-Search query
-Search result
Showing 1 - 50 of 2,388 items for (author: liang & h)

EMDB-42291: 
Structure of the human INTS9-INTS11-BRAT1 complex
Method: single particle / : Lin M, Tong L

EMDB-42292: 
Structure of the Drosophila IntS11-CG7044(dBRAT1) complex
Method: single particle / : Lin M, Tong L

EMDB-60100: 
SARS-CoV-2 spike trimer (6P) in complex with three R1-26 Fabs
Method: single particle / : Yan Q, Gao X, Liu B, Hou R, He P, Li Z, Chen Q, Wang J, He J, Chen L, Zhao J, Xiong X

EMDB-60101: 
SARS-CoV-2 spike trimer (6P) in complex with R1-26 Fab, head-to-head aggregate
Method: single particle / : Yan Q, Gao X, Liu B, Hou R, He P, Li Z, Chen Q, Wang J, He J, Chen L, Zhao J, Xiong X

EMDB-60102: 
SARS-CoV-2 spike trimer (6P) in complex with R1-26 Fab, focused refinement of RBD-Fab region
Method: single particle / : Yan Q, Gao X, Liu B, Hou R, He P, Li Z, Chen Q, Wang J, He J, Chen L, Zhao J, Xiong X

EMDB-60103: 
SARS-CoV-2 spike trimer (6P) in complex with two H18 Fabs
Method: single particle / : Yan Q, Gao X, Liu B, Hou R, He P, Li Z, Chen Q, Wang J, He J, Chen L, Zhao J, Xiong X

EMDB-60104: 
SARS-CoV-2 spike trimer (6P) in complex with three H18 Fabs
Method: single particle / : Yan Q, Gao X, Liu B, Hou R, He P, Li Z, Chen Q, Wang J, He J, Chen L, Zhao J, Xiong X

EMDB-60105: 
SARS-CoV-2 spike trimer (6P) in complex with three H18 Fabs, head-to-head aggregate (C1 symmetry)
Method: single particle / : Yan Q, Gao X, Liu B, Hou R, He P, Li Z, Chen Q, Wang J, He J, Chen L, Zhao J, Xiong X

EMDB-60106: 
SARS-CoV-2 spike trimer (6P) in complex with three H18 Fabs, head-to-head aggregate (C3 symmetry)
Method: single particle / : Yan Q, Gao X, Liu B, Hou R, He P, Li Z, Chen Q, Wang J, He J, Chen L, Zhao J, Xiong X

EMDB-60107: 
SARS-CoV-2 spike trimer (6P) in complex with two H18 and two R1-32 Fabs
Method: single particle / : Yan Q, Gao X, Liu B, Hou R, He P, Li Z, Chen Q, Wang J, He J, Chen L, Zhao J, Xiong X

EMDB-60108: 
SARS-CoV-2 spike trimer (6P) in complex with three H18 and three R1-32 Fabs
Method: single particle / : Yan Q, Gao X, Liu B, Hou R, He P, Li Z, Chen Q, Wang J, He J, Chen L, Zhao J, Xiong X

EMDB-60109: 
SARS-CoV-2 spike trimer (6P) in complex with three H18 and three R1-32 Fabs (one RBD rotated)
Method: single particle / : Yan Q, Gao X, Liu B, Hou R, He P, Li Z, Chen Q, Wang J, He J, Chen L, Zhao J, Xiong X

EMDB-60110: 
SARS-CoV-2 S1 in complex with H18 and R1-32 Fab
Method: single particle / : Yan Q, Gao X, Liu B, Hou R, He P, Li Z, Chen Q, Wang J, He J, Chen L, Zhao J, Xiong X

EMDB-60111: 
Dimer of SARS-CoV-2 S1 in complex with H18 and R1-32 Fabs
Method: single particle / : Yan Q, Gao X, Liu B, Hou R, He P, Li Z, Chen Q, Wang J, He J, Chen L, Zhao J, Xiong X
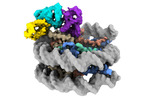
EMDB-38099: 
Cryo-EM structures of RNF168/UbcH5c-Ub in complex with H2AK13Ub nucleosomes determined by intein-based E2-Ub-NCP conjugation strategy
Method: single particle / : Ai HS, Tong ZB, Deng ZH, Pan M, Liu L
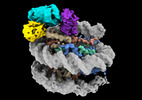
EMDB-38100: 
Cryo-EM structures of RNF168/UbcH5c-Ub/nucleosomes complex determined by activity-based chemical trapping strategy
Method: single particle / : Ai HS, Tong ZB, Deng ZH, Pan M, Liu L
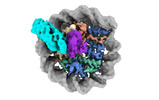
EMDB-38101: 
Cryo-EM structures of RNF168/UbcH5c-Ub in complex with H2AK13Ub nucleosomes determined by activity-based chemical trapping strategy (adjacent H2AK13/15 dual-monoubiquitination)
Method: single particle / : Ai HS, Tong ZB, Deng ZH, Pan M, Liu L
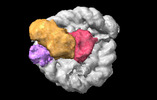
EMDB-38102: 
Cryo-EM map of RNF168/UbcH5c-Ub/nucleosome determined by E2-Ub-NCP conjugation strategy
Method: single particle / : Ai H, Zebin T, Deng Z, Pan M, Liu L

EMDB-38873: 
cryo-EM structure of Staphylococcus aureus(ATCC 29213) 50S ribosome in complex with MCX-190.
Method: single particle / : Li Y, Lu G, Li J, Pei X, Lin J

