-Search query
-Search result
Showing 1 - 50 of 233 items for (author: janowsk & r)
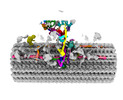
EMDB-41284: 
Combined linker domain of N-DRC and associated proteins Tetrahymena
Method: single particle / : Ghanaeian AG, Majhi SM, McCaffrey CM, Nami BN, Black CB, Yang SK, Legal TL, Papoulas OP, Janowska MJ, Valente-Paterno MV, Marcotte EM, Wloga DW, Bui KH

PDB-8tid: 
Combined linker domain of N-DRC and associated proteins Tetrahymena
Method: single particle / : Ghanaeian AG, Majhi SM, McCaffrey CM, Nami BN, Black CB, Yang SK, Legal TL, Papoulas OP, Janowska MJ, Valente-Paterno MV, Marcotte EM, Wloga DW, Bui KH

EMDB-41189: 
Baseplate of Nexin-dynein regulatory complex from Tetrahymena thermophila
Method: single particle / : Ghanaeian AG, Black CS, Yang SK, Bui KH

EMDB-41251: 
Linker domain of Nexin-dynein regulatory complex from Tetrahymena thermophila
Method: single particle / : Ghanaeian AG, Bui KH

EMDB-41270: 
Focused refinement of the N-DRC from the Tetrahymena WT subtomo
Method: subtomogram averaging / : Ghanaeian AG, Majhi SM, McCaffrey CM, Nami BN, Black CB, Yang SK, Legal TL, Papoulas OP, Janowska MJ, Valente-Paterno MV, Marcotte EM, Wloga DW, Bui KH

EMDB-41375: 
Linker domain of N-DRC complex from Tetrahymena thermophila
Method: single particle / : Ghanaeian AG, Bui KH

EMDB-41376: 
96nm repeat of Doublet microtubule from Tetrahymena thermophila
Method: single particle / : Ghanaeian AG, Bui KH

EMDB-41504: 
DRC9/10 baseplate from Tetrahymena thermophila
Method: single particle / : Ghanaeian AG, Bui KH

PDB-8tek: 
Baseplate of Nexin-dynein regulatory complex from Tetrahymena thermophila
Method: single particle / : Ghanaeian AG, Black CS, Yang SK, Bui KH

PDB-8th8: 
Linker domain of Nexin-dynein regulatory complex from Tetrahymena thermophila
Method: single particle / : Ghanaeian AG, Bui KH
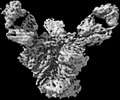
EMDB-27624: 
Cryo-EM structure of HIV-1 Env(CH848 10.17 DS.SOSIP_DT) in complex with DH1030.1 Fab
Method: single particle / : Gobeil S, Acharya P
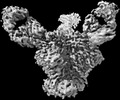
EMDB-27628: 
Cryo-EM structure of HIV-1 Env(BG505.T332N SOSIP) in complex with DH1030.1 Fab
Method: single particle / : Gobeil S, Acharya P

EMDB-29029: 
Langya Virus Fusion Protein (LayV-F) in Pre-Fusion Conformation
Method: single particle / : May AJ, Pothula KR, Janowska K, Acharya P

EMDB-29032: 
Langya Virus Fusion Protein (LayV-F) in Post-Fusion Conformation
Method: single particle / : May AJ, Pothula KR, Janowska K, Acharya P

PDB-8fej: 
Langya Virus Fusion Protein (LayV-F) in Pre-Fusion Conformation
Method: single particle / : May AJ, Pothula KR, Janowska K, Acharya P

PDB-8fel: 
Langya Virus Fusion Protein (LayV-F) in Post-Fusion Conformation
Method: single particle / : May AJ, Pothula KR, Janowska K, Acharya P

EMDB-28608: 
Cryo-EM structure of CH848 10.17DT DS-SOSIP-2P Env
Method: single particle / : Wrapp D, Acharya P, Haynes BF

