-Search query
-Search result
Showing 1 - 50 of 101 items for (author: effantin & g)

EMDB-19405: 
Tomograms of spoIVB Bacillus subtilis sporangia
Method: electron tomography / : Bauda E, Gallet B, Moravcova J, Effantin G, Chan H, Novacek J, Jouneau PH, Rodrigues CDA, Schoehn G, Moriscot C, Morlot C
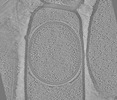
EMDB-19411: 
Tomograms of cotE Bacillus subtilis sporangia
Method: electron tomography / : Bauda E, Gallet B, Moravcova J, Effantin G, Chan H, Novacek J, Jouneau PH, Rodrigues CDA, Schoehn G, Moriscot C, Morlot C

EMDB-18453: 
Human Adenovirus type 11 fiber knob in complex with two copies of its cell receptor, Desmoglein-2
Method: single particle / : Effantin G

EMDB-18454: 
Human Adenovirus type 11 fiber knob in complex with two copies of its cell receptor, Desmoglein-2
Method: single particle / : Effantin G

EMDB-18455: 
Human Adenovirus type 11 fiber knob in complex with its cell receptors, Desmoglein-2 and CD46
Method: single particle / : Effantin G

PDB-8qjx: 
Human Adenovirus type 11 fiber knob in complex with two copies of its cell receptor, Desmoglein-2
Method: single particle / : Effantin G

PDB-8qjy: 
Human Adenovirus type 11 fiber knob in complex with two copies of its cell receptor, Desmoglein-2
Method: single particle / : Effantin G

PDB-8qk3: 
Human Adenovirus type 11 fiber knob in complex with its cell receptors, Desmoglein-2 and CD46
Method: single particle / : Effantin G

EMDB-15967: 
Cryo-EM structure of the proximal end of bacteriophage T5 tail : p142 tail terminator protein hexamer and pb6 tail tube protein trimer
Method: single particle / : Linares R, Effantin G, Breyton C, Darnault C, Epalle N, Boeri Erba E, Schoehn G

EMDB-15968: 
Cryo-EM structure of the proximal end of bacteriophage T5 tail, after interaction with its receptor : p142 tail terminator protein hexamer and pb6 tail tube protein trimer
Method: single particle / : Linares R, Effantin G, Breyton C, Darnault C, Epalle N, Boeri Erba E, Schoehn G

PDB-8bcp: 
Cryo-EM structure of the proximal end of bacteriophage T5 tail : p142 tail terminator protein hexamer and pb6 tail tube protein trimer
Method: single particle / : Linares R, Effantin G, Breyton C

PDB-8bcu: 
Cryo-EM structure of the proximal end of bacteriophage T5 tail, after interaction with its receptor : p142 tail terminator protein hexamer and pb6 tail tube protein trimer
Method: single particle / : Linares R, Effantin G, Breyton C

EMDB-15084: 
cryo-EM structure of thioredoxin glutathione reductase in complex with a non-competitive inhibitor
Method: single particle / : Ardini M, Angelucci F, Fata F, Gabriele F, Effantin G, Ling W, Williams DL, Petukhova VZ, Petukhov PA

PDB-8a1r: 
cryo-EM structure of thioredoxin glutathione reductase in complex with a non-competitive inhibitor
Method: single particle / : Ardini M, Angelucci F, Fata F, Gabriele F, Effantin G, Ling W, Williams DL, Petukhova VZ, Petukhov PA

EMDB-14733: 
Tail tip of siphophage T5 : tip proteins
Method: single particle / : Linares R, Arnaud CA, Effantin G, Darnault C, Epalle N, Boeri Erba E, Schoehn G, Breyton C

EMDB-14790: 
Tail tip of siphophage T5 : central fibre
Method: single particle / : Linares R, Arnaud CA, Effantin G, Epalle N, Boeri Erba E, Schoehn G, Breyton C

EMDB-14799: 
Tail tip of siphophage T5 : full complex after interaction with its bacterial receptor FhuA
Method: single particle / : Linares R, Arnaud CA, Effantin G, Darnault C, Epalle N, Boeri Erba E, Schoehn G, Breyton C

EMDB-14800: 
Tail tip of siphophage T5 : bent fibre after interaction with its bacterial receptor FhuA
Method: single particle / : Linares R, Arnaud CA, Effantin G, Darnault C, Epalle N, Boeri Erba E, Schoehn G, Breyton C

EMDB-14869: 
Tail tip of siphophage T5 : full structure
Method: single particle / : Linares R, Arnaud CA, Effantin G, Darnault C, Epalle N, Boeri Erba E, Schoehn G, Breyton C

EMDB-14873: 
Tail tip of siphophage T5 : open cone after interaction with bacterial receptor FhuA
Method: single particle / : Linares R, Arnaud CA, Effantin G, Darnault C, Epalle N, Boeri Erba E, Schoehn G, Breyton C

PDB-7zhj: 
Tail tip of siphophage T5 : tip proteins
Method: single particle / : Linares R, Arnaud CA, Effantin G, Darnault C, Epalle N, Boeri Erba E, Schoehn G, Breyton C

PDB-7zlv: 
Tail tip of siphophage T5 : central fibre protein pb4
Method: single particle / : Linares R, Arnaud CA, Effantin G, Epalle N, Boeri Erba E, Schoehn G, Breyton C

