-Search query
-Search result
Showing 1 - 50 of 791 items for (author: lim & a)

EMDB-17988: 
Mycobacterium smegmatis inosine monophosphate dehydrogenase (IMPDH) apo form
Method: single particle / : Bulvas O, Kouba T, Pichova I

EMDB-18184: 
Mycobacterium smegmatis inosine monophosphate dehydrogenase (IMPDH) ATP-bound form
Method: single particle / : Bulvas O, Kouba T, Pichova I

EMDB-18600: 
Mycobacterium smegmatis inosine monophosphate dehydrogenase (IMPDH) ATP+GTP-bound form, compressed
Method: single particle / : Bulvas O, Kouba T, Pichova I

EMDB-18601: 
Mycobacterium smegmatis inosine monophosphate dehydrogenase (IMPDH) ATP+GTP-bound form, less-compressed
Method: single particle / : Bulvas O, Kouba T, Pichova I

EMDB-18602: 
Mycobacterium smegmatis inosine monophosphate dehydrogenase (IMPDH) ATP+ppGpp-bound form, compressed
Method: single particle / : Bulvas O, Kouba T, Pichova I

EMDB-18604: 
Mycobacterium smegmatis inosine monophosphate dehydrogenase (IMPDH) ATP+ppGpp-bound form, less-compressed
Method: single particle / : Bulvas O, Kouba T, Pichova I

EMDB-18606: 
Mycobacterium smegmatis inosine monophosphate dehydrogenase (IMPDH) ATP+IMP-bound form, extended
Method: single particle / : Bulvas O, Kouba T, Pichova I

EMDB-18607: 
Mycobacterium smegmatis inosine monophosphate dehydrogenase (IMPDH) ATP-bound form, compressed
Method: single particle / : Bulvas O, Kouba T, Pichova I
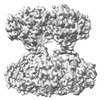
EMDB-18608: 
Mycobacterium smegmatis inosine monophosphate dehydrogenase (IMPDH) ATP+IMP-bound form, half-extended
Method: single particle / : Bulvas O, Kouba T, Pichova I

EMDB-43700: 
Cryo-EM map of LKB1-STRADalpha-MO25alpha from TFS Glacios with Gatan Alpine detector at 120 keV
Method: single particle / : Chan LM, Courteau BJ, Verba KA

EMDB-43701: 
Cryo-EM map of LKB1-STRADalpha-MO25alpha from TFS Glacios with Gatan Alpine detector at 200 keV
Method: single particle / : Chan LM, Courteau BJ, Verba KA

EMDB-43702: 
Cryo-EM map of LKB1-STRADalpha-MO25alpha from TFS Glacios with Gatan K3 detector at 200 keV
Method: single particle / : Chan LM, Courteau BJ, Verba KA

EMDB-43506: 
Cryo-EM structure of LKB1-STRADalpha-MO25alpha heterocomplex
Method: single particle / : Chan LM, Courteau BJ, Verba KA

PDB-8vsu: 
Cryo-EM structure of LKB1-STRADalpha-MO25alpha heterocomplex
Method: single particle / : Chan LM, Courteau BJ, Verba KA

EMDB-43527: 
Apoferritin at 100 keV on Alpine detector with a side-entry cryoholder
Method: single particle / : Wu M, Lander GC
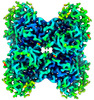
EMDB-43528: 
Aldolase at 100 keV on the Alpine detector with a side-entry cryoholder
Method: single particle / : Wu M, Lander GC
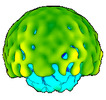
EMDB-50358: 
In vitro-induced genome-releasing intermediate of Rhodobacter microvirus Ebor computed with C5 symmetry
Method: single particle / : Bardy P, MacDonald CIW, Jenkins HT, Chechik M, Hart SJ, Turkenburg JP, Blaza JN, Fogg PCM, Antson AA

EMDB-50356: 
Empty capsid of Rhodobacter microvirus Ebor computed with I4 symmetry
Method: single particle / : Bardy P, MacDonald CIW, Jenkins HT, Byrom L, Chechik M, Hart SJ, Turkenburg JP, Blaza JN, Fogg PCM, Antson AA

