-Search query
-Search result
Showing 1 - 50 of 1,744 items for (author: gui & m)

EMDB-17375: 
Neisseria meningitidis Type IV pilus SB-GATDH variant
Method: helical / : Fernandez-Martinez D, Dumenil G

EMDB-17384: 
Neisseria meningitidis Type IV pilus SB-DATDH variant
Method: helical / : Fernandez-Martinez D, Dumenil G

EMDB-17386: 
Neisseria meningitidis Type IV pilus SA-GATDH variant
Method: helical / : Fernandez-Martinez D, Dumenil G

EMDB-17683: 
Neisseria meningitidis Type IV pilus SB-GATDH variant bound to the C24 nanobody
Method: helical / : Fernandez-Martinez D, Dumenil G

EMDB-17695: 
Neisseria meningitidis Type IV pilus SB-DATDH variant bound to the C24 nanobody
Method: helical / : Fernandez-Martinez D, Dumenil G

EMDB-17718: 
Neisseria meningitidis PilE, SB-GATDH variant, bound to the F10 nanobody
Method: helical / : Fernandez-Martinez D, Dumenil G

PDB-8p2v: 
Neisseria meningitidis Type IV pilus SB-GATDH variant
Method: helical / : Fernandez-Martinez D, Dumenil G

PDB-8p36: 
Neisseria meningitidis Type IV pilus SB-DATDH variant
Method: helical / : Fernandez-Martinez D, Dumenil G

PDB-8p3b: 
Neisseria meningitidis Type IV pilus SA-GATDH variant
Method: helical / : Fernandez-Martinez D, Dumenil G

PDB-8pij: 
Neisseria meningitidis Type IV pilus SB-GATDH variant bound to the C24 nanobody
Method: helical / : Fernandez-Martinez D, Dumenil G

PDB-8piz: 
Neisseria meningitidis Type IV pilus SB-DATDH variant bound to the C24 nanobody
Method: helical / : Fernandez-Martinez D, Dumenil G

PDB-8pjp: 
Neisseria meningitidis PilE, SB-GATDH variant, bound to the F10 nanobody
Method: helical / : Fernandez-Martinez D, Dumenil G

EMDB-36870: 
Structure of full Banna virus
Method: single particle / : Li Z, Cao S

EMDB-36871: 
In situ structure of RNA-dependent RNA polymerase in full BAV particles
Method: single particle / : Li Z, Cao S

EMDB-36872: 
Structure of VP9 in Banna virus
Method: single particle / : Li Z, Cao S

EMDB-36880: 
Structure of partial Banna virus
Method: single particle / : Li Z, Cao S

EMDB-36881: 
Structure of Banna virus core
Method: single particle / : Li Z, Cao S

EMDB-37378: 
Structure of full Banna virus
Method: single particle / : Li Z, Cao S

EMDB-37379: 
Structure of partial Banna virus
Method: single particle / : Li Z, Cao S

EMDB-37380: 
Structure of Banna virus core
Method: single particle / : Li Z, Cao S

PDB-8k43: 
In situ structure of RNA-dependent RNA polymerase in full BAV particles
Method: single particle / : Li Z, Cao S

EMDB-42812: 
PNMA2 capsid, overall icosahedral map
Method: single particle / : Wilkinson ME, Madigan V, Zhang Y, Zhang F

EMDB-42815: 
PNMA2 capsid, focussed refinement of a pentamer (C5 symmetry)
Method: single particle / : Wilkinson ME, Madigan V, Zhang Y, Zhang F

PDB-8uyo: 
Structure of a recombinant human PNMA2 capsid
Method: single particle / : Wilkinson ME, Madigan V, Zhang Y, Zhang F

EMDB-35832: 
Cryo-EM structure of the GPR34 receptor in complex with the antagonist YL-365
Method: single particle / : Jia GW, Wang X, Zhang CB, Dong HH, Su ZM

