-Search query
-Search result
Showing 1 - 50 of 387 items for (author: huang & rk)

EMDB-41877: 
Cryo-EM structure of long form insulin receptor (IR-B) in the apo state
Method: single particle / : An W, Hall C, Li J, Huang A, Wu J, Park J, Bai XC, Choi E

EMDB-41878: 
Cryo-EM structure of long form insulin receptor (IR-B) with four IGF2 bound, symmetric conformation.
Method: single particle / : An W, Hall C, Li J, Huang A, Wu J, Park J, Bai XC, Choi E

EMDB-41880: 
Cryo-EM structure of long form insulin receptor (IR-B) with three IGF2 bound, asymmetric conformation.
Method: single particle / : An W, Hall C, Li J, Huang A, Wu J, Park J, Bai XC, Choi E

EMDB-43279: 
Cryo-EM structure of short form insulin receptor (IR-A) with four IGF2 bound, symmetric conformation.
Method: single particle / : An W, Hall C, Li J, Huang A, Wu J, Park J, Bai XC, Choi E

EMDB-43280: 
Cryo-EM structure of short form insulin receptor (IR-A) with three IGF2 bound, asymmetric conformation.
Method: single particle / : An W, Hall C, Li J, Huang A, Wu J, Park J, Bai XC, Choi E

PDB-8u4b: 
Cryo-EM structure of long form insulin receptor (IR-B) in the apo state
Method: single particle / : An W, Hall C, Li J, Huang A, Wu J, Park J, Bai XC, Choi E

PDB-8u4c: 
Cryo-EM structure of long form insulin receptor (IR-B) with four IGF2 bound, symmetric conformation.
Method: single particle / : An W, Hall C, Li J, Huang A, Wu J, Park J, Bai XC, Choi E

PDB-8u4e: 
Cryo-EM structure of long form insulin receptor (IR-B) with three IGF2 bound, asymmetric conformation.
Method: single particle / : An W, Hall C, Li J, Huang A, Wu J, Park J, Bai XC, Choi E

PDB-8vjb: 
Cryo-EM structure of short form insulin receptor (IR-A) with four IGF2 bound, symmetric conformation.
Method: single particle / : An W, Hall C, Li J, Huang A, Wu J, Park J, Bai XC, Choi E

PDB-8vjc: 
Cryo-EM structure of short form insulin receptor (IR-A) with three IGF2 bound, asymmetric conformation.
Method: single particle / : An W, Hall C, Li J, Huang A, Wu J, Park J, Bai XC, Choi E

EMDB-43091: 
hGBP1 conformer on the bacterial outer membrane
Method: subtomogram averaging / : Zhu S, MacMicking J

EMDB-43153: 
hGBP1 conformer on the bacterial outer membrane
Method: subtomogram averaging / : Zhu S, MacMicking J

EMDB-41837: 
The structure of the PP2A-B56Delta holoenzyme mutant - E197K
Method: single particle / : Wu CG, Xing Y

PDB-8u1x: 
The structure of the PP2A-B56Delta holoenzyme mutant - E197K
Method: single particle / : Wu CG, Xing Y

EMDB-42018: 
The structure of the PP2A-B56Delta holoenzyme mutant - E197K
Method: single particle / : Wu CG, Xing Y

PDB-8u89: 
The structure of the PP2A-B56Delta holoenzyme mutant - E197K
Method: single particle / : Wu CG, Xing Y

EMDB-29783: 
Cryo-EM structure of T/F100 SOSIP.664 HIV-1 Env trimer with LMHS mutations in complex with 8ANC195 and 10-1074
Method: single particle / : Chen Y, Zhou F, Huang R, Tolbert W, Pazgier M

EMDB-41613: 
Cryo-EM structure of BG505 SOSIP.664 HIV-1 Env trimer in complex with temsavir, 8ANC195, and 10-1074
Method: single particle / : Tolbert WD, Pozharski E, Pazgier M
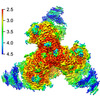
EMDB-28617: 
Cryo-EM structure of HIV-1 BG505 DS-SOSIP ENV trimer bound to VRC34.01 FAB
Method: single particle / : Pletnev S, Kwong P

EMDB-28618: 
Cryo-EM structure of HIV-1 BG505 DS-SOSIP ENV trimer bound to VRC34.01-COMBO1 FAB
Method: single particle / : Pletnev S, Kwong P

EMDB-28619: 
Cryo-EM structure of HIV-1 BG505 DS-SOSIP ENV trimer bound to VRC34.01-MM28 FAB
Method: single particle / : Pletnev S, Kwong P

PDB-8euu: 
Cryo-EM structure of HIV-1 BG505 DS-SOSIP ENV trimer bound to VRC34.01 FAB
Method: single particle / : Pletnev S, Kwong P

PDB-8euv: 
Cryo-EM structure of HIV-1 BG505 DS-SOSIP ENV trimer bound to VRC34.01-COMBO1 FAB
Method: single particle / : Pletnev S, Kwong P

PDB-8euw: 
Cryo-EM structure of HIV-1 BG505 DS-SOSIP ENV trimer bound to VRC34.01-MM28 FAB
Method: single particle / : Pletnev S, Kwong P

