-Search query
-Search result
Showing 1 - 50 of 51 items for (author: job & v)

EMDB-18381: 
UFL1 E3 ligase bound 60S ribosome
Method: single particle / : Makhlouf L, Zeqiraj E, Kulathu Y

EMDB-18382: 
UFL1 E3 ligase bound 60S ribosome
Method: single particle / : Makhlouf L, Kulathu Y, Zeqiraj E

PDB-8qfc: 
UFL1 E3 ligase bound 60S ribosome
Method: single particle / : Makhlouf L, Zeqiraj E, Kulathu Y

PDB-8qfd: 
UFL1 E3 ligase bound 60S ribosome
Method: single particle / : Makhlouf L, Kulathu Y, Zeqiraj E

EMDB-36137: 
Cryo-EM structure of Holo form of ScBfr with O symmetry
Method: single particle / : Jobichen C, Sivaraman J

EMDB-36139: 
Cryo-EM structure of Holo form of ScBfr in C1 symmetry
Method: single particle / : Jobichen C, Sivaraman J

EMDB-36140: 
Cryo-EM structure of Fe-biomineral from bacterioferritin
Method: single particle / : Jobichen C, Sivaraman J

PDB-8jax: 
Cryo-EM structure of Holo form of ScBfr with O symmetry
Method: single particle / : Jobichen C, Sivaraman J

PDB-8jb0: 
Cryo-EM structure of Holo form of ScBfr in C1 symmetry
Method: single particle / : Jobichen C, Sivaraman J

EMDB-33639: 
Cryo-EM structure of Apo form of ScBfr
Method: single particle / : Jobichen C, Sivaraman J

EMDB-33640: 
Cryo-EM structure of bacterioferritin holoform 1a
Method: single particle / : Jobichen C, Sivaraman J

EMDB-33645: 
Cryo-EM structure if bacterioferritin holoform
Method: single particle / : Jobichen C, Sivaraman J

PDB-7y6f: 
Cryo-EM structure of Apo form of ScBfr
Method: single particle / : Jobichen C, Sivaraman J

PDB-7y6g: 
Cryo-EM structure of bacterioferritin holoform 1a
Method: single particle / : Jobichen C, Sivaraman J

PDB-7y6p: 
Cryo-EM structure if bacterioferritin holoform
Method: single particle / : Jobichen C, Sivaraman J

EMDB-35191: 
Lb2Cas12a RNA DNA complex
Method: single particle / : Li J, Sivaraman J, Satoru M

PDB-8i54: 
Lb2Cas12a RNA DNA complex
Method: single particle / : Li J, Sivaraman J, Satoru M

EMDB-25448: 
Negative-stain EM reconstruction of SpFN_1B-06-PL, a SARS-CoV-2 spike fused to H.pylori ferritin nanoparticle vaccine candidate
Method: single particle / : Thomas PV, Smith C, Chen WH, Sankhala RS, Hajduczki A, Choe M, Martinez E, Chang W, Peterson CE, Karch C, Gohain N, Kannadka CB, de Val N, Joyce MG, Modjarrad K
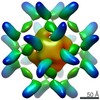
EMDB-25449: 
RFN_131, a Ferritin-based Nanoparticle Vaccine Candidate Displaying the SARS-CoV-2 Receptor-Binding Domain
Method: single particle / : Thomas PV, Smith C, Chen WH, Sankhala RS, Hajduczki A, Choe M, Martinez E, Chang W, Peterson CE, Karch C, Gohain N, Kannadka CB, de Val N, Joyce MG, Modjarrad K

EMDB-25450: 
pCoV146, a Ferritin-based Nanoparticle Vaccine Candidate Displaying the SARS-CoV-2 Spike Receptor-Binding and N-Terminal Domains
Method: single particle / : Thomas PV, Smith C, Chen WH, Sankhala RS, Hajduczki A, Choe M, Martinez E, Chang W, Peterson CE, Karch C, Gohain N, Kannadka CB, de Val N, Joyce MG, Modjarrad K

EMDB-25451: 
pCoV111, a Ferritin-based Nanoparticle Vaccine Candidate Displaying the SARS-CoV-2 Spike S1 Subunit
Method: single particle / : Thomas PV, Smith C, Chen WH, Sankhala RS, Hajduczki A, Choe M, Martinez E, Chang W, Peterson CE, Karch C, Gohain N, Kannadka CB, de Val N, Joyce MG, Modjarrad K

EMDB-11275: 
MreC
Method: helical / : Estrozi LF, Contreras-Martel C

PDB-6zlv: 
MreC
Method: helical / : Estrozi LF, Contreras-Martel C

EMDB-9910: 
Cryo-EM structure of Apo-bacterioferritin from Streptomyces coelicolor
Method: single particle / : Jobichen C, Sivaraman J

EMDB-9913: 
Cryo-EM structure of Holo-bacterioferritin-form-I from Streptomyces coelicolor
Method: single particle / : Jobichen C, Sivaraman J

EMDB-9915: 
Cryo-EM structure of Holo-bacterioferritin form-II from Streptomyces coelicolor
Method: single particle / : Jobichen C, Sivaraman J

PDB-6k3o: 
Cryo-EM structure of Apo-bacterioferritin from Streptomyces coelicolor
Method: single particle / : Jobichen C, Sivaraman J

PDB-6k43: 
Cryo-EM structure of Holo-bacterioferritin-form-I from Streptomyces coelicolor
Method: single particle / : Jobichen C, Sivaraman J