EMDB-50357: 
Native capsid of Rhodobacter microvirus Ebor computed with I4 symmetry
Method: single particle / : Bardy P, MacDonald CIW, Jenkins HT, Chechik M, Hart SJ, Turkenburg JP, Blaza JN, Fogg PCM, Antson AA
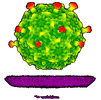
EMDB-50359: 
Rhodobacter microvirus Ebor attached to B10 host cell reconstructed by single particle analysis with applied C5 symmetry
Method: single particle / : Bardy P, MacDonald CIW, Jenkins HT, Chechik M, Hart SJ, Turkenburg JP, Blaza JN, Fogg PCM, Antson AA
EMDB-50360: 
Rhodobacter microvirus Ebor attached to the outer membrane vesicle
Method: subtomogram averaging / : Bardy P, Blaza JN, Jenkins HT, Nicholas TR, Konig HC, Alim NTB, Hart SJ, Turkenburg JP, Fogg PCM, Beatty JT, Antson AA

EMDB-50361: 
Rhodobacter microvirus Ebor attached to the host cell reconstructed by subtomogram averaging
Method: subtomogram averaging / : Bardy P, Traore DAK, Blaza JN, Jenkins HT, Nicholas TR, Hart SJ, Turkenburg JP, Fogg PCM, Antson AA

PDB-9ffg: 
Empty capsid of Rhodobacter microvirus Ebor computed with I4 symmetry
Method: single particle / : Bardy P, MacDonald CIW, Jenkins HT, Byrom L, Chechik M, Hart SJ, Turkenburg JP, Blaza JN, Fogg PCM, Antson AA

PDB-9ffh: 
Native capsid of Rhodobacter microvirus Ebor computed with I4 symmetry
Method: single particle / : Bardy P, MacDonald CIW, Jenkins HT, Chechik M, Hart SJ, Turkenburg JP, Blaza JN, Fogg PCM, Antson AA

EMDB-36429: 
Cryo-EM structure of dengue virus serotype 3 strain EHIE46200Y19 in complex with human antibody DENV-115 IgG at 4 deg C (subparticle LLR-LRR)
Method: single particle / : Fibriansah G, Ng TS, Tan AWK, Shi J, Lok SM

EMDB-36430: 
Cryo-EM structure of dengue virus serotype 3 strain 863DK in complex with human antibody DENV-115 Fab at 4 deg C (subparticle LLR-LRR)
Method: single particle / : Fibriansah G, Ng TS, Tan AWK, Shi J, Lok SM

EMDB-36431: 
Cryo-EM structure of dengue virus serotype 3 strain 863DK in complex with human antibody DENV-115 Fab at 37 deg C (subparticle LLR-LRR)
Method: single particle / : Fibriansah G, Ng TS, Tan AWK, Shi J, Lok SM

EMDB-36432: 
Cryo-EM structure of dengue virus serotype 3 strain 863DK in complex with human antibody DENV-290 Fab at 4 deg C (subparticle LLR-LRR)
Method: single particle / : Fibriansah G, Ng TS, Tan AWK, Shi J, Lok SM

EMDB-36433: 
Cryo-EM structure of dengue virus serotype 3 strain 863DK in complex with human antibody DENV-290 Fab at 37 deg C (subparticle LLR-LRR)
Method: single particle / : Fibriansah G, Ng TS, Tan AWK, Shi J, Lok SM

EMDB-36434: 
Cryo-EM structure of the small tail club shape particle of dengue virus serotype 3 strain CH53489 in complex with human antibody DENV-290 Fab at 37 deg C
Method: helical / : Fibriansah G, Ng TS, Tan AWK, Shi J, Lok SM

EMDB-36435: 
Cryo-EM structure of the big tail club shape particle of dengue virus serotype 3 strain CH53489 in complex with human antibody DENV-290 Fab at 37 deg C
Method: helical / : Fibriansah G, Ng TS, Tan AWK, Shi J, Lok SM

EMDB-36436: 
Cryo-EM structure of dengue virus serotype 3 strain EHIE46200Y19 in complex with human antibody DENV-115 IgG at 4 deg C
Method: single particle / : Fibriansah G, Ng TS, Tan AWK, Shi J, Lok SM

EMDB-36437: 
Cryo-EM structure of dengue virus serotype 3 strain 863DK in complex with human antibody DENV-115 Fab at 4 deg C
Method: single particle / : Fibriansah G, Ng TS, Tan AWK, Shi J, Lok SM

