-Search query
-Search result
Showing all 47 items for (author: dietrich & u)

EMDB-16451: 
Subtomogram average of the T. kivui 70S ribosome in situ
Method: subtomogram averaging / : Righetto RD, Dietrich HM, Kumar A, Wietrzynski W, Schuller SK, Trischler R, Wagner J, Schwarz FM, Mueller V, Schuller JM, Engel BD

EMDB-14169: 
Cryo-EM structure of Hydrogen-dependent CO2 reductase.
Method: single particle / : Dietrich HM, Righetto RD, Kumar A, Wietrzynski W, Schuller SK, Trischler R, Wagner J, Schwarz FM, Engel BD, Mueller V, Schuller JM

EMDB-15053: 
In situ structure of HDCR filaments
Method: subtomogram averaging / : Dietrich HM, Righetto RD, Kumar A, Wietrzynski W, Schuller SK, Trischler R, Wagner J, Schwarz FM, Engel BD, Mueller V, Schuller JM

EMDB-15054: 
In situ structure of the T. kivui ribosome
Method: subtomogram averaging / : Dietrich HM, Righetto RD, Kumar A, Wietrzynski W, Schuller SK, Trischler R, Wagner J, Schwarz FM, Engel BD, Mueller V, Schuller JM

EMDB-15055: 
In situ cryo-electron tomogram of a T. kivui cell 1
Method: electron tomography / : Dietrich HM, Righetto RD, Kumar A, Wietrzynski W, Schuller SK, Trischler R, Wagner J, Schwarz FM, Engel BD, Mueller V, Schuller JM

EMDB-15056: 
In situ cryo-electron tomogram of a T. kivui cell 2
Method: electron tomography / : Dietrich HM, Righetto RD, Kumar A, Wietrzynski W, Schuller SK, Trischler R, Wagner J, Schwarz FM, Engel BD, Mueller V, Schuller JM

PDB-7qv7: 
Cryo-EM structure of Hydrogen-dependent CO2 reductase.
Method: single particle / : Dietrich HM, Righetto RD, Kumar A, Wietrzynski W, Schuller SK, Trischler R, Wagner J, Schwarz FM, Engel BD, Mueller V, Schuller JM

EMDB-13246: 
cryoFIB milling/cryoET of amoeba infected with Legionella pneumophila
Method: electron tomography / : Boeck D, Huesler D

EMDB-13247: 
cryoFIB milling/cryoET of amoeba infected with Legionella pneumophila
Method: electron tomography / : Boeck D, Huesler D

EMDB-13248: 
cryoFIB milling/cryoET of amoeba infected with Legionella pneumophila
Method: electron tomography / : Boeck D, Huesler D

EMDB-13249: 
cryoFIB milling/cryoET of amoeba infected with Legionella pneumophila
Method: electron tomography / : Boeck D, Huesler D

EMDB-24642: 
SARS-CoV-2 Spike bound to Fab PDI 210
Method: single particle / : Pymm P, Glukhova A, Black K, Tham WH

EMDB-24643: 
SARS-CoV-2 Spike bound to Fab PDI 96
Method: single particle / : Pymm P, Glukhova A, Black K, Tham WH

EMDB-24644: 
SARS-CoV-2 Spike bound to Fab PDI 215
Method: single particle / : Black K, Glukhova A, Pymm P, Tham WH

EMDB-24645: 
SARS-CoV-2 Spike bound to Fab WCSL 119
Method: single particle / : Black K, Glukhova A, Pymm P, Tham WH

EMDB-24646: 
SARS-CoV-2 Spike bound to Fab WCSL 129
Method: single particle / : Black K, Glukhova A, Pymm P, Tham WH

EMDB-24647: 
SARS-CoV-2 Spike bound to Fab PDI 93
Method: single particle / : Black K, Glukhova A, Pymm P, Tham WH

EMDB-24648: 
SARS-CoV-2 Spike bound to Fab PDI 222
Method: single particle / : Glukhova A, Pymm P, Black K, Tham WH

EMDB-24649: 
SARS-CoV-2 receptor binding domain bound to Fab PDI 222
Method: single particle / : Pymm P, Glukhova A, Black KA, Tham WH

PDB-7rr0: 
SARS-CoV-2 receptor binding domain bound to Fab PDI 222
Method: single particle / : Pymm P, Glukhova A, Black KA, Tham WH

EMDB-11860: 
Subtomogram average structure of anammoxosomal nitrite oxidoreductase
Method: subtomogram averaging / : Dietrich L

EMDB-11861: 
Helical reconstruction of nitrite oxidoreductase from anammox bacteria
Method: helical / : Parey K

EMDB-23566: 
The SARS-CoV-2 spike protein receptor binding domain bound to neutralizing nanobodies WNb 2 and WNb 10
Method: single particle / : Pymm P, Glukhova A, Tham WH

EMDB-23567: 
Cryo-EM map of SARS-CoV-2 Spike protein bound to neutralising nanobodies WNb2 and WNb10 (C3 symmetry)
Method: single particle / : Pymm P, Tham WH, Glukhova A

EMDB-23568: 
Cryo-EM map of SARS-CoV-2 Spike protein in complex with neutralising nanobodies WNb2 and WNb10 (C1 symmetry)
Method: single particle / : Pymm P, Tham WH, Glukhova A

PDB-7lx5: 
The SARS-CoV-2 spike protein receptor binding domain bound to neutralizing nanobodies WNb 2 and WNb 10
Method: single particle / : Pymm P, Glukhova A, Tham WH