PDB-6k4m: 
Cryo-EM structure of Holo-bacterioferritin form-II from Streptomyces coelicolor
Method: single particle / : Jobichen C, Sivaraman J

EMDB-0142: 
Cryo-EM map of in vitro assembled Measles virus N into nucleocapsid-like particles (NCLPs) bound to viral genomic 5-prime RNA hexamers.
Method: helical / : Desfosses A, Milles S, Ringkjobing Jensen M, Guseva S, Colletier JP, Maurin D, Schoehn G, Gutsche I, Ruigrok R, Blackledge M

PDB-6h5s: 
Cryo-EM map of in vitro assembled Measles virus N into nucleocapsid-like particles (NCLPs) bound to viral genomic 5-prime RNA hexamers.
Method: helical / : Desfosses A, Milles S, Ringkjobing Jensen M, Guseva S, Colletier JP, Maurin D, Schoehn G, Gutsche I, Ruigrok R, Blackledge M

EMDB-0326: 
Aeromonas hydrophila ExeD
Method: single particle / : Contreras-Martel C, Farias Estrozi L

EMDB-0327: 
Vibrio vulnificus EpsD
Method: single particle / : Contreras-Martel C, Farias Estrozi L

PDB-6i1x: 
Aeromonas hydrophila ExeD
Method: single particle / : Contreras-Martel C, Farias Estrozi L

PDB-6i1y: 
Vibrio vulnificus EpsD
Method: single particle / : Contreras-Martel C, Farias Estrozi L

EMDB-0141: 
Cryo-EM structure of in vitro assembled Measles virus N into nucleocapsid-like particles (NCLPs) bound to polyA RNA hexamers.
Method: helical / : Desfosses A, Milles S, Ringkjobing Jensen M, Guseva S, Colletier J, Maurin D, Schoehn G, Gutsche I, Ruigrok R, Blackledge M

PDB-6h5q: 
Cryo-EM structure of in vitro assembled Measles virus N into nucleocapsid-like particles (NCLPs) bound to polyA RNA hexamers.
Method: helical / : Desfosses A, Milles S, Ringkjobing Jensen M, Guseva S, Colletier J, Maurin D, Schoehn G, Gutsche I, Ruigrok R, Blackledge M

EMDB-6482: 
Cryo-electron microscopy of alpha Synuclein amyloid fibrils
Method: helical / : Dearborn AD, Wall JS, Cheng N, Heymann JB, Kajava AV, Varkey J, Langen R, Steven AC

EMDB-2628: 
Structural similarity of secretins from Type II and Type III Secretion Systems
Method: single particle / : Tosi T, Estrozi LF, Job V, Guilvout I, Pugsley AP, Schoehn G, Dessen A

EMDB-2629: 
Structural similarity of secretins from Type II and Type III Secretion Systems
Method: single particle / : Tosi T, Estrozi LF, Job V, Guilvout I, Pugsley AP, Schoehn G, Dessen A

EMDB-5936: 
Electron cryo-microscopy of the Moloney murine leukemia virus furin precursor Env in its native form
Method: single particle / : Sjoberg M, Wu SR, Loving R, Rantalainen K, Lindqvist B, Garoff H

EMDB-5937: 
Electron cryo-microscopy of the Moloney murine leukemia virus furin precursor Env in its isomerization arrested state (IAS) form
Method: single particle / : Sjoberg M, Wu SR, Loving R, Rantalainen K, Lindqvist B, Garoff H

EMDB-5938: 
Electron cryo-microscopy of the Moloney murine leukemia virus furin precursor Env in its native form in complex with 83A25 Fab
Method: single particle / : Sjoberg M, Wu SR, Loving R, Rantalainen K, Lindqvist B, Garoff H

EMDB-5939: 
Electron cryo-microscopy of the partially glycosylated gp80 form of the wild type furin precursor Env of Moloney murine leukemia virus
Method: single particle / : Sjoberg M, Wu SR, Loving R, Rantalainen K, Lindqvist B, Garoff H

EMDB-2065: 
Electron cryo-microscopy of R-peptide precursor of Moloney murine leukemia virus Env in its isomerization arrested intermediate state
Method: single particle / : Loving R, Wu SR, Sjoberg M, Lindqvist B, Garoff H

EMDB-2066: 
Electron cryo-microscopy of the L651A mutant R-peptide precursor Env of Moloney murine leukemia virus in its native state
Method: single particle / : Loving R, Wu SR, Sjoberg M, Lindqvist B, Garoff H

EMDB-2067: 
Electron cryo-microscopy of R-peptide precursor of Moloney murine leukemia virus Env in its native form
Method: single particle / : Loving R, Wu SR, Sjoberg M, Lindqvist B, Garoff H

EMDB-1800: 
Single particle cryo-electron microscopy analysis of CD4 bound HIV-1 Env
Method: single particle / : Wu SR, Loving R, Lindqvist B, Hebert H, Koeck P, Sjoberg M, Garoff H

EMDB-1801: 
Single particle cryo-electron microscopy analysis of unliganded HIV-1 Env
Method: single particle / : Wu SR, Loving R, Lindqvist B, Hebert H, Koeck P, Sjoberg M, Garoff H

EMDB-2863: 
Surface rendered vitrified native Env
Method: single particle / : Wu SR, Sjoberg M, Wallin M, Lindqvist B, Ekstrom M, Hebert H, Koeck P, Garoff H
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
