-Search query
-Search result
Showing 1 - 50 of 95 items for (author: bhattacharya & n)

EMDB-43013: 
Myxococcus xanthus HEnc-K417N(A) protein shell with icosahedral T=1 symmetry
Method: single particle / : Hernandez C, Jenkins MC, Kopylov M

EMDB-43016: 
Myxococcus xanthus HEnc-K417N(A) protein shell with tetrahedral symmetry (12 pentamers, 4 hexamers)
Method: single particle / : Hernandez C, Jenkins MC, Kopylov M

EMDB-43037: 
Myxococcus xanthus HEnc-K417N(A) protein shell with D3 symmetry (12 pentamers, 3 hexamers)
Method: single particle / : Hernandez C, Jenkins MC, Kopylov M

EMDB-43038: 
Myxococcus xanthus HEnc-K417N(A) protein shell with D6 symmetry (12 pentamers, 8 hexamers)
Method: single particle / : Hernandez C, Jenkins MC, Kopylov M

EMDB-43039: 
Myxococcus xanthus HEnc-K417N(A) protein shell with C2 symmetry (12 pentamers, 9 hexamers)
Method: single particle / : Hernandez C, Jenkins MC, Kopylov M

EMDB-43040: 
Myxococcus xanthus HEnc-K417N(A) protein shell with D3 symmetry (12 pentamers, 11 hexamers)
Method: single particle / : Hernandez C, Jenkins MC, Kopylov M

EMDB-43041: 
Myxococcus xanthus HEnc-K417N(A) protein shell with D2 symmetry (12 pentamers, 12 hexamers)
Method: single particle / : Hernandez C, Jenkins MC, Kopylov M

EMDB-43042: 
Myxococcus xanthus HEnc-K417N(A) protein shell with D2 symmetry (12 pentamers, 14 hexamers)
Method: single particle / : Hernandez C, Jenkins MC, Kopylov M

EMDB-43043: 
Myxococcus xanthus HEnc-K417N(A) protein shell with D5 symmetry (12 pentamers, 15 hexamers)
Method: single particle / : Hernandez C, Jenkins MC, Kopylov M

EMDB-43113: 
Myxococcus xanthus EncA WT protein shell with icosahedral symmetry T=3
Method: single particle / : Hernandez C, Jenkins MC, Kopylov M

EMDB-43036: 
Myxococcus xanthus HEnc-K417N(A) protein shell with icosahedral T=3 symmetry
Method: single particle / : Hernandez C, Jenkins MC, Kopylov M

EMDB-26390: 
SAAV pH 7.4 capsid structure
Method: single particle / : Mietzsch M, McKenna R

EMDB-26391: 
SAAV pH 6.0 capsid structure
Method: single particle / : Mietzsch M, McKenna R

EMDB-26392: 
SAAV pH 5.5 capsid structure
Method: single particle / : Mietzsch M, McKenna R

EMDB-26393: 
SAAV pH 4.0 capsid structure
Method: single particle / : Mietzsch M, McKenna R

EMDB-26394: 
The SAAV capsid in complex with 3'SLN
Method: single particle / : Mietzsch M, McKenna R

EMDB-26395: 
The SAAV capsid in complex with 6'SLN
Method: single particle / : Mietzsch M, McKenna R

PDB-7u94: 
SAAV pH 7.4 capsid structure
Method: single particle / : Mietzsch M, McKenna R

PDB-7u95: 
SAAV pH 6.0 capsid structure
Method: single particle / : Mietzsch M, McKenna R

PDB-7u96: 
SAAV pH 5.5 capsid structure
Method: single particle / : Mietzsch M, McKenna R

PDB-7u97: 
SAAV pH 4.0 capsid structure
Method: single particle / : Mietzsch M, McKenna R

EMDB-26467: 
SARS-CoV-2 6P Mut7 in complex with Fab THSC20.HVTR04 (1 RBD up and 1 RDB down)
Method: single particle / : Torres JL, Ward AB

EMDB-26470: 
SARS-CoV-2 6P Mut7 in complex with Fab THSC20.HVTR26 (1 RBD up, 1 RBD down)
Method: single particle / : Torres JL, Ward AB

EMDB-26472: 
SARS-CoV-2 6P Mut7 in complex with Fab THSC20.HVTR26 (1 RBD up)
Method: single particle / : Torres JL, Ward AB

EMDB-26473: 
SARS-CoV-2 6P Mut7 in complex with Fab THSC20.HVTR26 (3 RBD down)
Method: single particle / : Torres JL, Ward AB

EMDB-23986: 
Structure of the adeno-associated virus 9 capsid at pH 6.0
Method: single particle / : Penzes JJ, Chipman P, Bhattacharya N, Zeher A, Huang R, McKenna R, Agbandje-McKenna M

EMDB-23993: 
Structure of the adeno-associated virus 9 capsid at pH 5.5
Method: single particle / : Penzes JJ, Chipman P, Bhattacharya N, Zeher A, Huang R, McKenna R, Agbandje-McKenna M

EMDB-23999: 
Structure of the adeno-associated virus 9 capsid at pH 4.0
Method: single particle / : Penzes JJ, Chipman P, Bhattacharya N, Zeher A, Huang R, McKenna R, Agbandje-McKenna M