EMDB-38874: 
Cryo-EM structure of Staphylococcus aureus (15B196) 50S ribosome in complex with MCX-190.
Method: single particle / : Li Y, Lu G, Li J, Pei X, Lin J

EMDB-38875: 
Cryo-EM structure of Staphylococcus aureus 70S ribosome (strain 15B196) in complex with MCX-190.
Method: single particle / : Li Y, Lu G, Li J, Pei X, Lin J

EMDB-38876: 
cryo-EM structure of Staphylococcus aureus(ATCC 29213) 70S ribosome in complex with MCX-190.
Method: single particle / : Li Y, Lu G, Li J, Pei X, Lin J

PDB-8y36: 
cryo-EM structure of Staphylococcus aureus(ATCC 29213) 50S ribosome in complex with MCX-190.
Method: single particle / : Li Y, Lu G, Li J, Pei X, Lin J

PDB-8y37: 
Cryo-EM structure of Staphylococcus aureus (15B196) 50S ribosome in complex with MCX-190.
Method: single particle / : Li Y, Lu G, Li J, Pei X, Lin J

PDB-8y38: 
Cryo-EM structure of Staphylococcus aureus 70S ribosome (strain 15B196) in complex with MCX-190.
Method: single particle / : Li Y, Lu G, Li J, Pei X, Lin J

PDB-8y39: 
cryo-EM structure of Staphylococcus aureus(ATCC 29213) 70S ribosome in complex with MCX-190.
Method: single particle / : Li Y, Lu G, Li J, Pei X, Lin J

EMDB-44351: 
Synaptic Vesicle V-ATPase with synaptophysin and SidK, State 3, V1
Method: single particle / : Coupland EM, Rubinstein JL

PDB-9b8p: 
Synaptic Vesicle V-ATPase with synaptophysin and SidK, State 3, V1
Method: single particle / : Coupland EM, Rubinstein JL
EMDB-38784: 
The structure of fox ACE2 and PT RBD complex
Method: single particle / : sun JQ

EMDB-38792: 
The structure of fox ACE2 and SARS-CoV RBD complex
Method: single particle / : sun JQ

EMDB-42965: 
CryoEM structure of AriA-Ocr complex
Method: single particle / : Deep A, Corbett KD

EMDB-42966: 
CryoEM structure of AriA-AriB complex (Form I)
Method: single particle / : Deep A, Corbett KD

EMDB-42967: 
CryoEM structure of AriA-AriB complex (Form II)
Method: single particle / : Deep A, Corbett KD

EMDB-42968: 
CryoEM structure of AriA-AriB complex (Form III)
Method: single particle / : Deep A, Corbett KD

EMDB-42969: 
CryoEM structure of AriA (E393Q) sensory subunit
Method: single particle / : Deep A, Corbett KD

EMDB-44350: 
Synaptic Vesicle V-ATPase with synaptophysin and SidK, State 3, Vo
Method: single particle / : Coupland CE, Rubinstein JL

EMDB-44352: 
Synaptic Vesicle V-ATPase with synaptophysin and SidK, State 3, peripheral stalks
Method: single particle / : Coupland CE, Rubinstein JL

EMDB-44353: 
Synaptic Vesicle V-ATPase with synaptophysin and SidK, State 3
Method: single particle / : Coupland CE, Rubinstein JL

EMDB-44354: 
Synaptic Vesicle V-ATPase with synaptophysin and SidK, State 2
Method: single particle / : Coupland CE, Rubinstein JL

EMDB-44355: 
Synaptic Vesicle V-ATPase with synaptophysin and SidK, State 1
Method: single particle / : Coupland CE, Rubinstein JL

PDB-9b8o: 
Synaptic Vesicle V-ATPase with synaptophysin and SidK, State 3, Vo
Method: single particle / : Coupland CE, Rubinstein JL

PDB-9b8q: 
Synaptic Vesicle V-ATPase with synaptophysin and SidK, State 3, peripheral stalks
Method: single particle / : Coupland CE, Rubinstein JL

PDB-9brb: 
Synaptic Vesicle V-ATPase with synaptophysin and SidK, State 1
Method: single particle / : Coupland CE, Rubinstein JL

PDB-9brc: 
Synaptic Vesicle V-ATPase with synaptophysin and SidK, State 2
Method: single particle / : Coupland CE, Rubinstein JL

PDB-9brd: 
Synaptic Vesicle V-ATPase with synaptophysin and SidK, State 3
Method: single particle / : Coupland CE, Rubinstein JL

EMDB-39858: 
Cryo-EM structure of the insect olfactory receptor OR5-Orco heterocomplex from Acyrthosiphon pisum bound with geranyl acetate
Method: single particle / : Wang YD, Qiu L, Guan ZY, Wang Q, Wang GR, Yin P
EMDB-39873: 
Cryo-EM structure of the insect olfactory receptor OR5-Orco heterocomplex from Acyrthosiphon pisum
Method: single particle / : Wang YD, Qiu L, Guan ZY, Wang Q, Wang GR, Yin P

PDB-8z9a: 
Cryo-EM structure of the insect olfactory receptor OR5-Orco heterocomplex from Acyrthosiphon pisum bound with geranyl acetate
Method: single particle / : Wang YD, Qiu L, Guan ZY, Wang Q, Wang GR, Yin P
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search





 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
