-Search query
-Search result
Showing 1 - 50 of 95 items for (author: adams & p)

EMDB-41963: 
Preholo-Proteasome from Beta 3 D205 deletion
Method: single particle / : Walsh Jr RM, Rawson S, Velez B, Blickling M, Razi A, Hanna J

EMDB-41993: 
Proteasome 20S Core Particle from Beta 3 D205 deletion
Method: single particle / : Walsh Jr RM, Rawson S, Velez B, Blickling M, Razi A, Hanna J
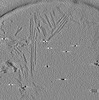
EMDB-28752: 
Cryo-electron tomography of wild type a-synuclein preformed fibrils
Method: electron tomography / : Jiang J, Boparai N, Dai W, Kim YS
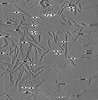
EMDB-28753: 
Cryo-electron tomography of S42Y a-synuclein preformed fibrils
Method: electron tomography / : Jiang J, Boparai N, Dai W, Kim YS

EMDB-40711: 
CryoEM structure of Western equine encephalitis virus VLP in complex with the chimeric Du-D1-Mo-D2 MXRA8 receptor
Method: single particle / : Zimmerman MI, Fremont DH, Center for Structural Genomics of Infectious Diseases (CSGID)

PDB-8sqn: 
CryoEM structure of Western equine encephalitis virus VLP in complex with the chimeric Du-D1-Mo-D2 MXRA8 receptor
Method: single particle / : Zimmerman MI, Fremont DH, Center for Structural Genomics of Infectious Diseases (CSGID), Center for Structural Biology of Infectious Diseases (CSBID)

EMDB-18400: 
GDNF/GFRa1 cell adhesion complex cryo-ET structure
Method: subtomogram averaging / : Houghton FM, Briggs DC, McDonald NQ

EMDB-18651: 
GDNF/GFRa1 cell adhesion complex bridging between adhering liposomes.
Method: electron tomography / : Houghton FM, Briggs DC, McDonald NQ

EMDB-26855: 
Arabidopsis DDM1 bound to nucleosome (H2A.W, H2B, H3.3, H4, with 147 bp DNA)
Method: single particle / : Ipsaro JJ, Adams DW, Joshua-Tor L

PDB-7ux9: 
Arabidopsis DDM1 bound to nucleosome (H2A.W, H2B, H3.3, H4, with 147 bp DNA)
Method: single particle / : Ipsaro JJ, Adams DW, Joshua-Tor L

EMDB-33718: 
ZIKV_Fab_G9E
Method: single particle / : Shu B, Thiam-Seng N, Lok S

PDB-7yar: 
ZIKV_Fab_G9E
Method: single particle / : Shu B, Thiam-Seng N, Lok S

EMDB-25487: 
SARS-CoV-2 Spike NTD in complex with neutralizing Fab SARS2-57 (local refinement)
Method: single particle / : Adams LJ, Fremont DH

EMDB-25488: 
SARS-CoV-2 Spike in complex with neutralizing Fab SARS2-57 (three down conformation)
Method: single particle / : Adams LJ, Fremont DH

EMDB-27438: 
Cryo-EM structure of SARS-CoV-2 Beta (B.1.351) spike protein in complex with VH domain F6
Method: single particle / : Zhu X, Saville JW, Mannar D, Berezuk AM, Subramaniam S

EMDB-27439: 
Cryo-EM structure of SARS-CoV-2 Beta (B.1.351) spike protein in complex with VH domain F6 (focused refinement of RBD and VH F6)
Method: single particle / : Zhu X, Saville JW, Mannar D, Berezuk AM, Subramaniam S

PDB-8di5: 
Cryo-EM structure of SARS-CoV-2 Beta (B.1.351) spike protein in complex with VH domain F6 (focused refinement of RBD and VH F6)
Method: single particle / : Zhu X, Saville JW, Mannar D, Berezuk AM, Subramaniam S

EMDB-26191: 
Cryo-EM structure of human SARM1 TIR domain in complex with 1AD
Method: single particle / : Saikot FK, Kobe B, Ve T, Nanson JD, Gu W, Luo Z, Brillault L, Landsberg MJ

EMDB-24272: 
Cryo-EM structure of activated human SARM1 in complex with NMN and 1AD (TIR:1AD)
Method: single particle / : Kerry PS, Nanson JD, Adams S, Cunnea K, Bosanac T, Kobe B, Hughes RO, Ve T

EMDB-24273: 
Cryo-EM structure of activated human SARM1 in complex with NMN and 1AD (ARM and SAM domains)
Method: single particle / : Kerry PS, Nanson JD, Adams S, Cunnea K, Bosanac T, Kobe B, Hughes RO, Ve T

EMDB-24274: 
Cryo-EM structure of activated human SARM1 in complex with NMN and 1AD (SAM-TIR:1AD)
Method: single particle / : Kerry PS, Adams S, Cunnea K, Brearley A, Bosanac T, Hughes RO, Ve T

PDB-7nak: 
Cryo-EM structure of activated human SARM1 in complex with NMN and 1AD (TIR:1AD)
Method: single particle / : Kerry PS, Nanson JD, Adams S, Cunnea K, Bosanac T, Kobe B, Hughes RO, Ve T

PDB-7nal: 
Cryo-EM structure of activated human SARM1 in complex with NMN and 1AD (ARM and SAM domains)
Method: single particle / : Kerry PS, Nanson JD, Adams S, Cunnea K, Bosanac T, Kobe B, Hughes RO, Ve T

EMDB-23812: 
Structure of the apo phosphoinositide 3-kinase p110 gamma (PIK3CG) p101 (PIK3R5) complex
Method: single particle / : Burke JE, Dalwadi U, Rathinaswamy MK, Yip CK

