-Search query
-Search result
Showing 1 - 50 of 208 items for (author: kikkawa & m)

EMDB-38291: 
Cryo-EM structure of human XKR8-basigin complex in lipid nanodisc
Method: single particle / : Sakuragi TS, Kanai RK, Kikkawa MK, Toyoshima CT, Nagata SN

PDB-8xej: 
Cryo-EM structure of human XKR8-basigin complex in lipid nanodisc
Method: single particle / : Sakuragi TS, Kanai RK, Kikkawa MK, Toyoshima CT, Nagata SN

EMDB-36048: 
Cryo-EM structure of hZnT7-Fab complex in zinc-unbound state, determined in outward-facing conformation
Method: single particle / : Han BB, Inaba K, Watanabe S

EMDB-36049: 
Cryo-EM structure of hZnT7-Fab complex in zinc-bound state, determined in outward-facing conformation
Method: single particle / : Han BB, Inaba K, Watanabe S

EMDB-36050: 
Cryo-EM structure of hZnT7-Fab complex in zinc-unbound state, determined in heterogeneous conformations- one subunit in an inward-facing and the other in an outward-facing conformation
Method: single particle / : Han BB, Inaba K, Watanabe S

EMDB-36051: 
Cryo-EM structure of hZnT7-Fab complex in zinc state 2, determined in heterogeneous conformations- one subunit in an inward-facing zinc-bound and the other in an outward-facing zinc-bound conformation
Method: single particle / : Han BB, Inaba K, Watanabe S

EMDB-36052: 
Cryo-EM structure of hZnT7DeltaHis-loop-Fab complex in zinc-unbound state, determined in outward-facing conformation
Method: single particle / : Han BB, Inaba K, Watanabe S

EMDB-36053: 
Cryo-EM structure of hZnT7DeltaHis-loop-Fab complex in zinc-bound state, determined in outward-facing conformation
Method: single particle / : Han BB, Inaba K, Watanabe S

EMDB-36055: 
Cryo-EM structure of hZnT7-Fab complex in zinc state 1, determined in heterogeneous conformations- one subunit in an inward-facing zinc-bound and the other in an outward-facing zinc-unbound conformation
Method: single particle / : Han BB, Inaba K, Watanabe S

PDB-8j7t: 
Cryo-EM structure of hZnT7-Fab complex in zinc-unbound state, determined in outward-facing conformation
Method: single particle / : Han BB, Inaba K, Watanabe S

PDB-8j7u: 
Cryo-EM structure of hZnT7-Fab complex in zinc-bound state, determined in outward-facing conformation
Method: single particle / : Han BB, Inaba K, Watanabe S

PDB-8j7v: 
Cryo-EM structure of hZnT7-Fab complex in zinc-unbound state, determined in heterogeneous conformations- one subunit in an inward-facing and the other in an outward-facing conformation
Method: single particle / : Han BB, Inaba K, Watanabe S

PDB-8j7w: 
Cryo-EM structure of hZnT7-Fab complex in zinc state 2, determined in heterogeneous conformations- one subunit in an inward-facing zinc-bound and the other in an outward-facing zinc-bound conformation
Method: single particle / : Han BB, Inaba K, Watanabe S

PDB-8j7x: 
Cryo-EM structure of hZnT7DeltaHis-loop-Fab complex in zinc-unbound state, determined in outward-facing conformation
Method: single particle / : Han BB, Inaba K, Watanabe S

PDB-8j7y: 
Cryo-EM structure of hZnT7DeltaHis-loop-Fab complex in zinc-bound state, determined in outward-facing conformation
Method: single particle / : Han BB, Inaba K, Watanabe S

PDB-8j80: 
Cryo-EM structure of hZnT7-Fab complex in zinc state 1, determined in heterogeneous conformations- one subunit in an inward-facing zinc-bound and the other in an outward-facing zinc-unbound conformation
Method: single particle / : Han BB, Inaba K, Watanabe S

EMDB-36340: 
Porcine uroplakin complex
Method: single particle / : Oda T, Yanagisawa H, Kikkawa M

PDB-8jj5: 
Porcine uroplakin complex
Method: single particle / : Oda T, Yanagisawa H, Kikkawa M

EMDB-34791: 
Doublet microtubule of zebrafish sperm axoneme, WT
Method: subtomogram averaging / : Yamaguchi H, Kikkawa M
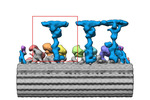

EMDB-34792: 
Radial spoke 1 of zebrafish sperm axoneme, WT
Method: subtomogram averaging / : Yamaguchi H, Kikkawa M

EMDB-34793: 
Radial spoke 2 of zebrafish sperm axoneme, WT
Method: subtomogram averaging / : Yamaguchi H, Kikkawa M