PDB-8eu8: 
Cryo-EM structure of CH848 10.17DT DS-SOSIP-2P Env
Method: single particle / : Wrapp D, Acharya P, Haynes BF

EMDB-26961: 
Triple mutant (K417N-E484K-N501Y) SARS-CoV-2 spike protein in the 3-RBD-Down conformation (S-GSAS-D614G-K417N-E484K-N501Y)
Method: single particle / : Gobeil S, Acharya P

EMDB-26433: 
SARS-CoV-2 Omicron-BA.2 3-RBD down Spike Protein Trimer without the P986-P987 stabilizing mutations (S-GSAS-Omicron-BA.2)
Method: single particle / : Stalls V, Acharya P
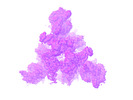
EMDB-26435: 
SARS-CoV-2 Omicron-BA.2 3-RBD down Spike Protein Trimer without the P986-P987 stabilizing mutations (S-GSAS-Omicron-BA.2)
Method: single particle / : Stalls V, Acharya P

EMDB-26436: 
SARS-CoV-2 Omicron-BA.2 3-RBD down Spike Protein Trimer without the P986-P987 stabilizing mutations (S-GSAS-Omicron-BA.2)
Method: single particle / : Stalls V, Acharya P

EMDB-26643: 
SARS-CoV-2 Omicron-BA.2 1.5-RBD up Spike Protein Trimer without the P986-P987 stabilizing mutations (S-GSAS-Omicron-BA.2)
Method: single particle / : Stalls V, Acharya P

EMDB-26644: 
SARS-CoV-2 Omicron-BA.2 1-RBD-up Spike Protein Trimer without the P986-P987 stabilizing mutations (S-GSAS-Omicron-BA.2)
Method: single particle / : Stalls V, Acharya P

EMDB-26647: 
SARS-CoV-2 Omicron-BA.2 1-RBD-up Spike Protein Trimer without the P986-P987 stabilizing mutations (S-GSAS-Omicron-BA.2)
Method: single particle / : Stalls V, Acharya P

PDB-7ub0: 
SARS-CoV-2 Omicron-BA.2 3-RBD down Spike Protein Trimer without the P986-P987 stabilizing mutations (S-GSAS-Omicron-BA.2)
Method: single particle / : Stalls V, Acharya P

PDB-7ub5: 
SARS-CoV-2 Omicron-BA.2 3-RBD down Spike Protein Trimer without the P986-P987 stabilizing mutations (S-GSAS-Omicron-BA.2)
Method: single particle / : Stalls V, Acharya P

PDB-7ub6: 
SARS-CoV-2 Omicron-BA.2 3-RBD down Spike Protein Trimer without the P986-P987 stabilizing mutations (S-GSAS-Omicron-BA.2)
Method: single particle / : Stalls V, Acharya P
EMDB-26600: 
SARS-CoV-2 Omicron-BA.1 2-RBD up Spike Protein Trimer without the P986-P987 stabilizing mutations (S-GSAS-Omicron-BA.1)
Method: single particle / : Stalls V, Acharya P

EMDB-25880: 
SARS-CoV-2 Omicron 1-RBD down Spike Protein Trimer without the P986-P987 stabilizing mutations (S-GSAS-Omicron)
Method: single particle / : Stalls V, Acharya P

PDB-7tge: 
SARS-CoV-2 Omicron 1-RBD down Spike Protein Trimer without the P986-P987 stabilizing mutations (S-GSAS-Omicron)
Method: single particle / : Stalls V, Acharya P

EMDB-25983: 
SARS-CoV-2 Omicron 3-RBD down Spike Protein Trimer without the P986-P987 stabilizing mutations (S-GSAS-Omicron)
Method: single particle / : Stalls V, Acharya P

EMDB-25984: 
SARS-CoV-2 Omicron 1-RBD up Spike Protein Trimer without the P986-P987 stabilizing mutations (S-GSAS-Omicron)
Method: single particle / : Stalls V, Acharya P