EMDB-11997: 
Structure of a nanoparticle for a COVID-19 vaccine candidate
Method: single particle / : Duyvesteyn HME, Stuart DI

PDB-7b3y: 
Structure of a nanoparticle for a COVID-19 vaccine candidate
Method: single particle / : Duyvesteyn HME, Stuart DI

EMDB-10840: 
Structure of RyR1 in apo/closed state in native membrane
Method: subtomogram averaging / : Chen W, Sanchez R, Zhang Y, Dietrich L, Kudryashev M

EMDB-20348: 
EM structure of human tumor suppressor BRCA2 protein bound to human DSS1 protein and ssDNA without crosslinking
Method: single particle / : Le HP, Ma X, Vaquero J, Brinkmeyer M, Guo F, Heyer WD, Liu J

EMDB-21998: 
EM structure of human tumor suppressor bound to human DSS1 protein and ssDNA with crosslinking
Method: single particle / : Le HP, Ma X, Vaquero J, Brinkmeyer M, Guo F, Heyer WD, Liu J

EMDB-10834: 
Tobacco mosaic virus (TMV) structure determined by a hybrid STA-SPA workflow
Method: subtomogram averaging / : Sanchez RM, Zhang Y, Chen W, Dietrich L, Kudryashev M

EMDB-4584: 
Structure and assembly of the mitochondrial membrane remodelling GTPase Mgm1
Method: subtomogram averaging / : Faelber K, Dietrich L, Noel J, Wollweber F, Pfitzner A, Muehleip A, Sanchez R, Kudryashev M, Chiaruttin N, Lilie H, Schleger J, Rosenbaum E, Hessenberger M, Matthaeus C, Noe F, Roux A, van der Laan M, Kuehlbrandt W, Daumke O
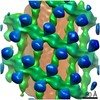
EMDB-10062: 
Structure of s-Mgm1 decorating the outer surface of tubulated lipid membranes
Method: subtomogram averaging / : Faelber K, Dietrich L, Noel JK, Sanchez R, Kudryashev M, Kuehlbrandt W, Daumke O
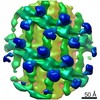
EMDB-10063: 
Structure of s-Mgm1 decorating the outer surface of tubulated lipid membranes in the GTPgammaS bound state
Method: subtomogram averaging / : Faelber K, Dietrich L, Noel JK, Sanchez R, Kudryashev M, Kuelbrandt W, Daumke O

EMDB-10064: 
Structure of s-Mgm1 decorating the inner surface of tubulated lipid membranes
Method: subtomogram averaging / : Faelber K, Dietrich L, Noel JK, Sanchez R, Kudryashev M, Kuelbrandt W, Daumke O

EMDB-10065: 
Structure of s-Mgm1 decorating the inner surface of tubulated lipid membranes in the GTPgammaS bound state
Method: subtomogram averaging / : Faelber K, Dietrich L, Noel JK, Sanchez R, Kudryashev M, Kuelbrandt W, Daumke O

PDB-6rzt: 
Structure of s-Mgm1 decorating the outer surface of tubulated lipid membranes
Method: subtomogram averaging / : Faelber K, Dietrich L, Noel JK, Sanchez R, Kudryashev M, Kuehlbrandt W, Daumke O

PDB-6rzu: 
Structure of s-Mgm1 decorating the outer surface of tubulated lipid membranes in the GTPgammaS bound state
Method: subtomogram averaging / : Faelber K, Dietrich L, Noel JK, Sanchez R, Kudryashev M, Kuelbrandt W, Daumke O

PDB-6rzv: 
Structure of s-Mgm1 decorating the inner surface of tubulated lipid membranes
Method: subtomogram averaging / : Faelber K, Dietrich L, Noel JK, Sanchez R, Kudryashev M, Kuelbrandt W, Daumke O

PDB-6rzw: 
Structure of s-Mgm1 decorating the inner surface of tubulated lipid membranes in the GTPgammaS bound state
Method: subtomogram averaging / : Faelber K, Dietrich L, Noel JK, Sanchez R, Kudryashev M, Kuelbrandt W, Daumke O
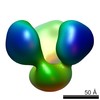
EMDB-2657: 
TEM of uncleaved soluble HIV envelope proteins
Method: single particle / : Arnold P, Himmels P, Weiss S, Decker TM, Markl J, Gatterdam V, Tampe R, Bartholomeaus P, Dietrich U, Duerr R

EMDB-2659: 
TEM of uncleaved soluble HIV envelope proteins
Method: single particle / : Arnold P, Himmels P, Weiss S, Decker TM, Markl J, Gatterdam V, Tampe R, Bartholomeaus P, Dietrich U, Duerr R
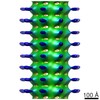
EMDB-2094: 
Central hub structure of Trichonympha cartwheel
Method: subtomogram averaging / : Guichard P, Desfosses A, Maheshwari A, Hachet V, Dietrich C, Brune A, Ishikawa T, Sachse C, Gonczy P

EMDB-1901: 
Insights into the structure of the CCR4-NOT Complex by electron microscopy
Method: single particle / : Nasertorabi F, Batisse C, Diepholz M, Suck D, Bottcher B

EMDB-5011: 
Crosslinked kinesin head-tail complex bound to the microtubule
Method: helical / : Dietrich KA, Sindelar CV, Brewer PD, Downing KH, Cremo CR, Rice SE

PDB-1mhs: 
Model of Neurospora crassa proton ATPase
Method: electron crystallography / : Kuhlbrandt W
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
