-Search query
-Search result
Showing 1 - 50 of 98 items for (author: chu & yk)

EMDB-19250: 
Pseudoatomic model of a second-order Sierpinski triangle formed by the citrate synthase from Synechococcus elongatus
Method: single particle / : Lo YK, Bohn S, Sendker FL, Schuller JM, Hochberg G

EMDB-19251: 
Structure of a first order Sierpinski triangle formed by the H369R mutant of the citrate synthase from Synechococcus elongatus
Method: single particle / : Lo YK, Bohn S, Sendker FL, Schuller JM, Hochberg G

EMDB-16004: 
Structure of hexameric subcomplexes (Truncation Delta2-6) of the fractal citrate synthase from Synechococcus elongatus PCC7942
Method: single particle / : Lo YK, Bohn S, Sendker FL, Schuller JM, Hochberg G

EMDB-15529: 
Structure of a first level Sierpinski triangle formed by a citrate synthase
Method: single particle / : Lo YK, Bohn S, Sendker FL, Schuller JM, Hochberg G

EMDB-42371: 
Mouse apoferritin imaged with a square aperture
Method: single particle / : Chua EYD, Alink LM, Kopylov M, Johnston J, Einsenstein F, de Marco A

EMDB-42372: 
Mouse apoferritin imaged with a round aperture
Method: single particle / : Chua EYD, Alink LM, Kopylov M, Johnston J, Einsenstein F, de Marco A

EMDB-42373: 
Mouse apoferritin imaged with a square aperture with P2 projection lens rotation, reconstructed from 25,000 particles
Method: single particle / : Chua EYD, Alink LM, Kopylov M, Johnston J, Einsenstein F, de Marco A

EMDB-42374: 
Mouse apoferritin imaged with a round aperture without P2 projection lens rotation, reconstructed from 25,000 particles
Method: single particle / : Chua EYD, Alink LM, Kopylov M, Johnston J, Einsenstein F, de Marco A

EMDB-36062: 
Asfv topoisomerase 2 - apo conformer Ia
Method: single particle / : Chang CW, Tsai MD

EMDB-36063: 
Asfv topoisomerase 2 - apo conformer Ib
Method: single particle / : Chang CW, Tsai MD

EMDB-36064: 
Cryo-EM structure of Asfv topoisomerase 2 - apo conformer IIa
Method: single particle / : Chang CW, Tsai MD

EMDB-36065: 
Cryo-EM structure of Asfv topoisomerase 2 - apo conformer IIb
Method: single particle / : Chang CW, Tsai MD

EMDB-36066: 
Cryo-EM structure of Asfv topoisomerase 2 - apo conformer IIIa
Method: single particle / : Chang CW, Tsai MD

EMDB-36067: 
Cryo-EM structure of Asfv topoisomerase 2 - apo conformer IIIb
Method: single particle / : Chang CW, Tsai MD

EMDB-36116: 
Cryo-EM structure of the African swine fever virus topoisomerase 2 complexed with Cut02aDNA and etoposide (EDI-1)
Method: single particle / : Chang CW, Tsai MD

EMDB-36117: 
Cryo-EM structure of the African swine fever virus topoisomerase 2 complexed with Cut02bDNA and etoposide (EDI-2)
Method: single particle / : Chang CW, Tsai MD

EMDB-36118: 
Cryo-EM structure of the African swine fever virus topoisomerase 2 complexed with Cut02aDNA and m-AMSA (EDI-3)
Method: single particle / : Chang CW, Tsai MD

EMDB-36473: 
Cryo-EM structure of the full-length African swine fever virus topoisomerase 2 complexed with Cut02aDNA and etoposide
Method: single particle / : Chang CW, Tsai MD

EMDB-42851: 
5x5 tiled montage tomogram of a holey carbon grid with apoferritin imaged with a square electron beam
Method: electron tomography / : Chua EYD, Alink LM, Kopylov M, Johnston J, Einsenstein F, de Marco A

EMDB-42879: 
3x3 tiled montage tomogram of a yeast lamella imaged with a square electron beam
Method: electron tomography / : Chua EYD, Alink LM, Kopylov M, Johnston J, Einsenstein F, de Marco A

EMDB-16024: 
SARS-CoV-2 S protein in complex with pT1644 Fab
Method: single particle / : Stroeh L, Hansen G, Vollmer B, Krey T, Benecke T, Gruenewald K

EMDB-16026: 
SARS-CoV-2 S protein in complex with pT1696 Fab
Method: single particle / : Hansen G, Benecke T, Vollmer B, Gruenewald K, Krey T

EMDB-41158: 
CS2it1p2_F7K local refinement for HCMV Trimer in complex with CS2it1p2_F7K Fab and CS4tt1p1_E3K Fab
Method: single particle / : Goldsmith JA, McLellan JS

