-Search query
-Search result
Showing 1 - 50 of 55 items for (author: bacia & k)

EMDB-17659: 
ACAD9-WT in complex with ECSIT-CTER
Method: single particle / : McGregor L, Acajjaoui S, Desfosses A, Saidi M, Bacia-Verloop M, Schwarz JJ, Juyoux P, Von Velsen J, Bowler MW, McCarthy A, Kandiah E, Gutsche I, Soler-Lopez M

EMDB-17660: 
Cryo-EM structure of human ACAD9-S191A
Method: single particle / : McGregor L, Acajjaoui S, Desfosses A, Saidi M, Bacia-Verloop M, Schwarz JJ, Juyoux P, Von Velsen J, Bowler MW, McCarthy A, Kandiah E, Gutsche I, Soler-Lopez M

EMDB-17661: 
ACAD9 homodimer WT
Method: single particle / : McGregor L, Acajjaoui S, Desfosses A, Saidi M, Bacia-Verloop M, Schwarz JJ, Juyoux P, Von Velsen J, Bowler MW, McCarthy A, Kandiah E, Gutsche I, Soler-Lopez M

PDB-8phe: 
ACAD9-WT in complex with ECSIT-CTER
Method: single particle / : McGregor L, Acajjaoui S, Desfosses A, Saidi M, Bacia-Verloop M, Schwarz JJ, Juyoux P, Von Velsen J, Bowler MW, McCarthy A, Kandiah E, Gutsche I, Soler-Lopez M

PDB-8phf: 
Cryo-EM structure of human ACAD9-S191A
Method: single particle / : McGregor L, Acajjaoui S, Desfosses A, Saidi M, Bacia-Verloop M, Schwarz JJ, Juyoux P, Von Velsen J, Bowler MW, McCarthy A, Kandiah E, Gutsche I, Soler-Lopez M

EMDB-13261: 
Providencia stuartii Arginine decarboxylase (Adc), decamer structure
Method: single particle / : Desfosses A, Jessop M, Bacia-Verloop M, Gutsche I

EMDB-13466: 
Providencia stuartii Arginine decarboxylase (Adc), stack structure
Method: single particle / : Jessop M, Desfosses A, Bacia-Verloop M, Gutsche I

PDB-7p9b: 
Providencia stuartii Arginine decarboxylase (Adc), decamer structure
Method: single particle / : Jessop M, Desfosses A, Bacia-Verloop M, Gutsche I

PDB-7pk6: 
Providencia stuartii Arginine decarboxylase (Adc), stack structure
Method: single particle / : Jessop M, Desfosses A, Bacia-Verloop M, Gutsche I

EMDB-10849: 
Inducible lysine decarboxylase LdcI decamer, pH 7.0
Method: single particle / : Jessop M, Felix J, Desfosses A, Effantin G, Gutsche I

EMDB-10850: 
Inducible lysine decarboxylase LdcI stacks, pH 5.7
Method: single particle / : Felix J, Jessop M, Desfosses A, Effantin G, Gutsche I

PDB-6yn5: 
Inducible lysine decarboxylase LdcI decamer, pH 7.0
Method: single particle / : Jessop M, Felix J, Desfosses A, Effantin G, Gutsche I

PDB-6yn6: 
Inducible lysine decarboxylase LdcI stacks, pH 5.7
Method: single particle / : Felix J, Jessop M, Desfosses A, Effantin G, Gutsche I

EMDB-12055: 
ACAD9-ECSIT-CTD (ACAD9 core)
Method: single particle / : Giachin G, Jessop M, Soler-Lopez M, Gutsche I

EMDB-10926: 
Structure of jumbo coliphage phAPEC6 capsid
Method: single particle / : Wagemans J, Tsonos J, Holtappels D, Fortuna K, Hernalsteens JP, De Greve H, Estrozi LF, Bacia-Verloop M, Moriscot C, Noben JP, Schoehn G, Lavigne R

