-Search query
-Search result
Showing all 27 items for (author: liao & hy)

EMDB-42149: 
S1V2-72 Fab bound to EHA2 from influenza B/Malaysia/2506/2004
Method: single particle / : Finney J, Kong S, Walsh Jr RM, Harrison SC, Kelsoe G

PDB-8udg: 
S1V2-72 Fab bound to EHA2 from influenza B/Malaysia/2506/2004
Method: single particle / : Finney J, Kong S, Walsh Jr RM, Harrison SC, Kelsoe G

EMDB-35609: 
Cryo-EM structure of phosphoketolase from Bifidobacterium longum in octameric assembly
Method: single particle / : Chang CW, Tsai MD

EMDB-35610: 
Cryo-EM structure of phosphoketolase from Bifidobacterium longum in dimeric assembly
Method: single particle / : Chang CW, Tsai MD

EMDB-35611: 
Cryo-EM structure of cyanobacteria phosphoketolase complexed with AMPPNP in dimeric assembly
Method: single particle / : Chang CW, Tsai MD

EMDB-35612: 
Cryo-EM structure of cyanobacteria phosphoketolase complexed with AMPPNPin dodecameric assembly
Method: single particle / : Chang CW, Tsai MD

EMDB-35613: 
Cryo-EM structure of cyanobacteria phosphoketolase
Method: single particle / : Chang CW, Tsai MD

EMDB-35617: 
Cryo-EM structure of cyanobacteria phosphoketolase in dodecameric assembly
Method: single particle / : Chang CW, Tsai MD

EMDB-32832: 
SARS-CoV-2 Spike in complex with Fab of m31A7
Method: single particle / : Wu YM, Chen X

EMDB-32825: 
Negative stain volume of the mono-GlcNAc-decorated SARS-CoV-2 Spike
Method: single particle / : Chen X, Huang HY

EMDB-31470: 
Cryo-EM structure of SARS-CoV-2 spike in complex with a neutralizing antibody chAb-25 (Focused refinement of S-RBD and chAb-25 region)
Method: single particle / : Yang TJ, Yu PY, Wu HC, Hsu STD

EMDB-31471: 
Cryo-EM structure of SARS-CoV-2 spike in complex with a neutralizing antibody chAb-45 (Focused refinement of S-RBD and chAb-45 region)
Method: single particle / : Yang TJ, Yu PY, Wu HC, Hsu STD

PDB-7f62: 
Cryo-EM structure of SARS-CoV-2 spike in complex with a neutralizing antibody chAb-25 (Focused refinement of S-RBD and chAb-25 region)
Method: single particle / : Yang TJ, Yu PY, Wu HC, Hsu STD

PDB-7f63: 
Cryo-EM structure of SARS-CoV-2 spike in complex with a neutralizing antibody chAb-45 (Focused refinement of S-RBD and chAb-45 region)
Method: single particle / : Yang TJ, Yu PY, Wu HC, Hsu STD
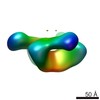
EMDB-8573: 
HIV-1 CH505 Transmitted Founder SOSIP.664 Env trimer in Complex with the DH576 Fab from the RV305 Trial
Method: single particle / : Fera D, Harrison SC

EMDB-6044: 
Ribosome conformation along minimum free-energy trajectory
Method: single particle / : Dashti A, Schwander P, Langlois R, Fung R, Li W, Hosseinizadeh A, Liao HY, Pallesen J, Sharma G, Stupina VA, Simon AE, Dinman J, Frank J, Ourmazd A

EMDB-2450: 
Cryo-EM map of the CSFV IRES in complex with the small ribosomal 40S subunit and DHX29
Method: single particle / : Hashem Y, des Georges A, Dhote A, Langlois R, Liao HY, Grassucci RA, Pestova TV, Hellen CUT, Frank J

EMDB-2451: 
Cryo-EM structure of the CSFV IRES in complex with eIF3, small ribosomal 40S subunit and DHX29
Method: single particle / : Hashem Y, des-Georges A, Dhote V, Langlois R, Liao HY, Grassucci RA, Pestova TV, Hellen CUT, Frank J

EMDB-5658: 
Cryo-electron microscopy structure of the mammalian 43S preinitiation complex bound to DHX29
Method: single particle / : Hashem Y, des Georges A, Dhote V, Langlois R, Liao HY, Grassucci RA, Hellen CUT, Pestova TV, Frank J

EMDB-2239: 
Cryo-electron microscopy structure of the Trypanosoma brucei 80S ribosome
Method: single particle / : Hashem Y, des Georges A, Fu J, Buss SN, Jossinet F, Jobe A, Zhang Q, Liao HY, Grassucci B, Bajaj C, Westhof E, Madison-Antenucci S, Frank J

EMDB-5359: 
New mRNA-tRNA translocation intermediates revealed by cryoEM: Class 4A (partially rotated (I) 70S ribosome with A-site and P-site tRNAs)
Method: single particle / : Agirrezabala X, Liao HY, Schreiner E, Fu J, Ortiz-Meoz RF, Schulten K, Green R, Frank J

EMDB-5360: 
New mRNA-tRNA translocation intermediates revealed by cryoEM: Class 3 (unrotated 70S ribosome with P-site tRNA)
Method: single particle / : Agirrezabala X, Liao HY, Schreiner E, Fu J, Ortiz-Meoz RF, Schulten K, Green R, Frank J

EMDB-5361: 
New mRNA-tRNA translocation intermediates revealed by cryoEM: Class 2 (unrotated 70S ribosome with A-site and P-site tRNAs)
Method: single particle / : Agirrezabala X, Liao HY, Schreiner E, Fu J, Ortiz-Meoz RF, Schulten K, Green R, Frank J

EMDB-5362: 
New mRNA-tRNA translocation intermediates revealed by cryoEM: Class 6 (rotated 70S ribosome with A-site and P-site tRNAs)
Method: single particle / : Agirrezabala X, Liao HY, Schreiner E, Fu J, Ortiz-Meoz RF, Schulten K, Green R, Frank J

EMDB-5363: 
New mRNA-tRNA translocation intermediates revealed by cryoEM: Class 5 (rotated 70S ribosome with A-site and P-site tRNAs)
Method: single particle / : Agirrezabala X, Liao HY, Schreiner E, Fu J, Ortiz-Meoz RF, Schulten K, Green R, Frank J

EMDB-5364: 
New mRNA-tRNA translocation intermediates revealed by cryoEM: Class 4B (partially rotated (II) 70S ribosome with A-site and P-site tRNAs)
Method: single particle / : Agirrezabala X, Liao HY, Schreiner E, Fu J, Ortiz-Meoz RF, Schulten K, Green R, Frank J
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search




 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
