-Search query
-Search result
Showing 1 - 50 of 201 items for (author: inoue & s)

EMDB entry, No image
EMDB-36467: 
Cryo-EM structure of the head region of full-length ERGIC-53 with MCFD2 (form A)
Method: single particle / : Watanabe S, Inaba K

EMDB entry, No image
EMDB-36468: 
Cryo-EM structures of the head region of full-length ERGIC-53 with MCFD2 (form B)
Method: single particle / : Watanabe S, Inaba K

EMDB entry, No image
EMDB-36469: 
Cryo-EM structures of the head region of full-length ERGIC-53 with MCFD2 (Substate A)
Method: single particle / : Watanabe S, Inaba K

EMDB entry, No image
EMDB-36470: 
Cryo-EM structure of the head region of full-length ERGIC-53 with MCFD2 (Substate B)
Method: single particle / : Watanabe S, Inaba K

EMDB entry, No image
EMDB-36471: 
Cryo-EM structure of the head region of full-length ERGIC-53 with MCFD2 (Substate C)
Method: single particle / : Watanabe S, Inaba K

EMDB entry, No image
EMDB-36472: 
Cryo-EM structure of the head region of full-length ERGIC-53 with MCFD2 (Substate D)
Method: single particle / : Watanabe S, Inaba K

EMDB entry, No image
EMDB-36479: 
Cryo-EM structure of full-length ERGIC-53 with MCFD2
Method: single particle / : Watanabe S, Inaba K

EMDB entry, No image
EMDB-36482: 
cryoEM structure of ERGIC-53 deltaH34 mutant with MCFD2
Method: single particle / : Watanabe S, Inaba K

PDB-8jp4: 
Cryo-EM structure of the head region of full-length ERGIC-53 with MCFD2 (form A)
Method: single particle / : Watanabe S, Inaba K

PDB-8jp5: 
Cryo-EM structures of the head region of full-length ERGIC-53 with MCFD2 (form B)
Method: single particle / : Watanabe S, Inaba K

PDB-8jp6: 
Cryo-EM structures of the head region of full-length ERGIC-53 with MCFD2 (Substate A)
Method: single particle / : Watanabe S, Inaba K

PDB-8jp7: 
Cryo-EM structure of the head region of full-length ERGIC-53 with MCFD2 (Substate B)
Method: single particle / : Watanabe S, Inaba K

PDB-8jp8: 
Cryo-EM structure of the head region of full-length ERGIC-53 with MCFD2 (Substate C)
Method: single particle / : Watanabe S, Inaba K

PDB-8jp9: 
Cryo-EM structure of the head region of full-length ERGIC-53 with MCFD2 (Substate D)
Method: single particle / : Watanabe S, Inaba K

PDB-8jpg: 
Cryo-EM structure of full-length ERGIC-53 with MCFD2
Method: single particle / : Watanabe S, Inaba K
EMDB-37881: 
GPR3/Gs complex
Method: single particle / : He Y, Xiong Y

EMDB-35234: 
Cryo-EM structure of Acipimox bound human hydroxy-carboxylic acid receptor 2 in complex with Gi heterotrimer
Method: single particle / : Park JH, Ishimoto N, Park SY

EMDB-35235: 
Cryo-EM structure of GSK256073 bound human hydroxy-carboxylic acid receptor 2 in complex with Gi heterotrimer
Method: single particle / : Park JH, Ishimoto N, Park SY

PDB-8i7v: 
Cryo-EM structure of Acipimox bound human hydroxy-carboxylic acid receptor 2 in complex with Gi heterotrimer
Method: single particle / : Park JH, Ishimoto N, Park SY

PDB-8i7w: 
Cryo-EM structure of GSK256073 bound human hydroxy-carboxylic acid receptor 2 in complex with Gi heterotrimer
Method: single particle / : Park JH, Ishimoto N, Park SY

