-Search query
-Search result
Showing all 31 items for (author: gilmore & r)

EMDB-25673: 
Prepore structure of pore-forming toxin Epx1
Method: single particle / : Xiong XZ, Yang P

PDB-7t4e: 
Prepore structure of pore-forming toxin Epx1
Method: single particle / : Xiong XZ, Yang P, Dong M, Abraham J

EMDB-8926: 
Mutated p53-DNA assembly
Method: single particle / : Kelly DF, Dearnaley WJ, Liang Y

EMDB-8927: 
Wild-type p53-DNA assembly
Method: single particle / : Kelly DF, Dearnaley WJ, Liang Y

EMDB-4306: 
Subtomogram average of unsorted ribosome-translocon complexes from STT3B(-/-) HEK cells
Method: subtomogram averaging / : Pfeffer S, Foerster F

EMDB-4307: 
Subtomogram average of OST-free ribosome-translocon complexes from STT3B(-/-) HEK cells
Method: subtomogram averaging / : Pfeffer S, Foerster F

EMDB-4308: 
Subtomogram average of OST-containing ribosome-translocon complexes from STT3B(-/-) HEK cells
Method: subtomogram averaging / : Pfeffer S, Foerster F
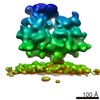
EMDB-4309: 
Subtomogram average of unsorted ribosome-translocon complexes from STT3A(-/-) HEK cells
Method: subtomogram averaging / : Pfeffer S, Foerster F
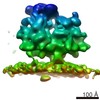
EMDB-4310: 
Subtomogram average of OST-free ribosome-translocon complexes from STT3A(-/-) HEK cells
Method: subtomogram averaging / : Pfeffer S, Foerster F

EMDB-4311: 
Subtomogram average of ribosome-translocon complexes from STT3A(-/-) HEK cells containing an unidentified ER-lumenal component
Method: subtomogram averaging / : Pfeffer S, Foerster F
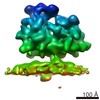
EMDB-4312: 
Subtomogram average of unsorted ribosome-translocon complexes from wild type HEK cells
Method: subtomogram averaging / : Pfeffer S, Foerster F
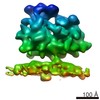
EMDB-4313: 
Subtomogram average of OST-free ribosome-translocon complexes from wild type HEK cells
Method: subtomogram averaging / : Pfeffer S, Foerster F
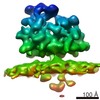
EMDB-4314: 
Subtomogram average of OST-containing ribosome-translocon complexes from wild type HEK cells
Method: subtomogram averaging / : Pfeffer S, Foerster F

EMDB-4315: 
Subtomogram average of OST-containing ribosome-translocon complexes from canine rough microsomal membranes
Method: subtomogram averaging / : Pfeffer S, Foerster F

EMDB-4316: 
Cryo-EM Structure of the Mammalian Oligosaccharyltransferase Bound to Sec61 and the Programmed 80S Ribosome
Method: single particle / : Braunger K, Becker T, Beckmann R

EMDB-4317: 
Cryo-EM Structure of the Mammalian Oligosaccharyltransferase Bound to Sec61 and the Non-programmed 80S Ribosome
Method: single particle / : Braunger K, Becker T, Beckmann R

PDB-6ftg: 
Subtomogram average of OST-containing ribosome-translocon complexes from canine rough microsomal membranes
Method: subtomogram averaging / : Pfeffer S, Foerster F

PDB-6fti: 
Cryo-EM Structure of the Mammalian Oligosaccharyltransferase Bound to Sec61 and the Programmed 80S Ribosome
Method: single particle / : Braunger K, Becker T, Beckmann R

PDB-6ftj: 
Cryo-EM Structure of the Mammalian Oligosaccharyltransferase Bound to Sec61 and the Non-programmed 80S Ribosome
Method: single particle / : Braunger K, Becker T, Beckmann R

EMDB-8833: 
Mutated BRCA1-BARD1
Method: single particle / : Kelly DF, Dearnaley WJ

