-Search query
-Search result
Showing 1 - 50 of 102 items for (author: guan & cc)

EMDB-33883: 
Structural basis of human PRPS2 filaments
Method: single particle / : Lu GM, Hu HH, Liu JL

EMDB-14922: 
cryo-EM structure of omicron spike in complex with de novo designed binder, full map
Method: single particle / : Pablo G, Sarah W, Alexandra VH, Anthony M, Andreas S, Zander H, Dongchun N, Shuguang T, Freyr S, Casper G, Priscilla T, Alexandra T, Stephane R, Sandrine G, Jane M, Aaron P, Zepeng X, Yan C, Pu H, George G, Elisa O, Beat F, Didier T, Henning S, Michael B, Bruno EC

PDB-7zrv: 
cryo-EM structure of omicron spike in complex with de novo designed binder, full map
Method: single particle / : Pablo G, Sarah W, Alexandra VH, Anthony M, Andreas S, Zander H, Dongchun N, Shuguang T, Freyr S, Casper G, Priscilla T, Alexandra T, Stephane R, Sandrine G, Jane M, Aaron P, Zepeng X, Yan C, Pu H, George G, Elisa O, Beat F, Didier T, Henning S, Michael B, Bruno EC

EMDB-14930: 
cryo-EM structure of omicron spike in complex with de novo designed binder, local
Method: single particle / : Pablo G, Sarah W, Alexandra VH, Anthony M, Andreas S, Zander H, Dongchun N, Shuguang T, Freyr S, Casper G, Priscilla T, Alexandra T, Stephane R, Sandrine G, Jane M, Aaron P, Zepeng X, Yan C, Pu H, George G, Elisa O, Beat F, Didier T, Henning S, Michael B, Bruno EC

EMDB-14947: 
cryo-EM structure of D614 spike in complex with de novo designed binder, full and local maps(addition)
Method: single particle / : Pablo G, Sarah W, Alexandra VH, Anthony M, Andreas S, Zander H, Dongchun N, Shuguang T, Freyr S, Casper G, Priscilla T, Alexandra T, Stephane R, Sandrine G, Jane M, Aaron P, Zepeng X, Yan C, Pu H, George G, Elisa O, Beat F, Didier T, Henning S, Michael B, Bruno EC

PDB-7zsd: 
cryo-EM structure of omicron spike in complex with de novo designed binder, local
Method: single particle / : Pablo G, Sarah W, Alexandra VH, Anthony M, Andreas S, Zander H, Dongchun N, Shuguang T, Freyr S, Casper G, Priscilla T, Alexandra T, Stephane R, Sandrine G, Jane M, Aaron P, Zepeng X, Yan C, Pu H, George G, Elisa O, Beat F, Didier T, Henning S, Michael B, Bruno EC

PDB-7zss: 
cryo-EM structure of D614 spike in complex with de novo designed binder
Method: single particle / : Pablo G, Sarah W, Alexandra VH, Anthony M, Andreas S, Zander H, Dongchun N, Shuguang T, Freyr S, Casper G, Priscilla T, Alexandra T, Stephane R, Sandrine G, Jane M, Aaron P, Zepeng X, Yan C, Pu H, George G, Elisa O, Beat F, Didier T, Henning S, Michael B, Bruno EC

EMDB-28092: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-093
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-33374: 
Focused refinement cryo-EM map of the A/B/C subunits of the T=4 lake sinai virus 2 virus-like particle at pH 7.5
Method: single particle / : Chen NC, Wang CH, Chen CJ, Yoshimura M, Guan HH, Chuankhayan P, Lin CC

EMDB-33375: 
Focus refinement cryo-EM map of the D/D/D subunits of the T=4 lake sinai virus 2 virus-like particle
Method: single particle / : Chen NC, Wang CH, Chen CJ, Yoshimura M, Guan HH, Chuankhayan P, Lin CC

EMDB-33376: 
Focused refinement cryo-EM map of the A/B/C subunits of the T=3 lake sinai virus 2 virus-like particle at pH 7.5
Method: single particle / : Chen NC, Wang CH, Chen CJ, Yoshimura M, Guan HH, Chuankhayan P, Lin CC

EMDB-33377: 
Focused refinement cryo-EM map of the A/B/C subunits of the T=4 lake sinai virus 2 virus-like particle at pH 6.5
Method: single particle / : Chen NC, Wang CH, Chen CJ, Yoshimura M, Guan HH, Chuankhayan P, Lin CC

EMDB-33378: 
Focused refinement cryo-EM map of the D/D/D subunits of the T=4 lake sinai virus 2 virus-like particle at pH 6.5
Method: single particle / : Chen NC, Wang CH, Chen CJ, Yoshimura M, Guan HH, Chuankhayan P, Lin CC

EMDB-33379: 
Focused refinement cryo-EM map of the A/B/C subunits of the T=3 lake sinai virus 2 virus-like particle at pH 6.5
Method: single particle / : Chen NC, Wang CH, Chen CJ, Yoshimura M, Guan HH, Chuankhayan P, Lin CC

EMDB-33380: 
Focused refinement cryo-EM map of the A/B/C subunits of the T=4 lake sinai virus 2 virus-like particle at pH 8.5
Method: single particle / : Chen NC, Wang CH, Chen CJ, Yoshimura M, Guan HH, Chuankhayan P, Lin CC