PDB-7zn2: 
Tail tip of siphophage T5 : full complex after interaction with its bacterial receptor FhuA
Method: single particle / : Linares R, Arnaud CA, Effantin G, Darnault C, Epalle N, Boeri Erba E, Schoehn G, Breyton C

PDB-7zn4: 
Tail tip of siphophage T5 : bent fibre after interaction with its bacterial receptor FhuA
Method: single particle / : Linares R, Arnaud CA, Effantin G, Darnault C, Epalle N, Boeri Erba E, Schoehn G, Breyton C

PDB-7zqb: 
Tail tip of siphophage T5 : full structure
Method: single particle / : Linares R, Arnaud CA, Effantin G, Darnault C, Epalle N, Boeri Erba E, Schoehn G, Breyton C

PDB-7zqp: 
Tail tip of siphophage T5 : open cone after interaction with bacterial receptor FhuA
Method: single particle / : Linares R, Arnaud CA, Effantin G, Darnault C, Epalle N, Boeri Erba E, Schoehn G, Breyton C

EMDB-15802: 
T5 Receptor Binding Protein pb5 in complex with its E. coli receptor FhuA
Method: single particle / : Degroux S, Effantin G, Linares R, Schoehn G, Breyton C

PDB-8b14: 
T5 Receptor Binding Protein pb5 in complex with its E. coli receptor FhuA
Method: single particle / : Degroux S, Effantin G, Linares R, Schoehn G, Breyton C

EMDB-14630: 
Membrane-bound CHMP2A-CHMP3 filament (430 Angstrom diameter)
Method: helical / : Azad K, Desfosses A, Effantin G, Schoehn G, Weissenhorn W

EMDB-14631: 
Membrane-bound CHMP2A-CHMP3 filament (410 Angstrom diameter)
Method: helical / : Azad K, Desfosses A, Effantin G, Schoehn G, Weissenhorn W

PDB-7zcg: 
CHMP2A-CHMP3 heterodimer (430 Angstrom diameter)
Method: helical / : Azad K, Desfosses A, Effantin G, Schoehn G, Weissenhorn W

PDB-7zch: 
CHMP2A-CHMP3 heterodimer (410 Angstrom diameter)
Method: helical / : Azad K, Desfosses A, Effantin G, Schoehn G, Weissenhorn W

EMDB-13953: 
Tail tip of siphophage T5 : common core proteins
Method: single particle / : Linares R, Arnaud CA, Effantin G, Darnault C, Epalle N, Boeri Erba E, Schoehn G, Breyton C

PDB-7qg9: 
Tail tip of siphophage T5 : common core proteins
Method: single particle / : Linares R, Arnaud CA, Effantin G, Darnault C, Epalle N, Boeri Erba E, Schoehn G, Breyton C

EMDB-4928: 
Cryo-EM reconstruction of TMV coat protein
Method: helical / : Kandiah E, Effantin G

PDB-6rlp: 
Cryo-EM reconstruction of TMV coat protein
Method: helical / : Kandiah E, Effantin G

EMDB-14571: 
Negative stain 3D structure of the plastid-encoded RNA polymerase from Sinapis alba
Method: single particle / : Effantin G, Fenel D

EMDB-12775: 
Cryo-EM structure of the plectasin fibril (double strands)
Method: helical / : Effantin G

EMDB-12776: 
Cryo-EM structure of the plectasin fibril (single strand)
Method: helical / : Effantin G

PDB-7oae: 
Cryo-EM structure of the plectasin fibril (double strands)
Method: helical / : Effantin G

PDB-7oag: 
Cryo-EM structure of the plectasin fibril (single strand)
Method: helical / : Effantin G

EMDB-13776: 
Structure of formaldehyde cross-linked SARS-CoV-2 S glycoprotein
Method: single particle / : Sulbaran G, Effantin G, Schoehn G, Weissenhorn W

PDB-7q1z: 
Structure of formaldehyde cross-linked SARS-CoV-2 S glycoprotein
Method: single particle / : Sulbaran G, Effantin G, Schoehn G, Weissenhorn W

EMDB-13120: 
Mature capsid of bacteriophage phiRSA1
Method: single particle / : Effantin G, Fujiwara A, Kawsaki T, Yamada T, Schoehn G

PDB-7oz4: 
Mature capsid of bacteriophage phiRSA1
Method: single particle / : Effantin G, Fujiwara A, Kawsaki T, Yamada T, Schoehn G

EMDB-11179: 
Head of Semi-jumbo phage RP13
Method: single particle / : Neumann E, Kawasaki T, Effantin G, Estrozi L, Chatchawankanphanich O, Yamada T, Schoehn G

EMDB-11180: 
Head reconstruction of full jumbo phage XacN1
Method: single particle / : Neumann E, Kawasaki T, Effantin G, Estrozi L, Chatchawankanphanich O, Yamada T, Schoehn G

EMDB-10849: 
Inducible lysine decarboxylase LdcI decamer, pH 7.0
Method: single particle / : Jessop M, Felix J, Desfosses A, Effantin G, Gutsche I

EMDB-10850: 
Inducible lysine decarboxylase LdcI stacks, pH 5.7
Method: single particle / : Felix J, Jessop M, Desfosses A, Effantin G, Gutsche I

PDB-6yn5: 
Inducible lysine decarboxylase LdcI decamer, pH 7.0
Method: single particle / : Jessop M, Felix J, Desfosses A, Effantin G, Gutsche I
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
