-Search query
-Search result
Showing all 38 items for (author: bess & j)

EMDB-16512: 
MiniCoV-ADDomer, a SARS-CoV-2 epitope presenting viral like particle
Method: single particle / : Bufton JC, Capin J, Boruku U, Garzoni F, Schaffitzel C, Berger I

EMDB-16522: 
Structure of ADDoCoV-ADAH11
Method: single particle / : Yadav KNS, Buzas D, Berger-Schaffitzel C, Berger I

PDB-8c9n: 
MiniCoV-ADDomer, a SARS-CoV-2 epitope presenting viral like particle
Method: single particle / : Bufton JC, Capin J, Boruku U, Garzoni F, Schaffitzel C, Berger I

EMDB-29292: 
HIV-1 envelope trimer bound to one CD4 molecule on membranes
Method: subtomogram averaging / : Li W, Qin Z, Nand E, Grunst MW, Grover JR, Uchil PD, Mothes W

EMDB-29293: 
HIV-1 envelope trimer bound to two CD4 molecules on membranes
Method: subtomogram averaging / : Li W, Qin Z, Nand E, Grunst MW, Grover JR, Uchil PD, Mothes W

EMDB-29294: 
HIV-1 envelope trimer bound to three CD4 molecules on membranes
Method: subtomogram averaging / : Li W, Qin Z, Nand E, Grunst MW, Grover JR, Uchil PD, Mothes W

EMDB-29295: 
Consensus structure of HIV-1 envelope trimer bound to CD4 receptors on membranes
Method: subtomogram averaging / : Li W, Qin Z, Nand E, Grunst MW, Grover JR, Uchil PD, Mothes W

EMDB-41045: 
Unliganded HIV-1 Env on virion
Method: subtomogram averaging / : Li W, Qin Z, Nand E, Grunst MW, Grover JR, Uchil PD, Mothes W

EMDB-27735: 
Cryo-EM structure of SIVmac239 SOS-2P Env trimer in complex with human bNAb PGT145
Method: single particle / : Gorman J, Kwong PD

PDB-8dvd: 
Cryo-EM structure of SIVmac239 SOS-2P Env trimer in complex with human bNAb PGT145
Method: single particle / : Gorman J, Kwong PD

EMDB-27718: 
SIV E660.CR54 SOS-2P Env Trimer with ITS92.02
Method: single particle / : Gorman J, Kwong PD

PDB-8dua: 
SIV E660.CR54 SOS-2P Env Trimer with ITS92.02
Method: single particle / : Gorman J, Kwong PD

EMDB-25062: 
In-situ structure of SIV trimer
Method: subtomogram averaging / : Gorman J, Wang CY, Mason RD, Nazzari AF, Welles H, Zhou TQ, Jr JB, Tsybovsky Y, Verardi R, Yang YP, Zhang BS, Lifson JD, Liu J, Roederer M, Kwong PD

EMDB-25063: 
in situ SIVmac239-Env trimers on the surface of AT-2-inactivated virions
Method: subtomogram averaging / : Liu J, Wang CY

EMDB-25064: 
in situ SIVmac239-Env trimers on the surface of AT-2-inactivated virions
Method: subtomogram averaging / : Liu J, Wang CY

EMDB-25065: 
In-situ structure of SIVmac239-Env trimers with ITS90.03 associated
Method: subtomogram averaging / : Liu J, Wang CY

EMDB-14571: 
Negative stain 3D structure of the plastid-encoded RNA polymerase from Sinapis alba
Method: single particle / : Effantin G, Fenel D

EMDB-27631: 
SIV mac239 SOS-2P K169T Env trimer with bNAb PGT145 and jacalin
Method: single particle / : Gorman J, Kwong PD

EMDB-26608: 
Structure of the VP5*/VP8* assembly from the human rotavirus strain CDC-9 in complex with antibody 41 - Upright conformation
Method: single particle / : Jenni S, Zongli L, Wang Y, Bessey T, Salgado EN, Schmidt AG, Greenberg HB, Jiang B, Harrison SC