PDB-7tl1: 
SARS-CoV-2 Omicron 3-RBD down Spike Protein Trimer without the P986-P987 stabilizing mutations (S-GSAS-Omicron)
Method: single particle / : Stalls V, Acharya P

PDB-7tl9: 
SARS-CoV-2 Omicron 1-RBD up Spike Protein Trimer without the P986-P987 stabilizing mutations (S-GSAS-Omicron)
Method: single particle / : Stalls V, Acharya P

EMDB-25846: 
SARS-CoV-2 Omicron 1-RBD up Spike Protein Trimer without the P986-P987 stabilizing mutations (S-GSAS-Omicron)
Method: single particle / : Stalls V, Acharya P

EMDB-25865: 
SARS-CoV-2 Omicron 3-RBD down Spike Protein Trimer without the P986-P987 stabilizing mutations (S-GSAS-Omicron)
Method: single particle / : Stalls V, Acharya P

EMDB-25904: 
CryoEM map of SARS-CoV-2 S protein in complex with Receptor Binding Domain antibody DH1042
Method: single particle / : Manne K, May A, Acharya P

PDB-7tei: 
SARS-CoV-2 Omicron 1-RBD up Spike Protein Trimer without the P986-P987 stabilizing mutations (S-GSAS-Omicron)
Method: single particle / : Stalls V, Acharya P

PDB-7tf8: 
SARS-CoV-2 Omicron 3-RBD down Spike Protein Trimer without the P986-P987 stabilizing mutations (S-GSAS-Omicron)
Method: single particle / : Stalls V, Acharya P

PDB-7tht: 
CryoEM structure of SARS-CoV-2 S protein in complex with Receptor Binding Domain antibody DH1042
Method: single particle / : Manne K, May A, Acharya P

EMDB-25893: 
Structure of RBD directed antibody DH1042 in complex with SARS-CoV-2 spike: Local refinement of RBD-Fab interface
Method: single particle / : May AJ, Manne K, Acharya P

PDB-7the: 
Structure of RBD directed antibody DH1042 in complex with SARS-CoV-2 spike: Local refinement of RBD-Fab interface
Method: single particle / : May AJ, Manne K, Acharya P

EMDB-26038: 
Delta (B.1.617.2) SARS-CoV-2 variant spike protein (S-GSAS-Delta) in the 3-RBD-down conformation; consensus state D1
Method: single particle / : Gobeil S, Acharya P

EMDB-26039: 
Delta (B.1.617.2) SARS-CoV-2 variant spike protein (S-GSAS-Delta) in the 1-RBD-up conformation; consensus state D2
Method: single particle / : Gobeil S, Acharya P

EMDB-26040: 
Delta (B.1.617.2) SARS-CoV-2 variant spike protein (S-GSAS-Delta) in the 3-RBD-down conformation; Subclassification D5 state
Method: single particle / : Gobeil S, Acharya P

EMDB-26041: 
Delta (B.1.617.2) SARS-CoV-2 variant spike protein (S-GSAS-Delta) in the 3-RBD-down conformation; Subclassification D6 state
Method: single particle / : Gobeil S, Acharya P

EMDB-26042: 
Delta (B.1.617.2) SARS-CoV-2 variant spike protein (S-GSAS-Delta) in the 3-RBD-down conformation; Subclassification D7 state
Method: single particle / : Gobeil S, Acharya P

EMDB-26043: 
Delta (B.1.617.2) SARS-CoV-2 variant spike protein (S-GSAS-Delta) in the 3-RBD-down conformation; Subclassification D8 state
Method: single particle / : Gobeil S, Acharya P

EMDB-26045: 
Delta (B.1.617.2) SARS-CoV-2 variant spike protein (S-GSAS-Delta) in the 3-RBD-down conformation; Subclassification D9 state
Method: single particle / : Gobeil S, Acharya P
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
