-Search query
-Search result
Showing 1 - 50 of 129 items for (author: cheng & rh)
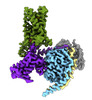
EMDB-43647: 
CryoEM structure of Gi-coupled TAS2R14 with cholesterol and an intracellular tastant
Method: single particle / : Kim Y, Gumpper RH, Roth BL
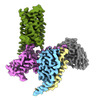
EMDB-43650: 
CryoEM structure of Ggust-coupled TAS2R14 with cholesterol and an intracellular tastant
Method: single particle / : Kim Y, Gumpper RH, Roth BL

EMDB-43656: 
CryoEM structure of Gi-coupled TAS2R14 with cholesterol and an intracellular tastant (Locally refined map)
Method: single particle / : Kim Y, Gumpper RH, Roth BL

EMDB-43657: 
CryoEM structure of Ggust-coupled TAS2R14 with cholesterol and an intracellular tastant (Locally refined map)
Method: single particle / : Kim Y, Gumpper RH, Roth BL

EMDB-40711: 
CryoEM structure of Western equine encephalitis virus VLP in complex with the chimeric Du-D1-Mo-D2 MXRA8 receptor
Method: single particle / : Zimmerman MI, Fremont DH, Center for Structural Genomics of Infectious Diseases (CSGID)
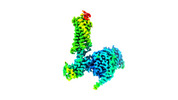
EMDB-29301: 
Neurotensin receptor allosterism revealed in complex with a biased allosteric modulator
Method: single particle / : Krumm BE, Diberto JF, Olsen RHJ, Kang H, Slocum ST, Zhang S, Strachan RT, Fay JF, Roth BL

EMDB-29302: 
CryoEM structure of Go-coupled NTSR1 with a biased allosteric modulator
Method: single particle / : Krumm BE, DiBerto JF, Olsen RHJ, Kang H, Slocum ST, Zhang S, Strachan RT, Fay JF, Roth BL
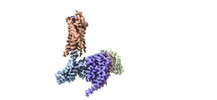
EMDB-29303: 
CryoEM structure of Go-coupled NTSR1
Method: single particle / : Krumm BE, DiBerto JF, Olsen RHJ, Kang H, Slocum ST, Zhang S, Strachan RT, Fay JF, Roth BL

EMDB-27966: 
CryoEM structure of miniGq-coupled hM3Dq in complex with DCZ
Method: single particle / : Zhang S, Fay JF, Roth BL

EMDB-27967: 
CryoEM structure of miniGo-coupled hM4Di in complex with DCZ
Method: single particle / : Zhang S, Fay JF, Roth BL

EMDB-27968: 
CryoEM structure of miniGq-coupled hM3Dq in complex with CNO
Method: single particle / : Zhang S, Fay JF, Roth BL

EMDB-27969: 
CryoEM structure of miniGq-coupled hM3R in complex with Iperoxo
Method: single particle / : Zhang S, Fay JF, Roth BL

EMDB-27970: 
CryoEM structure of miniGq-coupled hM3R in complex with iperoxo (local refinement)
Method: single particle / : Zhang S, Fay JF, Roth BL

EMDB-27752: 
CryoEM structure of Gq-coupled MRGPRX1 with peptide agonist BAM8-22
Method: single particle / : Liu Y, Cao C, Fay JF, Roth BL

EMDB-27753: 
CryoEM structure of Gq-coupled MRGPRX1 with peptide ligand BAM8-22 and positive allosteric modulator ML382
Method: single particle / : Liu Y, Cao C, Fay JF, Roth BL

EMDB-27754: 
CryoEM structure of Gq-coupled MRGPRX1 with ligand Compound-16
Method: single particle / : Liu Y, Cao C, Fay JF, Roth BL

EMDB-25401: 
5-HT2B receptor bound to LSD obtained by cryo-electron microscopy (cryoEM)
Method: single particle / : Barros-Alvarez X, Cao C, Panova O, Roth BL, Skiniotis G

