-Search query
-Search result
Showing 1 - 50 of 7,274 items for (author: chen & f)

EMDB-45962: 
Cryo-EM structure of Tulane virus 9-6-17 variant capsid protein VP1 5-12-18
Method: single particle / : Sun C, Jiang W

EMDB-45963: 
Cryo-EM structure of Tulane virus 9-6-17 variant capsid protein VP1 9-14-18
Method: single particle / : Sun C, Jiang W

EMDB-45964: 
Cryo-EM structure of Tulane virus 9-6-17 variant capsid protein VP1 9-14-18, DTT-treated
Method: single particle / : Sun C, Jiang W
EMDB-36364: 
G protein-coupled receptor 1
Method: single particle / : Liu A, Liu Y, Chen G, Ye F

EMDB-41542: 
Polyclonal immune complex of Fab binding the H2 HA from serum of subject 3-3 at week 4
Method: single particle / : Yang YR, Han J, Richey ST, Ward AB

EMDB-38845: 
Icosahedrally averaged cryo-EM reconstruction of PhiKZ capsid before applying the "block-based" reconstruction method
Method: single particle / : Yang Y, Shao Q, Guo M, Han L, Zhao X, Wang A, Li X, Wang B, Pan J, Chen Z, Fokine A, Sun L, Fang Q

EMDB-38846: 
Block 1 of PhiKZ capsid
Method: single particle / : Yang Y, Shao Q, Guo M, Han L, Zhao X, Wang A, Li X, Wang B, Pan J, Chen Z, Fokine A, Sun L, Fang Q

EMDB-38848: 
Block 2 of PhiKZ capsid
Method: single particle / : Yang Y, Shao Q, Guo M, Han L, Zhao X, Wang A, Li X, Wang B, Pan J, Chen Z, Fokine A, Sun L, Fang Q

EMDB-39002: 
Composite cryo-EM map of PhiKZ capsid after applying the "block-based" reconstruction method
Method: single particle / : Yang Y, Shao Q, Guo M, Han L, Zhao X, Wang A, Li X, Wang B, Pan J, Chen Z, Fokine A, Sun L, Fang Q

PDB-8y6v: 
Near-atomic structure of icosahedrally averaged jumbo bacteriophage PhiKZ capsid
Method: single particle / : Yang Y, Shao Q, Guo M, Han L, Zhao X, Wang A, Li X, Wang B, Pan J, Chen Z, Fokine A, Sun L, Fang Q

EMDB-41770: 
Apo form of human ATE1
Method: single particle / : Huang W, Zhang Y, Taylor DJ

EMDB-42071: 
human ATE1 in complex with Arg-tRNA and a peptide substrate
Method: single particle / : Huang W, Zhang Y, Taylor DJ

PDB-8uau: 
human ATE1 in complex with Arg-tRNA and a peptide substrate
Method: single particle / : Huang W, Zhang Y, Taylor DJ
EMDB-44246: 
Cryo-EM structure of HIV-1 JRFL v6 Env in complex with vaccine-elicited, Membrane Proximal External Region (MPER) directed antibody DH1317.4.
Method: single particle / : Acharya P, Parsons R, Janowska K, Williams WB, Alam M, Haynes BF
EMDB-37139: 
Structure of SARS-CoV Spike protein complexed with antibody PW5-5
Method: single particle / : Sun L, Mao Q, Wang Y
EMDB-37143: 
The local refined map of SARS-CoV-2 XBB Variant Spike protein complexed with antibody PW5-535
Method: single particle / : Sun L, Mao Q, Wang Y
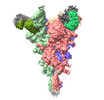
EMDB-37144: 
Trimer state of SARS-CoV Spike protein complexed with antibody PW5-535
Method: single particle / : Sun L, Mao Q, Wang Y
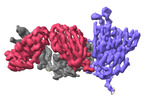

