-Search query
-Search result
Showing 1 - 50 of 67 items for (author: ekiert & dc)

EMDB-28966: 
CryoEM map of de novo designed oligomeric protein C4-71_6x
Method: single particle / : Redler RL, Edman NI, Baker D, Ekiert DC, Bhabha G

EMDB-28967: 
CryoEM map of de novo designed oligomeric protein C4-71_8x
Method: single particle / : Redler RL, Edman NI, Baker D, Ekiert DC, Bhabha G
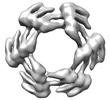
EMDB-28968: 
CryoEM map of de novo designed oligomeric protein C6-71
Method: single particle / : Redler RL, Edman NI, Baker D, Ekiert DC, Bhabha G

EMDB-28969: 
CryoEM map of de novo designed oligomeric protein C6-71_6x
Method: single particle / : Redler RL, Edman NI, Baker D, Ekiert DC, Bhabha G

EMDB-28970: 
CryoEM map of de novo designed oligomeric protein C6-71_8x
Method: single particle / : Redler RL, Edman NI, Baker D, Ekiert DC, Bhabha G

EMDB-28971: 
CryoEM map of de novo designed oligomeric protein C8-71_6x
Method: single particle / : Redler RL, Edman NI, Baker D, Ekiert DC, Bhabha G

EMDB-28972: 
CryoEM map of de novo designed oligomeric protein C8-71_8x
Method: single particle / : Redler RL, Edman NI, Baker D, Ekiert DC, Bhabha G
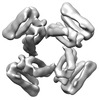
EMDB-28973: 
CryoEM map of de novo designed oligomeric protein C4-81
Method: single particle / : Redler RL, Edman NI, Baker D, Ekiert DC, Bhabha G

EMDB-28974: 
CryoEM map of designed oligomeric protein C4-71
Method: single particle / : Redler RL, Edman NI, Baker D, Ekiert DC, Bhabha G

EMDB-40208: 
Backbone model of de novo-designed chlorophyll-binding nanocage O32-15
Method: single particle / : Redler RL, Ennist NM, Wang S, Baker D, Ekiert DC, Bhabha G

EMDB-40209: 
Chlorophyll-binding region of de novo-designed nanocage O32-15
Method: single particle / : Redler RL, Ennist NM, Wang S, Baker D, Ekiert DC, Bhabha G

EMDB-28958: 
CryoEM structure of designed modular protein oligomer C4-131
Method: single particle / : Redler RL, Edman NI, Baker D, Ekiert D, Bhabha G

EMDB-28889: 
CryoEM structure of designed modular protein oligomer C6-79
Method: single particle / : Redler RL, Edman NI, Baker D, Ekiert D, Bhabha G

EMDB-28888: 
CryoEM structure of designed modular protein oligomer C8-71
Method: single particle / : Redler RL, Edman NI, Baker D, Ekiert D, Bhabha G

EMDB-40162: 
E. coli MlaC bound to MlaFEDB
Method: single particle / : MacRae MR, Coudray N, Ekiert D, Bhabha G

EMDB-29023: 
Structure of Mce1-LucB complex from Mycobacterium smegmatis (Map1)
Method: single particle / : Chen J, Bhabha G, Ekiert DC

EMDB-29024: 
Structure of Mce1 transporter from Mycobacterium smegmatis in the absence of LucB (Map2)
Method: single particle / : Chen J, Bhabha G, Ekiert DC

EMDB-29025: 
Structure of Mce1 transporter from Mycobacterium smegmatis (Map0)
Method: single particle / : Chen J, Bhabha G, Ekiert DC

EMDB-29228: 
Local refinement of MceG in Msmeg Mce1 transporter (Map0a)
Method: single particle / : Chen J, Bhabha G, Ekiert DC

EMDB-29229: 
Local refinement of YrbE and MCE ring in Msmeg Mce1 transporter (Map0b)
Method: single particle / : Chen J, Bhabha G, Ekiert DC

EMDB-29230: 
Local refinement of MCE ring and needle in Msmeg Mce1 transporter (Map0c)
Method: single particle / : Chen J, Bhabha G, Ekiert DC

EMDB-29231: 
Local refinement of MCE needle and portal in Msmeg Mce1 transporter (Map0d)
Method: single particle / : Chen J, Bhabha G, Ekiert DC

EMDB-29232: 
Raw, consensus refinement of Msmeg Mce1 transporter (Map0e)
Method: single particle / : Chen J, Bhabha G, Ekiert DC

EMDB-29233: 
Local refinement of MceG in Msmeg Mce1-LucB complex (Map1a)
Method: single particle / : Chen J, Bhabha G, Ekiert DC

EMDB-29234: 
Local refinement of YrbE, MCE ring, LucB in Msmeg Mce1-LucB complex (Map1b)
Method: single particle / : Chen J, Bhabha G, Ekiert DC

