-Search query
-Search result
Showing 1 - 50 of 154 items for (author: bowman & v)

EMDB-36766: 
Untilted (0 Degree Tilted) Single-Particle CryoEM Reconstruction of AAV2 Viral Capsid
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36767: 
10 Degree Tilted Single-Particle CryoEM Reconstruction of AAV2 Viral Capsid
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36768: 
20 Degree Tilted Single-Particle CryoEM Reconstruction of AAV2 Viral Capsid
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36769: 
30 Degree Tilted Single-Particle CryoEM Reconstruction of AAV2 Viral Capsid
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36770: 
40 Degree Tilted Single-Particle CryoEM Reconstruction of AAV2 Viral Capsid
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36771: 
50 Degree Tilted Single-Particle CryoEM Reconstruction of AAV2 Viral Capsid
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36772: 
60 Degree Tilted Single-Particle CryoEM Reconstruction of AAV2 Viral Capsid
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36807: 
Untilted (0 Degree Tilted) Single-Particle CryoEM Reconstruction of Apoferritin
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36809: 
10 Degree Tilted Single-Particle CryoEM Reconstruction of Apoferritin
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D
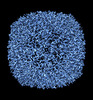
EMDB-36810: 
20 Degree Tilted Single-Particle CryoEM Reconstruction of Apoferritin
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36811: 
30 Degree Tilted Single-Particle CryoEM Reconstruction of Apoferritin
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36812: 
40 Degree Tilted Single-Particle CryoEM Reconstruction of Apoferritin
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36813: 
60 Degree Tilted Single-Particle CryoEM Reconstruction of Apoferritin
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D
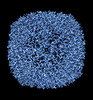
EMDB-36814: 
50 Degree Tilted Single-Particle CryoEM Reconstruction of Apoferritin
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36816: 
Untilted (0 Degree Tilted) Single-Particle CryoEM Reconstruction of DPS
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36817: 
10 Degree Tilted Single-Particle CryoEM Reconstruction of DPS
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36818: 
20 Degree Tilted Single-Particle CryoEM Reconstruction of DPS
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36819: 
30 Degree Tilted Single-Particle CryoEM Reconstruction of DPS
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36820: 
40 Degree Tilted Single-Particle CryoEM Reconstruction of DPS
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36821: 
50 Degree Tilted Single-Particle CryoEM Reconstruction of DPS
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36822: 
60 Degree Tilted Single-Particle CryoEM Reconstruction of DPS
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-41230: 
Untilted (0 Degree Tilted) Single-Particle CryoEM Reconstruction of Apoferritin
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-41231: 
30 Degree Tilted Single-Particle CryoEM Reconstruction of Apoferritin
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-41695: 
60 Degree Tilted Single-Particle CryoEM Reconstruction of RNA polymerase
Method: single particle / : Aiyer S, Shan Z, Oh J, Wang D, Tan YZ, Lyumkis D

PDB-8txo: 
E. coli DNA-directed RNA polymerase transcription elongation complex bound to the unnatural dZ-PTP base pair in the active site
Method: single particle / : Aiyer S, Shan Z, Oh J, Wang D, Tan YZ, Lyumkis D
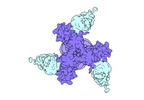
EMDB-27706: 
Vaccine elicited Antibody MU89 bound to CH848.D949.10.17_N133D_N138T.DS.SOSIP.664 HIV-1 Env trimer
Method: single particle / : Stalls V, Acharya P
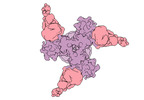
EMDB-27776: 
Vaccine elicited Antibody MU89+S27Y bound to CH848.D949.10.17_N133D_N138T.DS.SOSIP.664 HIV-1 Env trimer
Method: single particle / : Stalls V, Acharya P
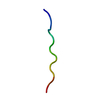
PDB-7t3h: 
MicroED structure of Dynobactin
Method: electron crystallography / : Yoo BK, Kaiser JT, Rees DC, Miller RD, Iinishi A, Lewis K, Bowman S

EMDB-14242: 
E. coli BAM complex (BamABCDE) bound to dynobactin A
Method: single particle / : Jakob RP, Hiller S