PDB-7mtg: 
Structure of the adeno-associated virus 9 capsid at pH 6.0
Method: single particle / : Penzes JJ, Chipman P, Bhattacharya N, Zeher A, Huang R, McKenna R, Agbandje-McKenna M

PDB-7mtp: 
Structure of the adeno-associated virus 9 capsid at pH 5.5
Method: single particle / : Penzes JJ, Chipman P, Bhattacharya N, Zeher A, Huang R, McKenna R, Agbandje-McKenna M

PDB-7mtw: 
Structure of the adeno-associated virus 9 capsid at pH 4.0
Method: single particle / : Penzes JJ, Chipman P, Bhattacharya N, Zeher A, Huang R, McKenna R, Agbandje-McKenna M

EMDB-24000: 
Structure of the adeno-associated virus 9 capsid at pH pH 7.4 in complex with terminal galactose
Method: single particle / : Penzes JJ, Chipman P, Bhattacharya N, Zeher A, Huang R, McKenna R, Agbandje-McKenna M

EMDB-24003: 
Structure of the adeno-associated virus 9 capsid at pH pH 5.5 in complex with terminal galactose
Method: single particle / : Penzes JJ, Chipman P, Bhattacharya N, Zeher A, Huang R, McKenna R, Agbandje-McKenna M

PDB-7mtz: 
Structure of the adeno-associated virus 9 capsid at pH pH 7.4 in complex with terminal galactose
Method: single particle / : Penzes JJ, Chipman P, Bhattacharya N, Zeher A, Huang R, McKenna R, Agbandje-McKenna M

PDB-7mua: 
Structure of the adeno-associated virus 9 capsid at pH pH 5.5 in complex with terminal galactose
Method: single particle / : Penzes JJ, Chipman P, Bhattacharya N, Zeher A, Huang R, McKenna R, Agbandje-McKenna M

EMDB-23973: 
Structure of the adeno-associated virus 9 capsid at pH 7.4
Method: single particle / : Penzes JJ, Chipman P, Bhattacharya N, Zeher A, Huang R, McKenna R, Agbandje-McKenna M

PDB-7mt0: 
Structure of the adeno-associated virus 9 capsid at pH 7.4
Method: single particle / : Penzes JJ, Chipman P, Bhattacharya N, Zeher A, Huang R, McKenna R, Agbandje-McKenna M
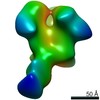
EMDB-22496: 
Base-directed response of rabbit #1 week 6 polyclonal fragment antigen binding in complex with 1PGE-THIVC SOSIP
Method: single particle / : Sewell LM, Ozorowski G, Ward AB

EMDB-22498: 
N611-site directed response of rabbit #1 week 6 polyclonal fragment antigen binding in complex with 1PGE-THIVC SOSIP
Method: single particle / : Sewell LM, Ozorowski G, Ward AB

EMDB-22499: 
Base-directed response of rabbit #1 week 12 polyclonal fragment antigen binding in complex with 1PGE-THIVC SOSIP
Method: single particle / : Sewell LM, Ozorowski G, Ward AB

EMDB-22500: 
N611-site directed response of rabbit #1 week 12 polyclonal fragment antigen binding in complex with 1PGE-THIVC SOSIP
Method: single particle / : Sewell LM, Ozorowski G, Ward AB

EMDB-22501: 
C3/V5-directed response of rabbit #1 week 12 polyclonal fragment antigen binding in complex with 1PGE-THIVC SOSIP
Method: single particle / : Sewell LM, Ozorowski G, Ward AB
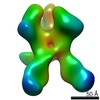
EMDB-22502: 
Base-directed response of rabbit #1 week 22 polyclonal fragment antigen binding in complex with 1PGE-THIVC SOSIP
Method: single particle / : Sewell LM, Ozorowski G, Ward AB

EMDB-22503: 
N611-site, base, and V1V3 directed response of rabbit #1 week 22 polyclonal fragment antigen binding in complex with 1PGE-THIVC SOSIP
Method: single particle / : Sewell LM, Ozorowski G, Ward AB
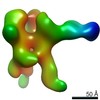
EMDB-22504: 
C3/V5 and base directed response of rabbit #1 week 22 polyclonal fragment antigen binding in complex with 1PGE-THIVC SOSIP
Method: single particle / : Sewell LM, Ozorowski G, Ward AB

EMDB-22505: 
V1/V3 and base directed response of rabbit #1 week 22 polyclonal fragment antigen binding in complex with 1PGE-THIVC SOSIP
Method: single particle / : Sewell LM, Ozorowski G, Ward AB

EMDB-23159: 
C3/V5-directed response of human donor G37080 polyclonal fragment antigen binding in complex with 1PGE-THIVC SOSIP
Method: single particle / : Sewell LM, Ozorowski G, Ward AB

EMDB-23460: 
Gorilla Bocavirus 1 Capsid
Method: single particle / : Yu JC, Mietzsch M

PDB-7lnk: 
Gorilla Bocavirus 1 Capsid
Method: single particle / : Yu JC, Mietzsch M, Agbandje-McKenna M

EMDB-22040: 
Asterix/Gtsf1 from mouse (full-length protein) bound to co-purifying tRNA
Method: single particle / : Ipsaro JJ, Joshua-Tor L
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