EMDB-10929: 
3D structure of bacteriophage phAPEC6 tail
Method: single particle / : Wagemans J, Tsonos J, Holtappels D, Fortuna K, Hernalsteens JP, De Greve H, Estrozi LF, Bacia-Verloop M, Moriscot C, Noben JP, Schoehn G, Lavigne R

EMDB-10351: 
MoxR AAA-ATPase RavA, C2-symmetric closed ring conformation
Method: single particle / : Jessop M, Felix J, Gutsche I

EMDB-10352: 
MoxR AAA-ATPase RavA, spiral open ring conformation
Method: single particle / : Jessop M, Felix J, Gutsche I

PDB-6sza: 
MoxR AAA-ATPase RavA, C2-symmetric closed ring conformation
Method: single particle / : Jessop M, Felix J, Gutsche I

PDB-6szb: 
MoxR AAA-ATPase RavA, spiral open ring conformation
Method: single particle / : Jessop M, Felix J, Gutsche I

EMDB-4469: 
Spiral structure of E. coli RavA in the RavA-LdcI cage-like complex
Method: single particle / : Arragain B, Felix J, Malet H, Gutsche I, Jessop M

EMDB-4470: 
Spiral structure of E. coli RavA in the RavA-LdcI cage-like complex
Method: single particle / : Arragain B, Felix J, Malet H, Gutsche I, Jessop M

PDB-6q7l: 
Spiral structure of E. coli RavA in the RavA-LdcI cage-like complex
Method: single particle / : Arragain B, Felix J, Malet H, Gutsche I, Jessop M

PDB-6q7m: 
Spiral structure of E. coli RavA in the RavA-LdcI cage-like complex
Method: single particle / : Arragain B, Felix J, Malet H, Gutsche I, Jessop M

EMDB-4468: 
Lysine decarboxylase A from Pseudomonas aeruginosa
Method: single particle / : Kandiah E, Gutsche I

PDB-6q6i: 
Lysine decarboxylase A from Pseudomonas aeruginosa
Method: single particle / : Kandiah E, Gutsche I

EMDB-10160: 
In Situ Core-Signalling Unit of E. coli Chemoreceptor Array
Method: subtomogram averaging / : Burt A, Desfosses A, Gutsche I

EMDB-3204: 
Structures of E.coli lysine decarboxylases
Method: single particle / : Kandiah E, Carriel D, Perard J, Malet H, Bacia M, Liu K, Chan WSS, Houry AW, Ollagnier de Choudens S, Elsen S, Gutsche I

EMDB-3205: 
Structure of E.coli Constitutive lysine decarboxylase
Method: single particle / : Kandiah E, Carriel D, Perard J, Malet H, Bacia M, Liu K, Chan WSS, Houry AW, Ollagnier de Choudens S, Elsen S, Gutsche I

EMDB-3206: 
Revisited cryo-EM structure of Inducible lysine decarboxylase complexed with LARA domain of RavA ATPase
Method: single particle / : Kandiah E, Carriel D, Perard J, Malet H, Bacia M, Liu K, Chan SWS, Houry WA, Ollagnier de Choudens S, Elsen S, Gutsche I

PDB-5fkx: 
Structure of E.coli inducible lysine decarboxylase at active pH
Method: single particle / : Kandiah E, Carriel D, Perard J, Malet H, Bacia M, Liu K, Chan SWS, Houry WA, Ollagnier de Choudens S, Elsen S, Gutsche I

PDB-5fkz: 
Structure of E.coli Constitutive lysine decarboxylase
Method: single particle / : Kandiah E, Carriel D, Perard J, Malet H, Bacia M, Liu K, Chan SWS, Houry WA, Ollagnier de Choudens S, Elsen S, Gutsche I

PDB-5fl2: 
Revisited cryo-EM structure of Inducible lysine decarboxylase complexed with LARA domain of RavA ATPase
Method: single particle / : Kandiah E, Carriel D, Perard J, Malet H, Bacia M, Liu K, Chan SWS, Houry WA, Ollagnier de Choudens S, Elsen S, Gutsche I