EMDB-29235: 
Local refinement of MCE ring and needle in Msmeg Mce1-LucB complex (Map1c)
Method: single particle / : Chen J, Bhabha G, Ekiert DC

EMDB-29236: 
Local refinement of MCE needle and portal in Msmeg Mce1-LucB complex (Map1d)
Method: single particle / : Chen J, Bhabha G, Ekiert DC

EMDB-29237: 
Raw, consensus refinement of Msmeg Mce1-LucB complex (Map1e)
Method: single particle / : Chen J, Bhabha G, Ekiert DC

EMDB-29238: 
Local refinement of MceG in Msmeg Mce1 transporter in the absence of LucB (Map2a)
Method: single particle / : Chen J, Bhabha G, Ekiert DC

EMDB-29239: 
Local refinement of YrbE and MCE ring in Msmeg Mce1 transporter in the absence of LucB (Map2b)
Method: single particle / : Chen J, Bhabha G, Ekiert DC

EMDB-29240: 
Local refinement of MCE ring and needle in Msmeg Mce1 transporter in the absence of LucB (Map2c)
Method: single particle / : Chen J, Bhabha G, Ekiert DC

EMDB-29241: 
Local refinement of MCE needle and portal in Msmeg Mce1 transporter in the absence of LucB (Map2d)
Method: single particle / : Chen J, Bhabha G, Ekiert DC

EMDB-29242: 
Raw, consensus refinement of Msmeg Mce1 transporter in the absence of LucB (Map2e)
Method: single particle / : Chen J, Bhabha G, Ekiert DC

EMDB-29041: 
E. coli MlaFEDB bound to MlaC
Method: single particle / : MacRae MR, Coudray N, Ekiert D, Bhabha G

EMDB-25066: 
Structure of E. coli LetB delta (Ring6) mutant, Ring1 in the closed state (Model 1)
Method: single particle / : Vieni C, Coudray N

EMDB-25067: 
Structure of E. coli LetB delta (Ring6) mutant, Ring 1 in the open state (Model 2, Rings 1-3 only)
Method: single particle / : Vieni C, Coudray N

EMDB-23838: 
Signal subtracted reconstruction of AAA2, AAA3, and AAA4 domains of dynein in the presence of a pyrazolo-pyrimidinone-based compound, Map 4
Method: single particle / : Santarossa CC, Coudray N, Urnavicius L, Ekiert DC, Bhabha G, Kapoor TM

EMDB-23841: 
Yeast dynein motor domain in the presence of a pyrazolo-pyrimidinone-based compound, Map 1
Method: single particle / : Santarossa CC, Urnavicius L, Coudray N, Ekeirt DC, Bhabha G, Kapoor TM

EMDB-23842: 
Signal subtracted reconstruction of AAA5 and AAA6 domains of dynein in the presence of a pyrazolo-pyrimidinone-based compound, Map 5
Method: single particle / : Santarossa CC, Coudray N, Urnavicius L, Ekiert DC, Bhabha G, Kapoor TM

EMDB-23844: 
Signal subtracted reconstruction of the AAA1 domain of dynein in the presence of a pyrazolo-pyrimidinone-based compound, Map 2
Method: single particle / : Santarossa CC, Urnavicius L, Coudray N, Ekiert DC, Bhabha G, Kapoor TM

EMDB-23846: 
Signal subtracted reconstruction of the AAA1 domain of dynein in the presence of a pyrazolo-pyrimidinone-based compound, Map 3
Method: single particle / : Santarossa CC, Coudray N, Urnavicius L, Ekiert DC, Bhabha G, Kapoor TM

EMDB-22116: 
Cryo-EM structure of MlaFEDB in nanodiscs with phospholipid substrates
Method: single particle / : Coudray N, Isom GL

EMDB-20993: 
Lipophilic Envelope-spanning Tunnel B (LetB), Map 1
Method: single particle / : Isom GL, Coudray N

EMDB-20994: 
Lipophilic Envelope-spanning Tunnel B (LetB), Map 2
Method: single particle / : Isom GL, Coudray N

EMDB-20995: 
Lipophilic Envelope-spanning Tunnel B (LetB), Map 3
Method: single particle / : Isom GL, Coudray N

EMDB-20996: 
Lipophilic Envelope-spanning Tunnel B (LetB), Map 4
Method: single particle / : Isom GL, Coudray N

EMDB-20997: 
Lipophilic Envelope-spanning Tunnel B (LetB), Map 5
Method: single particle / : Isom GL, Coudray N

EMDB-20998: 
Lipophilic Envelope-spanning Tunnel B (LetB), Map 6
Method: single particle / : Isom GL, Coudray N

EMDB-20999: 
Lipophilic Envelope-spanning Tunnel B (LetB), Map 7
Method: single particle / : Isom GL, Coudray N

EMDB-21000: 
Lipophilic Envelope-spanning Tunnel B (LetB), Map 8
Method: single particle / : Isom GL, Coudray N
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