PDB-8iyx: 
Cryo-EM structure of the GPR34 receptor in complex with the antagonist YL-365
Method: single particle / : Jia GW, Wang X, Zhang CB, Dong HH, Su ZM
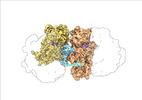
EMDB-19054: 
Escherichia coli paused disome complex (Non-rotated disome interface class 1)
Method: single particle / : Fluegel T, Schacherl M

PDB-8rcl: 
Escherichia coli paused disome complex (Non-rotated disome interface class 1)
Method: single particle / : Fluegel T, Schacherl M
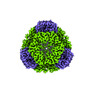
EMDB-42286: 
T33-ml28 - Designed Tetrahedral Protein Cage Using Machine Learning Algorithms
Method: single particle / : Castells-Graells R, Meador K, Sawaya MR, Yeates TO
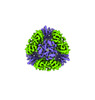
EMDB-42355: 
T33-ml30 - Designed Tetrahedral Protein Cage Using Machine Learning Algorithms
Method: single particle / : Castells-Graells R, Meador K, Sawaya MR, Yeates TO
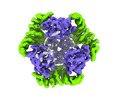
EMDB-42382: 
T33-ml35 Assembly Intermediate - Designed Tetrahedral Protein Cage Using Machine Learning Algorithms
Method: single particle / : Castells-Graells R, Meador K, Sawaya MR, Yeates TO
EMDB-42390: 
T33-ml23 Assembly Intermediate - Designed Tetrahedral Protein Cage Using Machine Learning Algorithms
Method: single particle / : Castells-Graells R, Meador K, Sawaya MR, Yeates TO

PDB-8ui2: 
T33-ml28 - Designed Tetrahedral Protein Cage Using Machine Learning Algorithms
Method: single particle / : Castells-Graells R, Meador K, Sawaya MR, Yeates TO

PDB-8ukm: 
T33-ml30 - Designed Tetrahedral Protein Cage Using Machine Learning Algorithms
Method: single particle / : Castells-Graells R, Meador K, Sawaya MR, Yeates TO

PDB-8umr: 
T33-ml35 Assembly Intermediate - Designed Tetrahedral Protein Cage Using Machine Learning Algorithms
Method: single particle / : Castells-Graells R, Meador K, Sawaya MR, Yeates TO

PDB-8un1: 
T33-ml23 Assembly Intermediate - Designed Tetrahedral Protein Cage Using Machine Learning Algorithms
Method: single particle / : Castells-Graells R, Meador K, Sawaya MR, Yeates TO

EMDB-29951: 
CryoEM structure of beta-2-adrenergic receptor in complex with nucleotide-free Gs heterotrimer (#1 of 20)
Method: single particle / : Papasergi-Scott MM, Skiniotis G

EMDB-29952: 
CryoEM structure of beta-2-adrenergic receptor in complex with nucleotide-free Gs heterotrimer (#2 of 20)
Method: single particle / : Papasergi-Scott MM, Skiniotis G

EMDB-29953: 
CryoEM structure of beta-2-adrenergic receptor in complex with nucleotide-free Gs heterotrimer (#3 of 20)
Method: single particle / : Papasergi-Scott MM, Skiniotis G

EMDB-29954: 
CryoEM structure of beta-2-adrenergic receptor in complex with nucleotide-free Gs heterotrimer (#4 of 20)
Method: single particle / : Papasergi-Scott MM, Skiniotis G

EMDB-29955: 
CryoEM structure of beta-2-adrenergic receptor in complex with nucleotide-free Gs heterotrimer (#5 of 20)
Method: single particle / : Papasergi-Scott MM, Skiniotis G

EMDB-29956: 
CryoEM structure of beta-2-adrenergic receptor in complex with nucleotide-free Gs heterotrimer (#6 of 20)
Method: single particle / : Papasergi-Scott MM, Skiniotis G
EMDB-29958: 
CryoEM structure of beta-2-adrenergic receptor in complex with nucleotide-free Gs heterotrimer (#7 of 20)
Method: single particle / : Papasergi-Scott MM, Skiniotis G
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search










 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
