM structure of HU-CB2-G protein complex
Method: single particle / : Wu LJ, Hua T, Liu ZJ, Li XT, Chang H

PDB-8gut: 
Cryo-EM structure of LEI-CB2-Gi complex
Method: single particle / : Liu ZJ, Hua T, Li XT, Chang H, Wu LJ
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search




 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