EMDB-28036: 
Cryo-EM structure of the full-length human NF1 dimer
Method: single particle / : Darling JE, Merk A, Grisshammer R, Ognjenovic J

PDB-8edm: 
Cryo-EM structure of the full-length human NF1 dimer
Method: single particle / : Darling JE, Merk A, Grisshammer R, Ognjenovic J

EMDB-41182: 
Cryo-EM map of the Unmodified nucleosome core particle in 100 mM KCl with local resolution values
Method: single particle / : Huang SK, Kay LE, Rubinstein JL
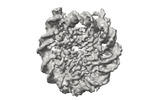
EMDB-41183: 
Cryo-EM map of the PARylated nucleosome core particle in 100 mM KCl with local resolution values
Method: single particle / : Huang SK, Kay LE, Rubinstein JL
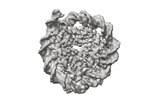
EMDB-41184: 
Cryo-EM map of the Unmodified nucleosome core particle in 5 mM KCl with local resolution values
Method: single particle / : Huang SK, Kay LE, Rubinstein JL

EMDB-27596: 
Cryo-EM structure of T/F100 SOSIP.664 HIV-1 Env trimer in complex with 8ANC195 and 10-1074
Method: single particle / : Chen Y, Zhou F, Huang R, Tolbert W, Pazgier M
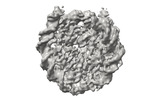
EMDB-41178: 
Cryo-EM map of the PARylated nucleosome core particle in 5 mM KCl with local resolution values
Method: single particle / : Huang SK, Kay LE, Rubinstein JL

EMDB-35828: 
Cryo-EPty SPA at CSA of 1.03 mrad
Method: single particle / : Pei X, Wang P

EMDB-35916: 
Cryo-EPty SPA at CSA of 3.26 mrad
Method: single particle / : Pei XD, Wang P

EMDB-35917: 
Cryo-EPty SPA at CSA of 4.83 mrad
Method: single particle / : Pei XD, Wang P

EMDB-27103: 
Cryo-EM structure of T/F100 SOSIP.664 HIV-1 Env trimer with LMHS mutations in complex with Temsavir, 8ANC195, and 10-1074
Method: single particle / : Chen Y, Pozharski E, Tolbert W, Pazgier M

EMDB-27826: 
Cryo-EM structure of the full-length human NF1 dimer
Method: single particle / : Darling JE, Merk A, Grisshammer R, Ognjenovic J

PDB-8e20: 
Cryo-EM structure of the full-length human NF1 dimer
Method: single particle / : Darling JE, Merk A, Grisshammer R, Ognjenovic J

EMDB-29396: 
Antibody vFP53.02 in complex with HIV-1 envelope trimer BG505 DS-SOSIP
Method: single particle / : Wang S, Kwong PD
EMDB-29836: 
vFP52.02 Fab in complex with BG505 DS-SOSIP Env trimer
Method: single particle / : Gorman J, Kwong PD

EMDB-29880: 
Cryo-EM structure of vFP49.02 Fab in complex with HIV-1 Env BG505 DS-SOSIP.664 (conformation 1)
Method: single particle / : Changela A, Gorman J, Kwong PD

EMDB-29881: 
Cryo-EM structure of vFP49.02 Fab in complex with HIV-1 Env BG505 DS-SOSIP.664 (conformation 2)
Method: single particle / : Changela A, Gorman J, Kwong PD

EMDB-29882: 
Cryo-EM structure of vFP49.02 Fab in complex with HIV-1 Env BG505 DS-SOSIP.664 (conformation 3)
Method: single particle / : Changela A, Gorman J, Kwong PD

EMDB-29905: 
vFP48.02 Fab in complex with BG505 DS-SOSIP Env trimer
Method: single particle / : Gorman J, Kwong PD

PDB-8fr6: 
Antibody vFP53.02 in complex with HIV-1 envelope trimer BG505 DS-SOSIP
Method: single particle / : Wang S, Kwong PD

PDB-8g85: 
vFP52.02 Fab in complex with BG505 DS-SOSIP Env trimer
Method: single particle / : Gorman J, Kwong PD

PDB-8g9w: 
Cryo-EM structure of vFP49.02 Fab in complex with HIV-1 Env BG505 DS-SOSIP.664 (conformation 1)
Method: single particle / : Changela A, Gorman J, Kwong PD

PDB-8g9x: 
Cryo-EM structure of vFP49.02 Fab in complex with HIV-1 Env BG505 DS-SOSIP.664 (conformation 2)
Method: single particle / : Changela A, Gorman J, Kwong PD

PDB-8g9y: 
Cryo-EM structure of vFP49.02 Fab in complex with HIV-1 Env BG505 DS-SOSIP.664 (conformation 3)
Method: single particle / : Changela A, Gorman J, Kwong PD

PDB-8gas: 
vFP48.02 Fab in complex with BG505 DS-SOSIP Env trimer
Method: single particle / : Gorman J, Kwong PD

EMDB-27025: 
Cryo-EM structure of Human 15-PGDH in complex with small molecule SW222746
Method: single particle / : Huang W, Taylor DJ
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