EMDB-36438: 
Cryo-EM structure of dengue virus serotype 3 strain 863DK in complex with human antibody DENV-115 Fab at 37 deg C
Method: single particle / : Fibriansah G, Ng TS, Tan AWK, Shi J, Lok SM

EMDB-36439: 
Cryo-EM structure of dengue virus serotype 3 strain 863DK in complex with human antibody DENV-290 Fab at 4 deg C
Method: single particle / : Fibriansah G, Ng TS, Tan AWK, Shi J, Lok SM

EMDB-36440: 
Cryo-EM structure of dengue virus serotype 3 strain 863DK in complex with human antibody DENV-290 Fab at 37 deg C
Method: single particle / : Fibriansah G, Ng TS, Tan AWK, Shi J, Lok SM

EMDB-36441: 
Cryo-EM structure of dengue virus serotype 3 strain CH53489 spherical particle in complex with human antibody DENV-290 Fab at 37 deg C
Method: single particle / : Fibriansah G, Ng TS, Tan AWK, Shi J, Lok SM

EMDB-44207: 
Ubiquitin E1-Ub-E2 tetrahedral transthiolation intermediate mimic (doubly Ub-loaded) - consensus map and model
Method: single particle / : Kochanczyk T, Lima CD

EMDB-44208: 
Ubiquitin E1-Ub-E2 tetrahedral transthiolation intermediate mimic (doubly Ub-loaded) - Ub(T) class 1 map and model from consensus
Method: single particle / : Kochanczyk T, Lima CD

EMDB-44209: 
Ubiquitin E1-Ub-E2 tetrahedral transthiolation intermediate mimic (doubly Ub-loaded) - Ub(T) class 10 map and model from consensus
Method: single particle / : Kochanczyk T, Lima CD

EMDB-44210: 
Ubiquitin E1-Ub-E2 tetrahedral transthiolation intermediate mimic (doubly Ub-loaded) - cluster 1 map and model (Ub(A)/ATP/Mg)
Method: single particle / : Kochanczyk T, Lima CD

EMDB-44211: 
Ubiquitin E1-Ub-E2 tetrahedral transthiolation intermediate mimic (doubly Ub-loaded) - cluster 2 map and model (Ub(A)/ATP/Mg)
Method: single particle / : Kochanczyk T, Lima CD

EMDB-44212: 
Ubiquitin E1-Ub-E2 tetrahedral transthiolation intermediate mimic (doubly Ub-loaded) - cluster 3 map and model (Ub(A)-AMP/PPi/Mg)
Method: single particle / : Kochanczyk T, Lima CD

EMDB-44213: 
Ubiquitin E1-Ub-E2 tetrahedral transthiolation intermediate mimic (doubly Ub-loaded) - cluster 4 map and model (Ub(A)-AMP/PPi/Mg)
Method: single particle / : Kochanczyk T, Lima CD

EMDB-44214: 
Ubiquitin E1-Ub-E2 tetrahedral transthiolation intermediate mimic (doubly Ub-loaded) - cluster 5 map and model (Ub(A)-AMP)
Method: single particle / : Kochanczyk T, Lima CD

EMDB-44215: 
Ubiquitin E1-Ub-E2 tetrahedral transthiolation intermediate mimic (doubly Ub-loaded) - Ub(T) class 1 map and model from cluster 1 (Ub(A)/ATP/Mg)
Method: single particle / : Kochanczyk T, Lima CD

EMDB-44216: 
Ubiquitin E1-Ub-E2 tetrahedral transthiolation intermediate mimic (doubly Ub-loaded) - Ub(T) class 10 map and model from cluster 5 (Ub(A)-AMP)
Method: single particle / : Kochanczyk T, Lima CD

PDB-9b5c: 
Ubiquitin E1-Ub-E2 tetrahedral transthiolation intermediate mimic (doubly Ub-loaded) - consensus map and model
Method: single particle / : Kochanczyk T, Lima CD

PDB-9b5d: 
Ubiquitin E1-Ub-E2 tetrahedral transthiolation intermediate mimic (doubly Ub-loaded) - Ub(T) class 1 map and model from consensus
Method: single particle / : Kochanczyk T, Lima CD

PDB-9b5e: 
Ubiquitin E1-Ub-E2 tetrahedral transthiolation intermediate mimic (doubly Ub-loaded) - Ub(T) class 10 map and model from consensus
Method: single particle / : Kochanczyk T, Lima CD
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