EMDB-8834: 
Wild type BRCA1-BARD1
Method: single particle / : Liang Y, Dearnaley WJ, Kelly D
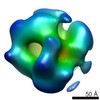
EMDB-6400: 
Electron microscopy of BRCA1(5832insC) mutant-RNAP II transcriptional complex
Method: single particle / : Winton CE, Gilmore BL, Demmert AC, Tanner JR, Bowman S, Karageorge V, Patel K, Sheng Z, Kelly DF

EMDB-6340: 
Electron cryo-microscopy of a BRCA1-RNAPII transcriptional complex
Method: single particle / : Gilmore BL, Winton CE, Demmert AC, Tanner JR, Bowman S, Karageorge V, Patel K, Sheng Z, Kelly DF

EMDB-1651: 
Cryo-EM structure of the programmed yeast 80 ribosome bound the Ssh1 complex
Method: single particle / : Becker T, Mandon E, Bhushan S, Jarasch A, Armache JP, Funes S, Jossinet F, Gumbart J, Mielke T, Berninghausen O, Schulten K, Westhof E, Gilmore R, Beckmann R

EMDB-1667: 
Cryo-EM structure of the active yeast Ssh1 complex bound to the programmed yeast 80S ribosome bearing a P-site tRNA
Method: single particle / : Becker T, Mandon E, Bhushan S, Jarasch A, Armache JP, Funes S, Jossinet F, Gumbart J, Mielke T, Berninghausen O, Schulten K, Westhof E, Gilmore R, Beckmann R

EMDB-1668: 
Cryo-EM structure of the active yeast 80S ribosome bearing a P-site tRNA and with the rRNA expansion segment ES27 in the exit conformation
Method: single particle / : Becker T, Mandon E, Bhushan S, Jarasch A, Armache JP, Funes S, Jossinet F, Gumbart J, Mielke T, Berninghausen O, Schulten K, Westhof E, Gilmore R, Beckmann R

EMDB-1669: 
Cryo-EM structures of the idle yeast Ssh1 complex bound to the yeast 80S ribosome
Method: single particle / : Becker T, Mandon E, Bhushan S, Jarasch A, Armache JP, Funes S, Jossinet F, Gumbart J, Mielke T, Berninghausen O, Schulten K, Westhof E, Gilmore R, Beckmann R

EMDB-1652: 
Cryo-EM structure of the mammalian Sec61 complex bound to the actively translating wheat germ 80S ribosome
Method: single particle / : Becker T, Mandon E, Bhushan S, Jarasch A, Armache JP, Funes S, Jossinet F, Gumbart J, Mielke T, Berninghausen O, Schulten K, Westhof E, Gilmore R, Beckmann R

PDB-2ww9: 
Cryo-EM structure of the active yeast Ssh1 complex bound to the yeast 80S ribosome
Method: single particle / : Becker T, Mandon E, Bhushan S, Jarasch A, Armache JP, Funes S, Jossinet F, Gumbart J, Mielke T, Berninghausen O, Schulten K, Westhof E, Gilmore R, Beckmann R

PDB-2wwa: 
Cryo-EM structure of idle yeast Ssh1 complex bound to the yeast 80S ribosome
Method: single particle / : Becker T, Mandon E, Bhushan S, Jarasch A, Armache JP, Funes S, Jossinet F, Gumbart J, Mielke T, Berninghausen O, Schulten K, Westhof E, Gilmore R, Beckmann R

PDB-2wwb: 
CRYO-EM STRUCTURE OF THE MAMMALIAN SEC61 COMPLEX BOUND TO THE ACTIVELY TRANSLATING WHEAT GERM 80S RIBOSOME
Method: single particle / : Becker T, Mandon E, Bhushan S, Jarasch A, Armache JP, Funes S, Jossinet F, Gumbart J, Mielke T, Berninghausen O, Schulten K, Westhof E, Gilmore R, Beckmann R
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
