+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-9565 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | EM Structure of VP1A and VP1B | |||||||||
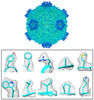
 Map data Map data | the structure of a cypovirus capsid of 750-%u212B diameter at 3.3-%u212B resolution using a 200 kV TEM | |||||||||
 Sample Sample |
| |||||||||
| Function / homology | : / : / CPV Capsid shell protein VP1, small protrusion domain / Inner layer core protein VP1-like, C-terminal / T=2 icosahedral viral capsid / viral inner capsid / VP1 / Capsid protein VP1 Function and homology information Function and homology information | |||||||||
| Biological species |   Bombyx mori cypovirus 1 Bombyx mori cypovirus 1 | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.3 Å | |||||||||
 Authors Authors | Li X / Zhou N / Xu B / Chen W / Zhu B / Wang X / Wang J / Liu H / Cheng L | |||||||||
 Citation Citation |  Journal: J Mol Biol / Year: 2017 Journal: J Mol Biol / Year: 2017Title: Near-Atomic Resolution Structure Determination of a Cypovirus Capsid and Polymerase Complex Using Cryo-EM at 200kV. Authors: Xiaowu Li / Niyun Zhou / Wenyuan Chen / Bin Zhu / Xurong Wang / Bin Xu / Jiawei Wang / Hongrong Liu / Lingpeng Cheng /  Abstract: Single-particle cryo-electron microscopy (cryo-EM) allows the high-resolution structural determination of biological assemblies in a near-native environment. However, all high-resolution (better than ...Single-particle cryo-electron microscopy (cryo-EM) allows the high-resolution structural determination of biological assemblies in a near-native environment. However, all high-resolution (better than 3.5Å) cryo-EM structures reported to date were obtained by using 300kV transmission electron microscopes (TEMs). We report here the structures of a cypovirus capsid of 750-Å diameter at 3.3-Å resolution and of RNA-dependent RNA polymerase (RdRp) complexes within the capsid at 3.9-Å resolution using a 200-kV TEM. The newly resolved structure revealed conformational changes of two subdomains in the RdRp. These conformational changes, which were involved in RdRp's switch from non-transcribing to transcribing mode, suggest that the RdRp may facilitate the unwinding of genomic double-stranded RNA. The possibility of 3-Å resolution structural determinations for biological assemblies of relatively small sizes using cryo-EM at 200kV was discussed. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_9565.map.gz emd_9565.map.gz | 1.2 GB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-9565-v30.xml emd-9565-v30.xml emd-9565.xml emd-9565.xml | 10.5 KB 10.5 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_9565.png emd_9565.png | 170.2 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-9565 http://ftp.pdbj.org/pub/emdb/structures/EMD-9565 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9565 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9565 | HTTPS FTP |
-Validation report
| Summary document |  emd_9565_validation.pdf.gz emd_9565_validation.pdf.gz | 743 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_9565_full_validation.pdf.gz emd_9565_full_validation.pdf.gz | 742.6 KB | Display | |
| Data in XML |  emd_9565_validation.xml.gz emd_9565_validation.xml.gz | 9.6 KB | Display | |
| Data in CIF |  emd_9565_validation.cif.gz emd_9565_validation.cif.gz | 11.1 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-9565 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-9565 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-9565 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-9565 | HTTPS FTP |