EMDB-25402: 
5-HT2B receptor bound to LSD in complex with heterotrimeric mini-Gq protein obtained by cryo-electron microscopy (cryoEM)
Method: single particle / : Barros-Alvarez X, Kim K, Panova O, Cao C, Roth BL, Skiniotis G

EMDB-25403: 
5-HT2B receptor bound to LSD in complex with beta-arrestin1 obtained by cryo-electron microscopy (cryoEM)
Method: single particle / : Barros-Alvarez X, Cao C, Panova O, Roth BL, Skiniotis G

EMDB-27864: 
Escherichia coli Rho-dependent transcription pre-termination complex containing 18 nt long RNA spacer, Mg-ADP-BeF3, and NusG; TEC part
Method: single particle / : Molodtsov V, Wang C

EMDB-27865: 
Escherichia coli Rho-dependent transcription pre-termination complex containing 18 nt long RNA spacer, Mg-ADP-BeF3, and NusG; Rho hexamer part
Method: single particle / : Molodtsov V, Wang C

EMDB-27897: 
Escherichia coli Rho-dependent transcription pre-termination complex containing 18 nt long RNA spacer, Mg-ADP-BeF3, and NusG
Method: single particle / : Molodtsov V, Wang C, Ebright RH

EMDB-27913: 
Escherichia coli Rho-dependent transcription pre-termination complex containing 21 nt long RNA spacer, Mg-ADP-BeF3, and NusG; TEC part
Method: single particle / : Molodtsov V, Wang C

EMDB-27914: 
Escherichia coli Rho-dependent transcription pre-termination complex containing 21 nt long RNA spacer, Mg-ADP-BeF3, and NusG; Rho hexamer part
Method: single particle / : Molodtsov V, Wang C

EMDB-27915: 
Escherichia coli Rho-dependent transcription pre-termination complex containing 21 nt long RNA spacer, Mg-ADP-BeF3, and NusG
Method: single particle / : Molodtsov V, Wang C, Ebright RH

EMDB-27916: 
Escherichia coli Rho-dependent transcription pre-termination complex containing 24 nt long RNA spacer, Mg-ADP-BeF3, and NusG; TEC part
Method: single particle / : Molodtsov V, Wang C

EMDB-27917: 
Escherichia coli Rho-dependent transcription pre-termination complex containing 24 nt long RNA spacer, Mg-ADP-BeF3, and NusG; Rho hexamer part
Method: single particle / : Molodtsov V, Wang C

EMDB-27918: 
Escherichia coli Rho-dependent transcription pre-termination complex containing 24 nt long RNA spacer, Mg-ADP-BeF3, and NusG
Method: single particle / : Molodtsov V, Wang C, Ebright RH

EMDB-27928: 
Escherichia coli Rho-dependent transcription pre-termination complex containing 18 nt long RNA spacer, lambda-tR1 rut RNA, Mg-ADP-BeF3, and NusG; Rho part
Method: single particle / : Molodtsov V, Wang C, Ebright RH

EMDB-27929: 
Escherichia coli Rho-dependent transcription pre-termination complex containing 18 nt long RNA spacer, lambda-tR1 rut RNA, Mg-ADP-BeF3, and NusG
Method: single particle / : Molodtsov V, Wang C, Ebright RH

EMDB-27930: 
Escherichia coli Rho-dependent transcription pre-termination complex containing 18 nt long RNA spacer, lambda-tR1 rut RNA, Mg-ADP-BeF3, and NusG; TEC part
Method: single particle / : Molodtsov V, Wang C

EMDB-27931: 
Escherichia coli Rho-dependent transcription pre-termination complex containing 18 nt long RNA spacer, dC75 rut mimic RNA, Mg-ADP-BeF3, and NusG; TEC part
Method: single particle / : Molodtsov V, Wang C, Ebright RH

