-Search query
-Search result
Showing 1 - 50 of 79 items for (author: jayati & sengupta)
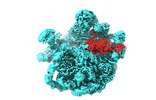
EMDB-37007: 
Mycobacterium smegmatis 50S ribosomal subunit-HflX complex
Method: single particle / : Srinivasan K, Banerjee A, Sengupta J
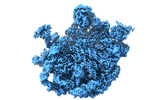
EMDB-38788: 
Mycobacterium smegmatis 50S ribosomal subunit with Erythromycin
Method: single particle / : Srinivasan K, Banerjee A, Sengupta J

PDB-8kab: 
Mycobacterium smegmatis 50S ribosomal subunit-HflX complex
Method: single particle / : Srinivasan K, Banerjee A, Sengupta J

PDB-8xz3: 
Mycobacterium smegmatis 50S ribosomal subunit with Erythromycin
Method: single particle / : Srinivasan K, Banerjee A, Sengupta J

EMDB-33096: 
Mycobacterium smegmatis 50S ribosomal subunit from Stationary phase of growth
Method: single particle / : Sengupta J, Baid P

EMDB-33599: 
Mycobacterium smegmatis 50S ribosomal subunit from Log Phase of growth
Method: single particle / : Sengupta J, Baid P

PDB-7xam: 
Mycobacterium smegmatis 50S ribosomal subunit from Stationary phase of growth
Method: single particle / : Sengupta J, Baid P

PDB-7y41: 
Mycobacterium smegmatis 50S ribosomal subunit from Log Phase of growth
Method: single particle / : Sengupta J, Baid P

EMDB-31004: 
Heme arrested off-pathway oligomer of alpha-synuclein
Method: single particle / : Dey S, Sil P, Paul S

EMDB-31023: 
on-pathway intermediate oligomer of alpha-synuclein
Method: single particle / : Dey S

EMDB-31024: 
an arrested off-pathway oligomer of alpha-synuclein upon heme treatment after fibril formation
Method: single particle / : Dey S

EMDB-30598: 
Cryo-EM structure of 70S ribosome in complex with peptide deformylase and trigger factor
Method: single particle / : Akbar S, Bhakta S, Sengupta J

EMDB-30611: 
Cryo-EM map of 70S ribosome in complex with peptide deformylase, trigger factor, and methionine aminopeptidase
Method: single particle / : Akbar S, Bhakta S

PDB-7d6z: 
Molecular model of the cryo-EM structure of 70S ribosome in complex with peptide deformylase and trigger factor
Method: single particle / : Akbar S, Bhakta S, Sengupta J

PDB-7d80: 
Molecular model of the cryo-EM structure of 70S ribosome in complex with peptide deformylase, trigger factor, and methionine aminopeptidase
Method: single particle / : Akbar S, Bhakta S, Sengupta J
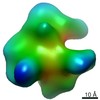
EMDB-9878: 
Cryo EM density map of Resveratrol-stabilized bioactive insulin oligomer
Method: single particle / : Sengupta J, Pathak BK

PDB-6jr3: 
Crystal structure of insulin hexamer fitted into cryo EM density map where each dimer was kept as rigid body
Method: single particle / : Sengupta J, Pathak BK, Bhakta S

PDB-6k4n: 
Cryo-EM structure of p300
Method: single particle / : Ghosh R, Roy S, Sengupta J

EMDB-9750: 
Cryo-EM density map of E. coli 70S ribosome in complex with peptide deformylase enzyme
Method: single particle / : Sengupta J, Akbar S

EMDB-9752: 
Cryo-EM density map of E. coli 70S ribosome in complex with methionine aminopeptidase enzyme
Method: single particle / : Sengupta J, Bhakta S

EMDB-9753: 
Cryo-EM density map of peptide deformylase and methionine aminopeptidase bound to the E. coli 70S ribosome
Method: single particle / : Sengupta J, Bhakta S

EMDB-9759: 
Cryo-EM density map of methionine aminopeptidase enzyme and chaperone trigger factor bound to the E. coli 70S ribosome
Method: single particle / : Sengupta J, Bhakta S

EMDB-9778: 
Cryo-EM density map of peptide deformylase enzyme and chaperone trigger factor bound to the E. coli 70S ribosome
Method: single particle / : Sengupta J, Bhakta S

PDB-6iy7: 
E. coli peptide deformylase crystal structure fitted into the cryo-EM density map of E. coli 70S ribosome in complex with peptide deformylase
Method: single particle / : Sengupta J, Akbar S, Bhakta S

PDB-6iz7: 
E. coli methionine aminopeptidase crystal structure fitted into the cryo-EM density map of E. coli 70S ribosome in complex with methionine aminopeptidase
Method: single particle / : Sengupta J, Bhakta S, Akbar S

PDB-6izi: 
Crystal structure of E. coli peptide deformylase and methionine aminopeptidase fitted into the cryo-EM density map of the complex
Method: single particle / : Sengupta J, Bhakta S, Akbar S

PDB-6j0a: 
Crystal structure of E. coli methionine aminopeptidase enzyme and chaperone trigger factor fitted into the cryo-EM density map of the complex
Method: single particle / : Sengupta J, Bhakta S, Akbar S

