-Search query
-Search result
Showing 1 - 50 of 15,555 items for (author: hu & b)

EMDB-44372: 
In-cell Saccharomyces cerevisiae nuclear pore complex with single nuclear ring
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-44377: 
In-cell Saccharomyces cerevisiae nuclear pore complex with double nuclear ring and basket
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-44379: 
In-cell Mus musculus nuclear pore complex with nuclear basket
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-44381: 
In-cell Toxoplasma gondii nuclear pore complex
Method: subtomogram averaging / : Singh D, Hutchings J, Li Z, Guo Q, Villa E

EMDB-45197: 
In-cell Saccharomyces cerevisiae symmetry-expanded nuclear pore complex with double nuclear ring and basket
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45198: 
In-cell Saccharomyces cerevisiae symmetry-expanded nuclear pore complex with single nuclear ring
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45199: 
In-cell Saccharomyces cerevisiae nuclear pore complex cytoplasmic ring focused refinement
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45200: 
In-cell Saccharomyces cerevisiae nuclear pore complex inner ring focused refinement
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45201: 
In-cell Saccharomyces cerevisiae nuclear pore complex single nuclear ring focused refinement
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45202: 
In-cell Saccharomyces cerevisiae nuclear pore complex double nuclear ring focused refinement
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45203: 
In-cell Saccharomyces cerevisiae nuclear pore complex nuclear basket focused refinement
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45204: 
In-cell Saccharomyces cerevisiae nuclear pore complex membrane focused refinement for single nuclear ring
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45205: 
In-cell Saccharomyces cerevisiae nuclear pore complex membrane focused refinement for double nuclear ring
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45216: 
In-cell Mus musculus nuclear pore complex with nuclear basket consensus map
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45219: 
In-cell Mus musculus nuclear pore complex cytoplasmic ring focused refinement
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45220: 
In-cell Mus musculus nuclear pore complex inner ring focused refinement
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45222: 
In-cell Mus musculus nuclear pore complex nuclear ring focused refinement
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45223: 
In-cell Mus musculus nuclear pore complex basket focused refinement
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45227: 
In-cell Mus musculus nuclear pore complex membrane focused refinement
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45228: 
In-cell Toxoplasma gondii symmetry-expanded nuclear pore complex
Method: subtomogram averaging / : Singh D, Hutchings J, Li Z, Guo Q, Villa E
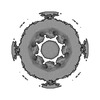
EMDB-45255: 
In-cell Saccharomyces cerevisiae C8-symmetrised nuclear pore complex consensus map
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45256: 
In-cell Saccharomyces cerevisiae symmetry-expanded nuclear pore complex consensus map
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45257: 
In-cell Mus musculus nuclear pore complex with nuclear basket consensus map
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45258: 
In-cell Mus musculus symmetry-expanded nuclear pore complex with nuclear basket consensus map
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45259: 
In-cell Toxoplasma gondii C8-symmetrised nuclear pore complex consensus map
Method: subtomogram averaging / : Singh D, Hutchings J, Li Z, Guo Q, Villa E
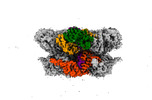
EMDB-42603: 
Human p97/VCP structure with a triazole inhibitor (NSC799462/hexamer)
Method: single particle / : Nandi P, DeVore K, Chiu PL
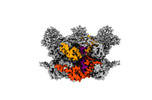
EMDB-42625: 
Human p97/VCP R155H mutant structure with a triazole inhibitor (NSC804515)
Method: single particle / : Nandi P, DeVore K, Chiu PL
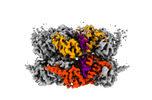
EMDB-42626: 
Human p97/VCP R155H mutant structure with a triazole inhibitor (NSC819701/up)
Method: single particle / : Nandi P, DeVore K, Chiu PL
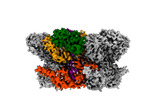
EMDB-42627: 
Human p97/VCP R155H mutant structure with a triazole inhibitor (NSC819701/down)
Method: single particle / : Nandi P, DeVore K, Chiu PL
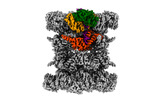
EMDB-44748: 
Human p97/VCP structure with a triazole inhibitor (NSC799462/dodecamer)
Method: single particle / : Nandi P, DeVore K, Chiu PL

EMDB-41766: 
CryoEM structure of D2 dopamine receptor in complex with GoA KE mutant, scFv16, and dopamine
Method: single particle / : Krumm BE, Kapolka NJ, Fay JF, Roth BL