-Related structure data
| Related structure data |  5h0sMC  9564C  5h0rC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_9565.map.gz / Format: CCP4 / Size: 1.3 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_9565.map.gz / Format: CCP4 / Size: 1.3 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | the structure of a cypovirus capsid of 750-%u212B diameter at 3.3-%u212B resolution using a 200 kV TEM | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.932 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Cypovirus
| Entire | Name:  Cypovirus (cytoplasmic polyhedrosis viruses) Cypovirus (cytoplasmic polyhedrosis viruses) |
|---|---|
| Components |
|
-Supramolecule #1: Cypovirus
| Supramolecule | Name: Cypovirus / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|
-Macromolecule #1: VP1
| Macromolecule | Name: VP1 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Bombyx mori cypovirus 1 Bombyx mori cypovirus 1 |
| Molecular weight | Theoretical: 148.696062 KDa |
| Recombinant expression | Organism:  Cypovirus (cytoplasmic polyhedrosis viruses) Cypovirus (cytoplasmic polyhedrosis viruses) |
| Sequence | String: MHSTNNNSNK RNNEEKHKQP EIDSSANNGE GTSGTRAQTV GDTATEAGVR NETKAGASTR RQTDGTGLSG TNAKIATASS ARQTDVEKP ADVTFTIENV DDVGIMQQKK PPTVVQSRTD VFNEQFANEA LHPMTKVIFN GLDVNTEVQP LSDDFKQISD P KGYLTYSV ...String: MHSTNNNSNK RNNEEKHKQP EIDSSANNGE GTSGTRAQTV GDTATEAGVR NETKAGASTR RQTDGTGLSG TNAKIATASS ARQTDVEKP ADVTFTIENV DDVGIMQQKK PPTVVQSRTD VFNEQFANEA LHPMTKVIFN GLDVNTEVQP LSDDFKQISD P KGYLTYSV KYEDQFTKKD KLRASEADDR IVGPTVNLFK YGAAVVNIDL NRDFFDTATG IDLTKGIPLV QDLLVPIGVT AG AEQSAEY VSGLLMVLFK VMTDNRLVIV GETTTPMSNT LSTVVNNVLR TTYHNNVGVN PALLRDFTQV NWLNRDITNM LQQ AGTKYG LGLTETRLDY VRLVKTIVGH ALNIDHFAAS VLNINLRALM EANVTADDRI KALQAHSMIS TQFHGPNQGA LRPE LAFDH DHIIRCLMLA AANYPRLEGI IVQINTGYVA SANVIRPVSE KRYFPENLEQ NQSAARLVSA VKARASEADI SSIHL AIAR EVSPMFNVHE LKKIAESFED PSSIVVVLEF ILFALFFPTE FNRIKGDIQN VLLLFFSRWY PVEYGIFIQR GATYTI NAA GEFEFSGRNE KWDQSLYLSE HFPALFSDVP LAGANTIIAI MRLFTPQGFL RTDDLAIAAN FPRASRNPQT YIPYTNQ RG TVTNEFASRF RTIVATLANV VNERAVQDDM QKATRSCTKQ WLRHLETQFD NIAVAHTDHL SVVYATMSNF MLNFTNNF S GNHATFKPDQ YVITSPEGSY KPIIERQGET VDGLTIIDTS IVWPILCQCT YPLVRQSGKG VDAVSIMEEI VYPDPSTTL SQSLSVAQVL SKLTLPDAFI NMILSGGDSV VMRTYQTEAD DDLDEGIRMT TYDQYLSHIR ERLHITNVPD PIYITGASTP DQIAASVQA THVAVVLYQS GVINGSASTY LRENEVLVVM PDYYDVVSRF ANANLQMNNN RYHESVLEIA DIFDQADFIQ T SDAVRQLR ALMPTLSTSQ IRHAIERIAQ ITDVDSTDYG KLTLRFLGTL TRSLKMQNAQ IRRIRPDGTV LRYDDQIDIE AF RWSRYFL DELRLRRLSV GLRLITNPRI ARRFDGVRIM YLTDDDPDPD FVPDVPEGYV AVQYAHRLFS SSLANKRNRV TYT HPPTGM AYPSPTGRPH VHMTINERAG MSKLVADNII ASVIKSNWVV DIHDIEYTAE VMTPSEGYTQ HVDAESIMTA PKGK LFHLQ FMDGLLRPEP SAFDPPASGE DMRLIYPLQP ISVARSMRAI VNHNEVDRPR GAVAPSSYEM DTGTLSRNGD LLYSP VANG QVGIPKLEVD HISFSNVVSM MTANIRTGDD MAVERVNPDD VRAINIRNA |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: NITROGEN |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI ARCTICA |
|---|---|
| Image recording | Film or detector model: FEI FALCON II (4k x 4k) / Average electron dose: 20.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.3 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 27000 |
|---|---|
| Initial angle assignment | Type: COMMON LINE |
| Final angle assignment | Type: COMMON LINE |
 Movie
Movie Controller
Controller