EMDB-24674: 
Seipin forms a flexible cage at lipid droplet formation sites
Method: single particle / : Arlt H, Sui X, Folger B, Adams C, Chen X, Remme R, Hamprecht FA, DiMaio F, Liao M, Goodman JM, Farese Jr RV, Walther TC

PDB-7rsl: 
Seipin forms a flexible cage at lipid droplet formation sites
Method: single particle / : Arlt H, Sui X, Folger B, Adams C, Chen X, Remme R, Hamprecht FA, DiMaio F, Liao M, Goodman JM, Farese Jr RV, Walther TC

EMDB-23379: 
Thermotoga maritima Encapsulin Nanocompartment Pore Mutant S7D
Method: single particle / : Andreas MP, Adamson L, Tasneem N, Close W, Giessen T, Lau YH

EMDB-23380: 
Thermotoga maritima Encapsulin Nanocompartment Pore Mutant S1K
Method: single particle / : Andreas MP, Adamson L, Tasneem N, Close W, Giessen T, Lau YH

EMDB-23381: 
Thermotoga maritima Encapsulin Nanocompartment Pore Mutant S1R
Method: single particle / : Andreas MP, Adamson L, Tasneem N, Close W, Giessen T, Lau YH

EMDB-23382: 
Thermotoga maritima Encapsulin Nanocompartment Pore Mutant SE3
Method: single particle / : Andreas MP, Adamson L, Tasneem N, Close W, Giessen T, Lau YH

EMDB-23383: 
Thermotoga maritima Encapsulin Nanocompartment Pore Mutant S6E
Method: single particle / : Andreas MP, Adamson L, Tasneem N, Close W, Giessen T, Lau YH

EMDB-23384: 
Thermotoga maritima Encapsulin Nanocompartment Pore Mutant S5D
Method: single particle / : Andreas MP, Adamson L, Tasneem N, Close W, Giessen T, Lau YH

EMDB-23385: 
Thermotoga maritima Encapsulin Nanocompartment Pore Mutant S7G
Method: single particle / : Andreas MP, Adamson L, Tasneem N, Close W, Giessen T, Lau YH

PDB-7lii: 
Thermotoga maritima Encapsulin Nanocompartment Pore Mutant S7D
Method: single particle / : Andreas MP, Adamson L, Tasneem N, Close W, Giessen T, Lau YH

PDB-7lij: 
Thermotoga maritima Encapsulin Nanocompartment Pore Mutant S1K
Method: single particle / : Andreas MP, Adamson L, Tasneem N, Close W, Giessen T, Lau YH

PDB-7lik: 
Thermotoga maritima Encapsulin Nanocompartment Pore Mutant S1R
Method: single particle / : Andreas MP, Adamson L, Tasneem N, Close W, Giessen T, Lau YH

PDB-7lil: 
Thermotoga maritima Encapsulin Nanocompartment Pore Mutant SE3
Method: single particle / : Andreas MP, Adamson L, Tasneem N, Close W, Giessen T, Lau YH

PDB-7lim: 
Thermotoga maritima Encapsulin Nanocompartment Pore Mutant S6E
Method: single particle / : Andreas MP, Adamson L, Tasneem N, Close W, Giessen T, Lau YH

PDB-7lis: 
Thermotoga maritima Encapsulin Nanocompartment Pore Mutant S5D
Method: single particle / : Andreas MP, Adamson L, Tasneem N, Close W, Giessen T, Lau YH

PDB-7lit: 
Thermotoga maritima Encapsulin Nanocompartment Pore Mutant S7G
Method: single particle / : Andreas MP, Adamson L, Tasneem N, Close W, Giessen T, Lau YH

EMDB-23808: 
Structure of the phosphoinositide 3-kinase p110 gamma (PIK3CG) p101 (PIK3R5) complex
Method: single particle / : Burke JE, Dalwadi U, Rathinaswamy MK, Yip CK

EMDB-12604: 
Cryo-EM structure of the mycolic acid transporter MmpL3 from M. tuberculosis
Method: single particle / : Adams O, Deme JC, Parker JL, Lea SM, Newstead S

PDB-7nvh: 
Cryo-EM structure of the mycolic acid transporter MmpL3 from M. tuberculosis
Method: single particle / : Adams O, Deme JC, Parker JL, Lea SM, Newstead S

EMDB-23898: 
SARS-CoV-2 Spike in complex with neutralizing Fab SARS2-38 (three down conformation)
Method: single particle / : Adams LJ, Fremont DH, Center for Structural Genomics of Infectious Diseases (CSGID)

EMDB-23899: 
SARS-CoV-2 Spike RBD in complex with neutralizing Fab SARS2-38 (local refinement)
Method: single particle / : Adams LJ, Fremont DH, Center for Structural Genomics of Infectious Diseases (CSGID)

PDB-7mkl: 
SARS-CoV-2 Spike in complex with neutralizing Fab SARS2-38 (three down conformation)
Method: single particle / : Adams LJ, Fremont DH, Center for Structural Genomics of Infectious Diseases (CSGID)

PDB-7mkm: 
SARS-CoV-2 Spike RBD in complex with neutralizing Fab SARS2-38 (local refinement)
Method: single particle / : Adams LJ, Fremont DH, Center for Structural Genomics of Infectious Diseases (CSGID)

EMDB-22410: 
Cryo-electron tomography of HIV-1 integrase mutant, W235E
Method: electron tomography / : Adams LJ, Steven AC

EMDB-22411: 
Cryo-electron tomography of HIV-1 integrase mutant, W235E
Method: electron tomography / : Adams LJ, Steven AC

EMDB-23211: 
Cryo-EM structure of human ACE2 receptor bound to protein encoded by vaccine candidate BNT162b1
Method: single particle / : Lees JA, Han S
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