EMDB-34794: 
Radial spoke 3 of zebrafish sperm axoneme, WT
Method: subtomogram averaging / : Yamaguchi H, Kikkawa M
EMDB-34795: 
Doublet microtubule of zebrafish sperm axoneme, calaxin-/-, OAD+ class
Method: subtomogram averaging / : Yamaguchi H, Kikkawa M
EMDB-34796: 
Doublet microtubule of zebrafish sperm axoneme, calaxin-/-, OAD- class
Method: subtomogram averaging / : Yamaguchi H, Kikkawa M
EMDB-34797: 
Doublet microtubule of zebrafish sperm axoneme, armc4-/-
Method: subtomogram averaging / : Yamaguchi H, Kikkawa M

EMDB-34798: 
Outer arm dynein of zebrafish sperm axoneme, WT
Method: subtomogram averaging / : Yamaguchi H, Kikkawa M
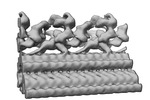

EMDB-34799: 
Outer arm dynein of zebrafish sperm axoneme, WT, refined focusing on the docking complex
Method: subtomogram averaging / : Yamaguchi H, Kikkawa M

EMDB-34800: 
Outer arm dynein of zebrafish sperm axoneme, WT, 1 mM calcium condition, refined focusing on the docking complex
Method: subtomogram averaging / : Yamaguchi H, Kikkawa M
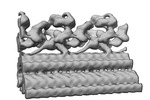

EMDB-34801: 
Outer arm dynein of zebrafish sperm axoneme, calaxin-/-, refined focusing on the docking complex
Method: subtomogram averaging / : Yamaguchi H, Kikkawa M

EMDB-34802: 
Outer arm dynein of zebrafish sperm axoneme, calaxin-/-, incubated with recombinant Calaxin protein, refined focusing on the docking complex
Method: subtomogram averaging / : Yamaguchi H, Kikkawa M

EMDB-33711: 
CryoEM structure of SPCA1a in E1-Ca-AMPPCP state subclass 1
Method: single particle / : Chen Z, Watanabe S, Inaba K

EMDB-33712: 
CryoEM structure of SPCA1a in E1-Ca-AMPPCP state subclass 2
Method: single particle / : Chen Z, Watanabe S, Inaba K

EMDB-33713: 
CryoEM structure of SPCA1a in E1-Ca-AMPPCP state subclass 3
Method: single particle / : Chen Z, Watanabe S, Inaba K

EMDB-33714: 
CryoEM structure of SPCA1a in E1-Mn-AMPPCP state subclass 1
Method: single particle / : Chen Z, Watanabe S, Inaba K

EMDB-33717: 
CryoEM structure of SPCA1a in E2P state
Method: single particle / : Chen Z, Watanabe S, Inaba K

PDB-7yag: 
CryoEM structure of SPCA1a in E1-Ca-AMPPCP state subclass 1
Method: single particle / : Chen Z, Watanabe S, Inaba K

PDB-7yah: 
CryoEM structure of SPCA1a in E1-Ca-AMPPCP state subclass 2
Method: single particle / : Chen Z, Watanabe S, Inaba K

PDB-7yai: 
CryoEM structure of SPCA1a in E1-Ca-AMPPCP state subclass 3
Method: single particle / : Chen Z, Watanabe S, Inaba K

PDB-7yaj: 
CryoEM structure of SPCA1a in E1-Mn-AMPPCP state subclass 1
Method: single particle / : Chen Z, Watanabe S, Inaba K

PDB-7yam: 
CryoEM structure of SPCA1a in E2P state
Method: single particle / : Chen Z, Watanabe S, Inaba K

EMDB-32019: 
Cryo-EM structure of SAM-Tom40 intermediate complex
Method: single particle / : Takeda H, Tsutsumi A, Kikkawa M, Endo T

PDB-7vku: 
Cryo-EM structure of SAM-Tom40 intermediate complex
Method: single particle / : Takeda H, Tsutsumi A, Kikkawa M, Endo T

EMDB-32347: 
The E1-BeF3- 2Ca2+ of SERCA2b
Method: single particle / : Zhang Y, Watanabe S, Tsutsumi A, Inaba K

EMDB-32348: 
The 'Ca2+-unbound' BeF3- of SERCA2b
Method: single particle / : Zhang Y, Watanabe S, Tsutsumi A, Inaba K

EMDB-32349: 
'late' E2P of SERCA2b
Method: single particle / : Zhang Y, Watanabe S, Tsutsumi A, Inaba K

EMDB-32350: 
E2 Pi of SERCA2b
Method: single particle / : Zhang Y, Watanabe S, Tsutsumi A, Inaba K

PDB-7w7t: 
The E1-BeF3- 2Ca2+ of SERCA2b
Method: single particle / : Zhang Y, Watanabe S, Tsutsumi A, Inaba K

PDB-7w7u: 
The 'Ca2+-unbound' BeF3- of SERCA2b
Method: single particle / : Zhang Y, Watanabe S, Tsutsumi A, Inaba K

PDB-7w7v: 
'late' E2P of SERCA2b
Method: single particle / : Zhang Y, Watanabe S, Tsutsumi A, Inaba K

PDB-7w7w: 
E2 Pi of SERCA2b
Method: single particle / : Zhang Y, Watanabe S, Tsutsumi A, Inaba K
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