PDB-6j45: 
Crystal structure of E. coli peptide deformylase enzyme and chaperone trigger factor fitted into the cryo-EM density map of the complex
Method: single particle / : Sengupta J, Bhakta S, Akbar S

EMDB-6791: 
Cryo-EM structure of p300-p53 protein complex
Method: single particle / : Ghosh R, Roy S, Sengupta J

EMDB-6792: 
Cryo-EM structure of p300
Method: single particle / : Ghosh R, Roy S, Sengupta J

PDB-5xzc: 
Cryo-EM structure of p300-p53 protein complex
Method: single particle / : Ghosh R, Roy S, Sengupta J

EMDB-6979: 
E. coli 50S subunit bound HflX protein in presence of ATP (AMP-PNP) and GTP (GMP-PNP) analogs.
Method: single particle / : Dey S

PDB-5zzm: 
E. coli 50S subunit bound HflX protein in presence of ATP (AMP-PNP) and GTP (GMP-PNP) analogs.
Method: single particle / : Dey S

EMDB-2970: 
Cryo-EM structure of E. coli 70S ribosome bound to additional non-ribosomal proteins.
Method: single particle / : Shasmal M, Dey S, Shaikh TR, Bhakta S, Sengupta J

EMDB-2972: 
Cryo-EM structure of E. coli 70S ribosome bound to additional non-ribosomal proteins.
Method: single particle / : Shasmal M, Dey S, Shaikh TR, Bhakta S, Sengupta J

PDB-4v69: 
Ternary complex-bound E.coli 70S ribosome.
Method: single particle / : Villa E, Sengupta J, Trabuco LG, LeBarron J, Baxter WT, Shaikh TR, Grassucci RA, Nissen P, Ehrenberg M, Schulten K, Frank J

PDB-4v47: 
Real space refined coordinates of the 30S and 50S subunits fitted into the low resolution cryo-EM map of the EF-G.GTP state of E. coli 70S ribosome
Method: single particle / : Gao H, Sengupta J, Valle M, Korostelev A, Eswar N, Stagg SM, Van Roey P, Agrawal RK, Harvey ST, Sali A, Chapman MS, Frank J

PDB-4v48: 
Real space refined coordinates of the 30S and 50S subunits fitted into the low resolution cryo-EM map of the initiation-like state of E. coli 70S ribosome
Method: single particle / : Gao H, Sengupta J, Valle M, Korostelev A, Eswar N, Stagg SM, Van Roey P, Agrawal RK, Harvey ST, Sali A, Chapman MS, Frank J

EMDB-5307: 
Three-dimensional cryo-electron microscopy density map of the 70S ribosome from Mycobacterium smegmatis
Method: single particle / : Shasmal M, Sengupta J

PDB-3iyx: 
Coordinates of the b1b bridge-forming protein structures fitted into the Cryo-EM map of E.coli 70S ribosome (EMD-1056)
Method: single particle / : Shasmal M, Chakraborty B, Sengupta J

PDB-3iyy: 
Coordinates of the b1b bridge-forming protein structures fitted into the Cryo-EM map of EFG.GDPNP-bound E.coli 70S ribosome(EMD-1363)
Method: single particle / : Shasmal M, Chakraborty B, Sengupta J

EMDB-5016: 
15.3A eEF2-80S Ribosome Transition State Complex by Cryo-electron Microscopy
Method: single particle / : Sengupta J, Nilsson J, Gursky R, Kjeldgaard M, Nissen P, Frank J

EMDB-5036: 
Aminoacyl-tRNA-EF-Tu-GDP-kir ternary complex-bound E. coli 70S ribosome
Method: single particle / : Villa E, Sengupta J, Trabuco LG, LeBarron J, Baxter WT, Shaikh TR, Grassucci RA, Nissen P, Ehrenberg M, Schulten K, Frank J

EMDB-5015: 
A 12.6A cryo-EM map of the 80S.eEF2.AlF4-GDP complex
Method: single particle / : Sengupta J, Nilsson J, Gursky R, Kjeldgaard M, Nissen P, Frank J

EMDB-5017: 
Segmented eEF2 density from the cryo-EM map of eEF2-bound 80S complex
Method: single particle / : Sengupta J, Nilsson J, Gursky R, Kjeldgaard M, Nissen P, Frank J

PDB-3dwu: 
Transition-state model conformation of the switch I region fitted into the cryo-EM map of the eEF2.80S.AlF4.GDP complex
Method: single particle / : Nissen P, Nyborg J, Kjeldgaard M

PDB-3dny: 
Fitting of the eEF2 crystal structure into the cryo-EM density map of the eEF2.80S.AlF4-.GDP complex
Method: single particle / : Sengupta J, Frank J

EMDB-1395: 
Incorporation of aminoacyl-tRNA into the ribosome as seen by cryo-electron microscopy.
Method: single particle / : Valle M, Zavialov A, Li W, Stagg SM, Sengupta J, Nielsen RC, Nissen P, Harvey SC, Ehrenberg M, Frank J

EMDB-1362: 
Locking and unlocking of ribosomal motions.
Method: single particle / : Mikel V, Andrey Z, Sengupta J, Rawat U, Ehrenberg M, Frank J

EMDB-1363: 
Locking and unlocking of ribosomal motions.
Method: single particle / : Mikel V, Andrey Z, Sengupta J, Rawat U, Ehrenberg M, Frank J
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