EMDB-36900: 
Cryo-EM structure of niacin bound human hydroxy-carboxylic acid receptor 2 (Local refinement)
Method: single particle / : Park JH, Ishimoto N, Park SY

EMDB-36901: 
Cryo-EM structure of Acipimox bound human hydroxy-carboxylic acid receptor 2 (Local refinement)
Method: single particle / : Park JH, Ishimoto N, Park SY

EMDB-36902: 
Cryo-EM structure of GSK256073 bound human hydroxy-carboxylic acid receptor 2 (Local refinement)
Method: single particle / : Park JH, Ishimoto N, Park SY

EMDB-34437: 
Cryo-EM structure of niacin bound human hydroxy-carboxylic acid receptor 2 in complex with Gi heterotrimer
Method: single particle / : Park JH, Ishimoto N, Park SY

EMDB-36048: 
Cryo-EM structure of hZnT7-Fab complex in zinc-unbound state, determined in outward-facing conformation
Method: single particle / : Han BB, Inaba K, Watanabe S

EMDB-36049: 
Cryo-EM structure of hZnT7-Fab complex in zinc-bound state, determined in outward-facing conformation
Method: single particle / : Han BB, Inaba K, Watanabe S

EMDB-36050: 
Cryo-EM structure of hZnT7-Fab complex in zinc-unbound state, determined in heterogeneous conformations- one subunit in an inward-facing and the other in an outward-facing conformation
Method: single particle / : Han BB, Inaba K, Watanabe S

EMDB-36051: 
Cryo-EM structure of hZnT7-Fab complex in zinc state 2, determined in heterogeneous conformations- one subunit in an inward-facing zinc-bound and the other in an outward-facing zinc-bound conformation
Method: single particle / : Han BB, Inaba K, Watanabe S

EMDB-36052: 
Cryo-EM structure of hZnT7DeltaHis-loop-Fab complex in zinc-unbound state, determined in outward-facing conformation
Method: single particle / : Han BB, Inaba K, Watanabe S

EMDB-36053: 
Cryo-EM structure of hZnT7DeltaHis-loop-Fab complex in zinc-bound state, determined in outward-facing conformation
Method: single particle / : Han BB, Inaba K, Watanabe S

EMDB-36055: 
Cryo-EM structure of hZnT7-Fab complex in zinc state 1, determined in heterogeneous conformations- one subunit in an inward-facing zinc-bound and the other in an outward-facing zinc-unbound conformation
Method: single particle / : Han BB, Inaba K, Watanabe S

PDB-8j7t: 
Cryo-EM structure of hZnT7-Fab complex in zinc-unbound state, determined in outward-facing conformation
Method: single particle / : Han BB, Inaba K, Watanabe S

PDB-8j7u: 
Cryo-EM structure of hZnT7-Fab complex in zinc-bound state, determined in outward-facing conformation
Method: single particle / : Han BB, Inaba K, Watanabe S

PDB-8j7v: 
Cryo-EM structure of hZnT7-Fab complex in zinc-unbound state, determined in heterogeneous conformations- one subunit in an inward-facing and the other in an outward-facing conformation
Method: single particle / : Han BB, Inaba K, Watanabe S

PDB-8j7w: 
Cryo-EM structure of hZnT7-Fab complex in zinc state 2, determined in heterogeneous conformations- one subunit in an inward-facing zinc-bound and the other in an outward-facing zinc-bound conformation
Method: single particle / : Han BB, Inaba K, Watanabe S

PDB-8j7x: 
Cryo-EM structure of hZnT7DeltaHis-loop-Fab complex in zinc-unbound state, determined in outward-facing conformation
Method: single particle / : Han BB, Inaba K, Watanabe S

PDB-8j7y: 
Cryo-EM structure of hZnT7DeltaHis-loop-Fab complex in zinc-bound state, determined in outward-facing conformation
Method: single particle / : Han BB, Inaba K, Watanabe S