EMDB-41776: 
CryoEM structure of D2 dopamine receptor in complex with GoA KE mutant and dopamine
Method: single particle / : Krumm BE, Kapolka NJ, Fay JF, Roth BL

EMDB-44551: 
Map of eastern equine encephalitis virus q3 spike protein in complex with VLDLR without masked refinement
Method: single particle / : Abraham J, Yang P, Li W, Fan X, Pan J

EMDB-45206: 
Reconstituted P400 Subcomplex of the human TIP60 complex
Method: single particle / : Yang Z, Mameri A, Florez Ariza AJ, Cote J, Nogales E

EMDB-45240: 
P400 subcomplex of the native human TIP60 complex
Method: single particle / : Yang Z, Mameri A, Florez Ariza AJ, Cote J, Nogales E

EMDB-45252: 
ARP module of the human TIP60 complex
Method: single particle / : Yang Z, Mameri A, Florez Ariza AJ, Cote J, Nogales E
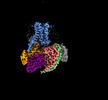
EMDB-60628: 
Carazolol-activated human beta3 adrenergic receptor
Method: single particle / : Zheng S, Zhang S, Dai S, Chen K, Gao K, Lin B, Liu X
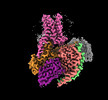
EMDB-60629: 
Epinephrine-activated human beta3 adrenergic receptor
Method: single particle / : Zheng S, Zhang S, Dai S, Chen K, Gao K, Lin B, Liu X

EMDB-42291: 
Structure of the human INTS9-INTS11-BRAT1 complex
Method: single particle / : Lin M, Tong L

EMDB-42292: 
Structure of the Drosophila IntS11-CG7044(dBRAT1) complex
Method: single particle / : Lin M, Tong L

EMDB-31367: 
Structure of mumps virus nucleoprotein without C-arm
Method: helical / : Shen Q, Shan H, Zhang N, Qin Y

EMDB-60100: 
SARS-CoV-2 spike trimer (6P) in complex with three R1-26 Fabs
Method: single particle / : Yan Q, Gao X, Liu B, Hou R, He P, Li Z, Chen Q, Wang J, He J, Chen L, Zhao J, Xiong X

EMDB-60101: 
SARS-CoV-2 spike trimer (6P) in complex with R1-26 Fab, head-to-head aggregate
Method: single particle / : Yan Q, Gao X, Liu B, Hou R, He P, Li Z, Chen Q, Wang J, He J, Chen L, Zhao J, Xiong X

EMDB-60102: 
SARS-CoV-2 spike trimer (6P) in complex with R1-26 Fab, focused refinement of RBD-Fab region
Method: single particle / : Yan Q, Gao X, Liu B, Hou R, He P, Li Z, Chen Q, Wang J, He J, Chen L, Zhao J, Xiong X

EMDB-60103: 
SARS-CoV-2 spike trimer (6P) in complex with two H18 Fabs
Method: single particle / : Yan Q, Gao X, Liu B, Hou R, He P, Li Z, Chen Q, Wang J, He J, Chen L, Zhao J, Xiong X

EMDB-60104: 
SARS-CoV-2 spike trimer (6P) in complex with three H18 Fabs
Method: single particle / : Yan Q, Gao X, Liu B, Hou R, He P, Li Z, Chen Q, Wang J, He J, Chen L, Zhao J, Xiong X

EMDB-60105: 
SARS-CoV-2 spike trimer (6P) in complex with three H18 Fabs, head-to-head aggregate (C1 symmetry)
Method: single particle / : Yan Q, Gao X, Liu B, Hou R, He P, Li Z, Chen Q, Wang J, He J, Chen L, Zhao J, Xiong X

EMDB-60106: 
SARS-CoV-2 spike trimer (6P) in complex with three H18 Fabs, head-to-head aggregate (C3 symmetry)
Method: single particle / : Yan Q, Gao X, Liu B, Hou R, He P, Li Z, Chen Q, Wang J, He J, Chen L, Zhao J, Xiong X

EMDB-60107: 
SARS-CoV-2 spike trimer (6P) in complex with two H18 and two R1-32 Fabs
Method: single particle / : Yan Q, Gao X, Liu B, Hou R, He P, Li Z, Chen Q, Wang J, He J, Chen L, Zhao J, Xiong X

EMDB-60108: 
SARS-CoV-2 spike trimer (6P) in complex with three H18 and three R1-32 Fabs
Method: single particle / : Yan Q, Gao X, Liu B, Hou R, He P, Li Z, Chen Q, Wang J, He J, Chen L, Zhao J, Xiong X
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