EMDB-33381: 
Focused refinement cryo-EM map of the D/D/D subunits of the T=4 lake sinai virus 2 virus-like particle at pH 8.5
Method: single particle / : Chen NC, Wang CH, Chen CJ, Yoshimura M, Guan HH, Chuankhayan P, Lin CC

EMDB-33382: 
Focused refinement cryo-EM map of the A/B/C subunits of the T=3 lake sinai virus 2 virus-like particle at pH 8.5
Method: single particle / : Chen NC, Wang CH, Chen CJ, Yoshimura M, Guan HH, Chuankhayan P, Lin CC

EMDB-33383: 
Focused refinement cryo-EM map of the A/B/C subunits of the T=3 lake sinai virus 1 (delta N-terminal 48 residues) virus-like particle at pH 6.5
Method: single particle / : Chen NC, Wang CH, Chen CJ, Yoshimura M, Guan HH, Chuankhayan P, Lin CC

EMDB-33384: 
Cryo-EM map of the T=4 lake sinai virus 1 (delta N-terminal 48 residues) virus-like particle at pH 6.5
Method: single particle / : Chen NC, Wang CH, Chen CJ, Yoshimura M, Guan HH, Chuankhayan P, Lin CC

EMDB-28090: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-040
Method: single particle / : Li H, Callaway H, Yu X, Shek J, Saphire EO

EMDB-28091: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-045
Method: single particle / : Li H, Callaway H, Yu X, Shek J, Saphire EO

EMDB-28093: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-156
Method: single particle / : Shek J, Callaway H, Li H, Yu X, Saphire EO

EMDB-33190: 
Cryo-EM structure of the T=4 lake sinai virus 2 virus-like capsid at pH 7.5
Method: single particle / : Chen NC, Wang CH, Chen CJ, Yoshimura M, Guan HH, Chuankhayan P, Lin CC

EMDB-33368: 
Cryo-EM structure of the T=3 lake sinai virus 2 virus-like capsid at pH 7.5
Method: single particle / : Chen NC, Wang CH, Chen CJ, Yoshimura M, Guan HH, Chuankhayan P, Lin CC

EMDB-33369: 
Cryo-EM structure of the T=4 lake sinai virus 2 virus-like capsid at pH 6.5
Method: single particle / : Chen NC, Wang CH, Chen CJ, Yoshimura M, Guan HH, Chuankhayan P, Lin CC

EMDB-33370: 
Cryo-EM structure of the T=3 lake sinai virus 2 virus-like capsid at pH 6.5
Method: single particle / : Chen NC, Wang CH, Chen CJ, Yoshimura M, Guan HH, Chuankhayan P, Lin CC

EMDB-33371: 
Cryo-EM structure of the T=4 lake sinai virus 2 virus-like capsid at pH 8.5
Method: single particle / : Chen NC, Wang CH, Chen CJ, Yoshimura M, Guan HH, Chuankhayan P, Lin CC

EMDB-33372: 
Cryo-EM structure of the T=3 lake sinai virus 2 virus-like capsid at pH 8.5
Method: single particle / : Chen NC, Wang CH, Chen CJ, Yoshimura M, Guan HH, Chuankhayan P, Lin CC

EMDB-33373: 
Cryo-EM structure of the T=3 lake sinai virus 1 (delta-N48) virus-like capsid at pH 6.5
Method: single particle / : Chen NC, Wang CH, Chen CJ, Yoshimura M, Guan HH, Chuankhayan P, Lin CC

EMDB-28094: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-234
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28095: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-260
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28096: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-279
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28097: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-290
Method: single particle / : Yu X, Callaway H, Li H, Shek J, Saphire EO
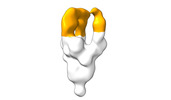
EMDB-28098: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-294
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28099: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-295
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28100: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-299
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28102: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-334
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO
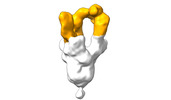
EMDB-28103: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-360
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28104: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-361
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28105: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-362
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28106: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-368
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28168: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-292
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28169: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-333
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO
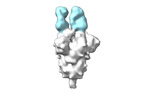
EMDB-28170: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-355
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28171: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-371
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-32541: 
Structural basis for ligand binding modes of CTP synthase
Method: single particle / : Liu JL, Guo CJ

EMDB-32542: 
Structural basis for ligand binding modes of CTP synthase
Method: single particle / : Liu JL, Guo CJ

EMDB-33305: 
E.coli phosphoribosylpyrophosphate (PRPP) synthetase type A filament bound with ADP, Pi and R5P
Method: single particle / : Hu HH, Lu GM, Chang CC, Liu JL

EMDB-33306: 
E.coli phosphoribosylpyrophosphate (PRPP) synthetase type A(AMP/ADP) filament bound with ADP, AMP and R5P
Method: single particle / : Hu HH, Lu GM, Chang CC, Liu JL

EMDB-33309: 
E.coli phosphoribosylpyrophosphate (PRPP) synthetase type B filament bound with Pi
Method: single particle / : Hu HH, Lu GM, Chang CC, Liu JL
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