EMDB-26609: 
Structure of the VP5*/VP8* assembly from the human rotavirus strain CDC-9 - Reversed conformation
Method: single particle / : Jenni S, Zongli L, Wang Y, Bessey T, Salgado EN, Schmidt AG, Greenberg HB, Jiang B, Harrison SC

PDB-7ums: 
Structure of the VP5*/VP8* assembly from the human rotavirus strain CDC-9 in complex with antibody 41 - Upright conformation
Method: single particle / : Jenni S, Zongli L, Wang Y, Bessey T, Salgado EN, Schmidt AG, Greenberg HB, Jiang B, Harrison SC

PDB-7umt: 
Structure of the VP5*/VP8* assembly from the human rotavirus strain CDC-9 - Reversed conformation
Method: single particle / : Jenni S, Zongli L, Wang Y, Bessey T, Salgado EN, Schmidt AG, Greenberg HB, Jiang B, Harrison SC

EMDB-12158: 
CryoEM structure of Mycobacterium tuberculosis UMP Kinase (UMPK) in complex with UDP and UTP
Method: single particle / : Bous J, Trapani S, Walter P, Bron P, Munier-Lehmann H

PDB-7bes: 
CryoEM structure of Mycobacterium tuberculosis UMP Kinase (UMPK) in complex with UDP and UTP
Method: single particle / : Bous J, Trapani S, Walter P, Bron P, Munier-Lehmann H

EMDB-32033: 
14pf microtubule decorated with EML1-GFP
Method: helical / : Bangera M, Sirajuddin M

EMDB-21411: 
HIV envelope glycoprotein bound with soluble CD4 (D1-D2) and antibody 17b on AT-2 treated BaL strain virus
Method: subtomogram averaging / : Jun L, Ze L, Li W

EMDB-21412: 
Ligand-free HIV envelope glycoprotein on AT-2 treated BaL strain virus
Method: subtomogram averaging / : Jun L, Ze L, Li W

EMDB-21413: 
HIV envelope glycoprotein bound with antibodies 10-1074 and 3BNC117 on AT-2 treated BaL strain virus
Method: subtomogram averaging / : Jun L, Ze L, Li W
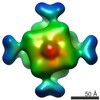
EMDB-2069: 
Octameric structure of IMPDH from Pseudomonas aeruginosa in its APO form
Method: single particle / : Labesse G, Vaupre L, Salard-Arnaud I, Alexandre T, Lai Kee Him J, Raynal B, Bron P, Munier-Lehmann H

EMDB-2070: 
Octameric structure of IMPDH from Pseudomonas aeruginosa in presence of Mg-ATP
Method: single particle / : Labesse G, Vaupre L, alard-Arnaud I, Alexandre T, Lai Kee Him J, Raynal B, Bron P, Munier-Lehmann H

EMDB-1846: 
Human native spliceosomal C complex assembled on PM5 pre-mRNA.
Method: single particle / : Golas MM, Sander B, Bessonov S, Grote M, Wolf E, Kastner B, Stark H, Luhrmann R

EMDB-1847: 
Human 35S U5 snRNP
Method: single particle / : Golas MM, Sander B, Bessonov S, Grote M, Wolf E, Kastner B, Stark H, Luhrmann R

EMDB-1848: 
High salt resistant human spliceosomal C complex core
Method: single particle / : Golas MM, Sander B, Bessonov S, Grote M, Wolf E, Kastner B, Stark H, Luhrmann R

EMDB-5272: 
Molecular Structure of Unliganded Native SIVmneE11S gp120 trimer: Spike region
Method: subtomogram averaging / : White TA, Bartesaghi A, Borgnia M, Subramaniam S
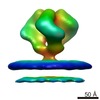
EMDB-5273: 
Molecular Structure of Unliganded Native SIVmac239 gp120 trimer: Spike region
Method: subtomogram averaging / : White TA, Bartesaghi A, Borgnia M, Subramaniam S

EMDB-5274: 
Molecular Structure of Unliganded Native CP-MAC gp120 trimer: Spike region
Method: subtomogram averaging / : White TA, Bartesaghi A, Borgnia M, Subramaniam S
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search





 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