EMDB-37145: 
The local refined map of SARS-CoV Spike protein complexed with antibody PW5-5
Method: single particle / : Sun L, Mao Q, Wang Y
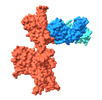
EMDB-37160: 
Monomer state of SARS-CoV-2 XBB Variant Spike protein trimer complexed with antibody PW5-5
Method: single particle / : Sun L, Mao Q, Wang Y
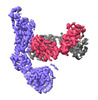
EMDB-37161: 
Monomer state of SARS-CoV Spike protein complexed with antibody PW5-535
Method: single particle / : Sun L, Mao Q, Wang Y
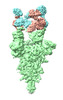
EMDB-37162: 
Structure of SARS-CoV-2 Omicron BA.1 Spike complexed with antibody PW5-570
Method: single particle / : Sun L, Mao Q, Wang Y
EMDB-37163: 
The local refined map of SARS-CoV-2 Omicron BA.1 Spike complexed with antibody PW5-570
Method: single particle / : Sun L, Mao Q, Wang Y
EMDB-37164: 
State 1 of SARS-CoV-2 XBB Variant Spike protein trimer complexed with antibody PW5-5
Method: single particle / : Sun L, Mao Q, Wang Y
EMDB-37165: 
Structure of SARS-CoV-2 XBB Variant Spike protein complexed with broadly neutralizing antibody PW5-535
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8kdm: 
Structure of SARS-CoV Spike protein complexed with antibody PW5-5
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8kdr: 
The local refined map of SARS-CoV-2 XBB Variant Spike protein complexed with antibody PW5-535
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8kds: 
Trimer state of SARS-CoV Spike protein complexed with antibody PW5-535
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8kdt: 
The local refined map of SARS-CoV Spike protein complexed with antibody PW5-5
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8kej: 
Monomer state of SARS-CoV-2 XBB Variant Spike protein trimer complexed with antibody PW5-5
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8kek: 
Monomer state of SARS-CoV Spike protein complexed with antibody PW5-535
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8keo: 
Structure of SARS-CoV-2 Omicron BA.1 Spike complexed with antibody PW5-570
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8kep: 
The local refined map of SARS-CoV-2 Omicron BA.1 Spike complexed with antibody PW5-570
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8keq: 
State 1 of SARS-CoV-2 XBB Variant Spike protein trimer complexed with antibody PW5-5
Method: single particle / : Sun L, Mao Q, Wang Y

PDB-8ker: 
Structure of SARS-CoV-2 XBB Variant Spike protein complexed with broadly neutralizing antibody PW5-535
Method: single particle / : Sun L, Mao Q, Wang Y

EMDB-17696: 
Structure of human 48S translation initiation complex in open codon scanning state (48S-1)
Method: single particle / : Petrychenko V, Yi SH, Liedtke D, Peng BZ, Rodnina MV, Fischer N

EMDB-17697: 
Structure of human 48S translation initiation complex in AUG recognition state after eIF5-induced GTP hydrolysis by eIF2 (48S-2)
Method: single particle / : Petrychenko V, Yi SH, Liedtke D, Peng BZ, Rodnina MV, Fischer N

EMDB-17698: 
Structure of human 48S translation initiation complex upon transfer of initiator tRNA to eIF5B (48S-3)
Method: single particle / : Petrychenko V, Yi SH, Liedtke D, Peng BZ, Rodnina MV, Fischer N

EMDB-17699: 
Structure of human 48S translation initiation complex after eIF5 release (48S-4)
Method: single particle / : Petrychenko V, Yi SH, Liedtke D, Peng BZ, Rodnina MV, Fischer N

EMDB-17700: 
Structure of human 48S translation initiation complex after eIF2 release prior 60S subunit joining (48S-5)
Method: single particle / : Petrychenko V, Yi SH, Liedtke D, Peng BZ, Rodnina MV, Fischer N

EMDB-17701: 
Structure of human 48S translation initiation complex with initiator tRNA, eIF1A and eIF3 (off-pathway)
Method: single particle / : Petrychenko V, Yi SH, Liedtke D, Peng BZ, Rodnina MV, Fischer N

EMDB-19128: 
Structure of human eIF3 core from closed 48S translation initiation complex
Method: single particle / : Petrychenko V, Yi SH, Liedtke D, Peng BZ, Rodnina MV, Fischer N

PDB-8pj1: 
Structure of human 48S translation initiation complex in open codon scanning state (48S-1)
Method: single particle / : Petrychenko V, Yi SH, Liedtke D, Peng BZ, Rodnina MV, Fischer N

PDB-8pj2: 
Structure of human 48S translation initiation complex in AUG recognition state after eIF5-induced GTP hydrolysis by eIF2 (48S-2)
Method: single particle / : Petrychenko V, Yi SH, Liedtke D, Peng BZ, Rodnina MV, Fischer N

PDB-8pj3: 
Structure of human 48S translation initiation complex upon transfer of initiator tRNA to eIF5B (48S-3)
Method: single particle / : Petrychenko V, Yi SH, Liedtke D, Peng BZ, Rodnina MV, Fischer N

PDB-8pj4: 
Structure of human 48S translation initiation complex after eIF5 release (48S-4)
Method: single particle / : Petrychenko V, Yi SH, Liedtke D, Peng BZ, Rodnina MV, Fischer N

PDB-8pj5: 
Structure of human 48S translation initiation complex after eIF2 release prior 60S subunit joining (48S-5)
Method: single particle / : Petrychenko V, Yi SH, Liedtke D, Peng BZ, Rodnina MV, Fischer N

PDB-8pj6: 
Structure of human 48S translation initiation complex with initiator tRNA, eIF1A and eIF3 (off-pathway)
Method: single particle / : Petrychenko V, Yi SH, Liedtke D, Peng BZ, Rodnina MV, Fischer N

PDB-8rg0: 
Structure of human eIF3 core from closed 48S translation initiation complex
Method: single particle / : Petrychenko V, Yi SH, Liedtke D, Peng BZ, Rodnina MV, Fischer N

EMDB-39299: 
Human resource SGLT1-MAP17 complex
Method: single particle / : Chen L, Zhang XZ
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search




 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