PDB-8j80: 
Cryo-EM structure of hZnT7-Fab complex in zinc state 1, determined in heterogeneous conformations- one subunit in an inward-facing zinc-bound and the other in an outward-facing zinc-unbound conformation
Method: single particle / : Han BB, Inaba K, Watanabe S

EMDB-34530: 
Membrane protein A
Method: single particle / : Tajima S, Kim Y, Yamashita K, Fukuda M, Deisseroth K, Kato HE

EMDB-34531: 
Membrane protein B
Method: single particle / : Tajima S, Kim Y, Yamashita K, Fukuda M, Deisseroth K, Kato HE

EMDB-35713: 
Cryo-EM structure of the potassium-selective channelrhodopsin HcKCR1 H225F mutant in lipid nanodisc
Method: single particle / : Tajima S, Kim Y, Nakamura S, Yamashita K, Fukuda M, Deisseroth K, Kato HE
EMDB-33972: 
Cryo-EM structure of the SARS-CoV-2 spike protein (3-up RBD) bound to neutralizing antibody CSW1-1805
Method: single particle / : Anzai I, Fujita J, Ono C, Kosaka Y, Miyamoto Y, Shichinohe S, Takada K, Torii S, Taguwa S, Suzuki K, Makino F, Kajita T, Inoue T, Namba K, Watanabe T, Matsuura Y
EMDB-33973: 
Cryo-EM structure of the SARS-CoV-2 spike protein (1-up RBD) bound to neutralizing antibody CSW2-1353
Method: single particle / : Anzai I, Fujita J, Ono C, Kosaka Y, Miyamoto Y, Shichinohe S, Takada K, Torii S, Taguwa S, Suzuki K, Makino F, Kajita T, Inoue T, Namba K, Watanabe T, Matsuura Y
EMDB-33974: 
Cryo-EM structure of the SARS-CoV-2 spike protein (2-up RBD) bound to neutralizing antibody CSW2-1353
Method: single particle / : Anzai I, Fujita J, Ono C, Kosaka Y, Miyamoto Y, Shichinohe S, Takada K, Torii S, Taguwa S, Suzuki K, Makino F, Kajita T, Inoue T, Namba K, Watanabe T, Matsuura Y
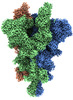
EMDB-33975: 
Cryo-EM structure of the SARS-CoV-2 spike protein (2-up RBD) with C480A mutation
Method: single particle / : Anzai I, Fujita J, Ono C, Kosaka Y, Miyamoto Y, Shichinohe S, Takada K, Torii S, Taguwa S, Suzuki K, Makino F, Kajita T, Inoue T, Namba K, Watanabe T, Matsuura Y

EMDB-34429: 
Cryo-EM structure of KpFtsZ-monobody double helical tube
Method: helical / : Fujita J, Amesaka H, Yoshizawa T, Kuroda N, Kamimura N, Hara M, Inoue T, Namba K, Tanaka S, Matsumura H

PDB-8h1o: 
Cryo-EM structure of KpFtsZ-monobody double helical tube
Method: helical / : Fujita J, Amesaka H, Yoshizawa T, Kuroda N, Kamimura N, Hara M, Inoue T, Namba K, Tanaka S, Matsumura H

EMDB-35344: 
Cryo-EM structure of KpFtsZ single filament
Method: helical / : Fujita J, Amesaka H, Yoshizawa T, Kuroda N, Kamimura N, Hibino K, Konishi T, Kato Y, Hara M, Inoue T, Namba K, Tanaka S, Matsumura H

PDB-8ibn: 
Cryo-EM structure of KpFtsZ single filament
Method: helical / : Fujita J, Amesaka H, Yoshizawa T, Kuroda N, Kamimura N, Hibino K, Konishi T, Kato Y, Hara M, Inoue T, Namba K, Tanaka S, Matsumura H
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search




 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