PDB-7r1w: 
E. coli BAM complex (BamABCDE) bound to dynobactin A
Method: single particle / : Jakob RP, Hiller S, Maier T

EMDB-23778: 
Negative stain EM map of BG505 SOSIP in complex with Rh4O9.8 monoclonal antibody (as Fab fragment)
Method: single particle / : Antanasijevic A, Ward AB

EMDB-23779: 
BG505 SOSIP.v5.2 in complex with the polyclonal Fab pAbC-1 from animal Rh.4O9
Method: single particle / : Antanasijevic A, Ozorowski G, Nogal B, Ward AB

EMDB-23780: 
BG505 SOSIP MD39 in complex with the monoclonal antibodies Rh.33104 mAb.1 and RM20A3
Method: single particle / : Antanasijevic A, Ward AB

EMDB-23801: 
BG505 SOSIP.v5.2(7S) in complex with the monoclonal antibodies Rh.33172 mAb.1 and RM19R
Method: single particle / : Antanasijevic A, Ward AB

PDB-7mdt: 
BG505 SOSIP.v5.2 in complex with the monoclonal antibody Rh4O9.8 (as Fab fragment)
Method: single particle / : Antanasijevic A, Ozorowski G, Nogal B, Ward AB

PDB-7mdu: 
BG505 SOSIP MD39 in complex with the monoclonal antibodies Rh.33104 mAb.1 and RM20A3
Method: single particle / : Antanasijevic A, Ward AB

PDB-7mep: 
BG505 SOSIP.v5.2(7S) in complex with the monoclonal antibodies Rh.33172 mAb.1 and RM19R
Method: single particle / : Antanasijevic A, Ward AB

EMDB-21061: 
RM19M Fab in complex with BG505 SOSIPv5.2
Method: single particle / : Cottrell CA, Torres JL, Patel R, Nogal B, Ward AB

EMDB-21062: 
RM19A1 Fab in complex with BG505 SOSIPv5.2
Method: single particle / : Cottrell CA, Ozorowski G, Ward AB

EMDB-21064: 
RM19P Fab in complex with BG505 SOSIPv5.2
Method: single particle / : Cottrell CA, Ozorowski G, Ward AB

EMDB-21065: 
RM19O Fab in complex with BG505 SOSIPv5.2
Method: single particle / : Cottrell CA, Torres JL, Patel R, Ward AB

EMDB-21066: 
RM19L Fab in complex with BG505 SOSIPv5.2
Method: single particle / : Cottrell CA, Torres JL, Patel R, Ward AB
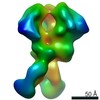
EMDB-21075: 
RM19B Fab in complex with BG505 SOSIPv5.2
Method: single particle / : Cottrell CA, Torres JL, Patel R, Ward AB

EMDB-21076: 
RM19K Fab in complex with BG505 SOSIPv5.2
Method: single particle / : Cottrell CA, Torres JL, Patel R, Ward AB

EMDB-21077: 
RM19B1 Fab in complex with BG505 SOSIPv5.2
Method: single particle / : Cottrell CA, Torres JL, Patel R, Ward AB

EMDB-21078: 
RM19C Fab in complex with BG505 SOSIPv5.2
Method: single particle / : Cottrell CA, Torres JL, Patel R, Ward AB
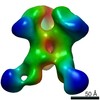
EMDB-21079: 
RM19C2 Fab in complex with BG505 SOSIPv5.2
Method: single particle / : Cottrell CA, Torres JL, Patel R, Ward AB
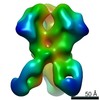
EMDB-21080: 
RM19E Fab in complex with BG505 SOSIPv5.2
Method: single particle / : Cottrell CA, Torres JL, Patel R, Ward AB

EMDB-21081: 
RM19G Fab in complex with BG505 SOSIPv5.2
Method: single particle / : Cottrell CA, Torres JL, Patel R, Ward AB
EMDB-21082: 
RM19C3 Fab in complex with BG505 SOSIPv5.2
Method: single particle / : Cottrell CA, Torres JL, Patel R, Ward AB
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