EMDB-2679: 
Electron cryo-microscopy of the complex formed between the hexameric AAA+ ATPase RavA and the decameric inducible decarboxylase LdcI
Method: single particle / : Malet H, Liu K, El Bakkouri M, Chan SWS, Effantin G, Bacia M, Houry WA, Gutsche I

EMDB-2681: 
Assembly principles of the unique cage formed by the AAA+ ATPase RavA hexamer and the lysine decarboxylase LdcI decamer
Method: single particle / : Malet H, Liu K, El Bakkouri M, Chan SWS, Effantin G, Bacia M, Houry WA, Gutsche I

PDB-4upb: 
Electron cryo-microscopy of the complex formed between the hexameric ATPase RavA and the decameric inducible decarboxylase LdcI
Method: single particle / : Malet H, Liu K, El Bakkouri M, Chan SWS, Effantin G, Bacia M, Houry WA, Gutsche I

PDB-4upf: 
Assembly principles of the unique cage formed by the ATPase RavA hexamer and the lysine decarboxylase LdcI decamer
Method: single particle / : Malet H, Liu K, El Bakkouri M, Chan SWS, Effantin G, Bacia M, Houry WA, Gutsche I

EMDB-2428: 
The structure of the COPII coat assembled on membranes
Method: subtomogram averaging / : Zanetti G, Prinz S, Daum S, Meister A, Schekman R, Bacia K, Briggs JAG

EMDB-2429: 
The structure of the COPII coat assembled on membranes
Method: subtomogram averaging / : Zanetti G, Prinz S, Daum S, Meister A, Schekman R, Bacia K, Briggs JAG

EMDB-2430: 
The structure of the COPII coat assembled on membranes
Method: subtomogram averaging / : Zanetti G, Prinz S, Daum S, Meister A, Schekman R, Bacia K, Briggs JAG
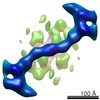
EMDB-2431: 
The structure of the COPII coat assembled on membranes
Method: subtomogram averaging / : Zanetti G, Prinz S, Daum S, Meister A, Schekman R, Bacia K, Briggs JAG

EMDB-2432: 
The structure of the COPII coat assembled on membranes
Method: electron tomography / : Zanetti G, Prinz S, Daum S, Meister A, Schekman R, Bacia K, Briggs JAG

PDB-4bzi: 
The structure of the COPII coat assembled on membranes
Method: electron tomography / : Zanetti G, Prinz S, Daum S, Meister A, Schekman R, Bacia K, Briggs JAG

PDB-4bzj: 
The structure of the COPII coat assembled on membranes
Method: subtomogram averaging / : Zanetti G, Prinz S, Daum S, Meister A, Schekman R, Bacia K, Briggs JAG

PDB-4bzk: 
The structure of the COPII coat assembled on membranes
Method: subtomogram averaging / : Zanetti G, Prinz S, Daum S, Meister A, Schekman R, Bacia K, Briggs JAG

EMDB-2243: 
Cryo-electron microscopy of phirsl1 jumbo phage
Method: single particle / : Effantin G, Hamasaki R, Kawasaki T, Bacia M, Moriscot C, Weissenhorn W, Yamada T, Schoehn G

EMDB-2244: 
Cryo-electron microscopy of phirsl1 jumbo phage
Method: helical / : Effantin G, Hamasaki R, Kawasaki T, Bacia M, Moriscot C, Weissenhorn W, Yamada T, Schoehn G

EMDB-2245: 
Cryo-electron microscopy of phirsl1 jumbo phage
Method: single particle / : Effantin G, Hamasaki R, Kawasaki T, Bacia M, Moriscot C, Weissenhorn W, Yamada T, Schoehn G

EMDB-2246: 
Cryo-electron microscopy of phirsl1 jumbo phage
Method: single particle / : Effantin G, Hamasaki R, Kawasaki T, Bacia M, Moriscot C, Weissenhorn W, Yamada T, Schoehn G

EMDB-2247: 
Cryo-electron microscopy of phirsl1 jumbo phage
Method: single particle / : Effantin G, Hamasaki R, Kawasaki T, Bacia M, Moriscot C, Weissenhorn W, Yamada T, Schoehn G
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