EMDB-41180: 
Global reconstruction for HCMV Pentamer in complex with CS2pt1p2_A10L Fab and CS3pt1p4_C1L Fab
Method: single particle / : Goldsmith JG, McLellan JS
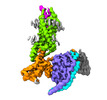
EMDB-33984: 
Complex structure of Neuropeptide Y Y2 receptor in complex with PYY(3-36) and Gi
Method: single particle / : Kang H, Park C, Kim J, Choi HJ

EMDB-33985: 
Complex structure of Neuropeptide Y Y2 receptor in complex with NPY and Gi
Method: single particle / : Kang H, Park C, Kim J, Choi HJ

EMDB-35494: 
Complex structure of Neuropeptide Y Y2 receptor in complex with NPY and Gi (Consensus map)
Method: single particle / : Kang H, Park C, Kim J, Choi HJ

EMDB-35495: 
Complex structure of Neuropeptide Y Y2 receptor in complex with NPY and Gi (Focused map on NPY-Y2R)
Method: single particle / : Kang H, Park C, Kim J, Choi HJ

EMDB-35496: 
Complex structure of Neuropeptide Y Y2 receptor in complex with NPY and Gi (Focused map on Gi-scFv16)
Method: single particle / : Kang H, Park C, Kim J, Choi HJ

EMDB-35497: 
Complex structure of Neuropeptide Y Y2 receptor in complex with PYY(3-36) and Gi (Consensus map)
Method: single particle / : Kang H, Park C, Kim J, Choi HJ

EMDB-35498: 
Complex structure of Neuropeptide Y Y2 receptor in complex with PYY(3-36) and Gi (Focused map on PYY(3-36)-Y2R)
Method: single particle / : Kang H, Park C, Kim J, Choi HJ

EMDB-35499: 
Complex structure of Neuropeptide Y Y2 receptor in complex with PYY(3-36) and Gi (Focused map on Gi-scFv16)
Method: single particle / : Kang H, Park C, Kim J, Choi HJ

EMDB-13485: 
Cryo-EM structure of Bestrhodopsin (rhodopsin-rhodopsin-bestrophin) complex
Method: single particle / : Matzov D, Kaczmarczyk I, Shalev-Benami M

PDB-7pl9: 
Cryo-EM structure of Bestrhodopsin (rhodopsin-rhodopsin-bestrophin) complex
Method: single particle / : Matzov D, Kaczmarczyk I, Shalev-Benami M

EMDB-14250: 
Structure of the SARS-CoV-2 spike glycoprotein in complex with the 87G7 antibody Fab fragment
Method: single particle / : Hurdiss DL, Drulyte I

EMDB-14271: 
Structure of the SARS-CoV-2 spike glycoprotein in complex with the 87G7 antibody Fab fragment (local refinement)
Method: single particle / : Hurdiss DL, Drulyte I

PDB-7r40: 
Structure of the SARS-CoV-2 spike glycoprotein in complex with the 87G7 antibody Fab fragment
Method: single particle / : Hurdiss DL

EMDB-30262: 
DXPS
Method: single particle / : Lau WCY

EMDB-23719: 
EBOV GP bound to rEBOV-442 and rEBOV-515 Fabs
Method: single particle / : Murin CD, Ward AB

PDB-7m8l: 
EBOV GP bound to rEBOV-442 and rEBOV-515 Fabs
Method: single particle / : Murin CD, Ward AB

EMDB-11588: 
Structure of low-light grown Chlorella ohadii Photosystem I
Method: single particle / : Caspy I, Nelson N, Nechushtai R, Neumann E, Shkolnisky Y

EMDB-11589: 
Structure of high-light grown Chlorella ohadii photosystem I
Method: single particle / : Caspy I, Nelson N, Nechushtai R, Shkolnisky Y, Neumann E

EMDB-11640: 
Structure of small high-light grown Chlorella ohadii photosystem I
Method: single particle / : Caspy I, Nelson N

PDB-6zzx: 
Structure of low-light grown Chlorella ohadii Photosystem I
Method: single particle / : Caspy I, Nelson N, Nechushtai R, Neumann E, Shkolnisky Y

PDB-6zzy: 
Structure of high-light grown Chlorella ohadii photosystem I
Method: single particle / : Caspy I, Nelson N, Nechushtai R, Shkolnisky Y, Neumann E

PDB-7a4p: 
Structure of small high-light grown Chlorella ohadii photosystem I
Method: single particle / : Caspy I, Nelson N, Nechushtai R, Shkolnisky Y, Neumann E

EMDB-30263: 
Hsp21-DXPS
Method: single particle / : Lau WCY

EMDB-22839: 
BDBV-289 bound to EBOV GPdMuc Makona
Method: single particle / : Murin CD, Ward AB

EMDB-12335: 
Structure of PSII-M
Method: single particle / : Zabret J, Bohn S, Schuller SK, Arnolds O, Chan A, Tajkhorshid E, Stoll R, Engel BD, Rudack T, Schuller JM, Nowaczyk MM
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search




 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