EMDB-27932: 
Escherichia coli Rho-dependent transcription pre-termination complex containing 18 nt long RNA spacer, dC75 rut mimic RNA, Mg-ADP-BeF3, and NusG; Rho hexamer part
Method: single particle / : Molodtsov V, Wang C, Ebright RH

EMDB-27933: 
Escherichia coli Rho-dependent transcription pre-termination complex containing 18 nt long RNA spacer, dC75 rut mimic RNA
Method: single particle / : Molodtsov V, Wang C, Ebright RH

EMDB-13604: 
Apo human Kv3.1 cryo-EM structure
Method: single particle / : Botte M, Huber S, Bucher D, Klint JK, Rodriguez D, Tagmose L, Chami M, Cheng R, Hennig M, Abdul Rhaman W

EMDB-13605: 
Ligand-bound human Kv3.1 cryo-EM structure (Lu AG00563)
Method: single particle / : Botte M, Huber S, Bucher D, Klint JK, Rodriguez D, Tagmose L, Chami M, Cheng R, Hennig M, Abdul Rhaman W

PDB-7pqt: 
Apo human Kv3.1 cryo-EM structure
Method: single particle / : Botte M, Huber S, Bucher D, Klint JK, Rodriguez D, Tagmose L, Chami M, Cheng R, Hennig M, Abdul Rhaman W

PDB-7pqu: 
Ligand-bound human Kv3.1 cryo-EM structure (Lu AG00563)
Method: single particle / : Botte M, Huber S, Bucher D, Klint JK, Rodriguez D, Tagmose L, Chami M, Cheng R, Hennig M, Abdul Rhaman W
EMDB-33202: 
S-ECD (Omicron) in complex with STS165
Method: single particle / : Li YN, Shen YP, Zhang YY, Yan RH
EMDB-33203: 
S-ECD (Omicron) in complex with PD of ACE2
Method: single particle / : Li YN, Shen YP, Zhang YY, Yan RH

EMDB-33204: 
S-ECD (Omicron) in complex with PD of ACE2 focused on RBD_PD sub-complex
Method: single particle / : Li YN, Shen YP, Zhang YY, Yan RH

EMDB-24896: 
CryoEM structure of Gq-coupled MRGPRX2 with peptide agonist Cortistatin-14
Method: single particle / : Cao C, Fay JF, Gumpper RH, Roth BL

EMDB-24897: 
CryoEM structure of Gi-coupled MRGPRX2 with peptide agonist Cortistatin-14
Method: single particle / : Cao C, Fay JF, Gumpper RH, Roth BL

EMDB-24898: 
CryoEM structure of Gq-coupled MRGPRX2 with small molecule agonist (R)-Zinc-3573
Method: single particle / : Cao C, Fay JF, Gumpper RH, Roth BL

EMDB-24899: 
CryoEM structure of Gi-coupled MRGPRX2 with small molecule agonist (R)-Zinc-3573
Method: single particle / : Cao C, Fay JF, Gumpper RH, Roth BL

EMDB-24900: 
CryoEM structure of Gq-coupled MRGPRX4 with small molecule agonist MS47134
Method: single particle / : Cao C, Fay JF, Gumpper RH, Roth BL

EMDB-22495: 
Cryo-EM Structure of CVB3 VLP
Method: single particle / : Baikoghli MA, Cheng RH

EMDB-30512: 
S protein of SARS-CoV-2 in complex bound with P2B-1A1
Method: single particle / : Yan RH, Zhang YY, Li YN, Zhou Q

EMDB-30513: 
cryo EM map of the S protein of SARS-CoV-2 in complex bound with P2B-1A10
Method: single particle / : Yan RH, Zhang YY, Li YN, Zhou Q

EMDB-30514: 
S protein of SARS-CoV-2 in complex bound with P5A-1B8_2B
Method: single particle / : Yan RH, Zhang YY, Li YN, Zhou Q
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
