[English] 日本語
 Yorodumi
Yorodumi- EMDB-8309: HIV-1 Env BG505 SOSIP.664 trimer in complex with 2 copies of rabb... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-8309 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | HIV-1 Env BG505 SOSIP.664 trimer in complex with 2 copies of rabbit antibody 12A fragment antigen binding | |||||||||
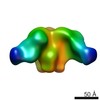
 Map data Map data | HIV-1 Env BG505 SOSIP.664 in complex with rabbit antibody 12A fragment antigen binding | |||||||||
 Sample Sample |
| |||||||||
| Biological species |    Human immunodeficiency virus 1 Human immunodeficiency virus 1 | |||||||||
| Method | single particle reconstruction / negative staining / Resolution: 22.0 Å | |||||||||
 Authors Authors | Ozorowski G / Ward AB | |||||||||
 Citation Citation |  Journal: Cell Rep / Year: 2016 Journal: Cell Rep / Year: 2016Title: Holes in the Glycan Shield of the Native HIV Envelope Are a Target of Trimer-Elicited Neutralizing Antibodies. Authors: Laura E McCoy / Marit J van Gils / Gabriel Ozorowski / Terrence Messmer / Bryan Briney / James E Voss / Daniel W Kulp / Matthew S Macauley / Devin Sok / Matthias Pauthner / Sergey Menis / ...Authors: Laura E McCoy / Marit J van Gils / Gabriel Ozorowski / Terrence Messmer / Bryan Briney / James E Voss / Daniel W Kulp / Matthew S Macauley / Devin Sok / Matthias Pauthner / Sergey Menis / Christopher A Cottrell / Jonathan L Torres / Jessica Hsueh / William R Schief / Ian A Wilson / Andrew B Ward / Rogier W Sanders / Dennis R Burton /    Abstract: A major advance in the search for an HIV vaccine has been the development of a near-native Envelope trimer (BG505 SOSIP.664) that can induce robust autologous Tier 2 neutralization. Here, potently ...A major advance in the search for an HIV vaccine has been the development of a near-native Envelope trimer (BG505 SOSIP.664) that can induce robust autologous Tier 2 neutralization. Here, potently neutralizing monoclonal antibodies (nAbs) from rabbits immunized with BG505 SOSIP.664 are shown to recognize an immunodominant region of gp120 centered on residue 241. Residue 241 occupies a hole in the glycan defenses of the BG505 isolate, with fewer than 3% of global isolates lacking a glycan site at this position. However, at least one conserved glycan site is missing in 89% of viruses, suggesting the presence of glycan holes in most HIV isolates. Serum evidence is consistent with targeting of holes in natural infection. The immunogenic nature of breaches in the glycan shield has been under-appreciated in previous attempts to understand autologous neutralizing antibody responses and has important potential consequences for HIV vaccine design. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_8309.map.gz emd_8309.map.gz | 11.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-8309-v30.xml emd-8309-v30.xml emd-8309.xml emd-8309.xml | 12.7 KB 12.7 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_8309.png emd_8309.png | 79.5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-8309 http://ftp.pdbj.org/pub/emdb/structures/EMD-8309 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8309 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8309 | HTTPS FTP |
-Validation report
| Summary document |  emd_8309_validation.pdf.gz emd_8309_validation.pdf.gz | 78.2 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_8309_full_validation.pdf.gz emd_8309_full_validation.pdf.gz | 77.3 KB | Display | |
| Data in XML |  emd_8309_validation.xml.gz emd_8309_validation.xml.gz | 494 B | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-8309 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-8309 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-8309 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-8309 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_8309.map.gz / Format: CCP4 / Size: 15.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_8309.map.gz / Format: CCP4 / Size: 15.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | HIV-1 Env BG505 SOSIP.664 in complex with rabbit antibody 12A fragment antigen binding | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.57 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : HIV-1 Env BG505 SOSIP.664 in complex with rabbit antibody 12A fra...
| Entire | Name: HIV-1 Env BG505 SOSIP.664 in complex with rabbit antibody 12A fragment antigen binding |
|---|---|
| Components |
|
-Supramolecule #1: HIV-1 Env BG505 SOSIP.664 in complex with rabbit antibody 12A fra...
| Supramolecule | Name: HIV-1 Env BG505 SOSIP.664 in complex with rabbit antibody 12A fragment antigen binding type: complex / ID: 1 / Parent: 0 |
|---|---|
| Molecular weight | Theoretical: 570 KDa |
-Supramolecule #2: Rabbit antibody 12A fragment antigen binding
| Supramolecule | Name: Rabbit antibody 12A fragment antigen binding / type: complex / ID: 2 / Parent: 1 Details: Fab fragment generated by proteolytic cleavage of IgG antibody |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  Homo sapiens (human) / Recombinant strain: HEK293F / Recombinant plasmid: pCMV3 Homo sapiens (human) / Recombinant strain: HEK293F / Recombinant plasmid: pCMV3 |
| Molecular weight | Theoretical: 50 KDa |
-Supramolecule #3: HIV-1 Env BG505 SOSIP.664
| Supramolecule | Name: HIV-1 Env BG505 SOSIP.664 / type: complex / ID: 3 / Parent: 1 |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
| Recombinant expression | Organism:  Homo sapiens (human) / Recombinant strain: HEK293F / Recombinant plasmid: pPPI4 Homo sapiens (human) / Recombinant strain: HEK293F / Recombinant plasmid: pPPI4 |
| Molecular weight | Theoretical: 420 KDa |
-Experimental details
-Structure determination
| Method | negative staining |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.03 mg/mL |
|---|---|
| Buffer | pH: 7.4 / Details: 50 mM Tris-HCl, pH 7.4, 150 mM NaCl |
| Staining | Type: NEGATIVE / Material: uranyl formate Details: 2% uranyl formate used to stain sample for 60 seconds. |
| Grid | Model: EMS Cu400 / Material: COPPER / Support film - Material: PARLODION / Support film - topology: CONTINUOUS / Pretreatment - Type: PLASMA CLEANING |
| Details | Complex formed by overnight incubation of Fab with Env at a 6x molar excess of Fab to Env and dilution to 0.03 mg/mL prior to grid adsorption. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: FEI CETA (4k x 4k) / Average electron dose: 25.0 e/Å2 Details: Stage tilts of -50, -40, -30, -20, -10, and 0 degrees were performed to increase orientation sampling. |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.5 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 92000 |
| Sample stage | Specimen holder model: SIDE ENTRY, EUCENTRIC |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: NONE Details: Initial model created using representative 2D class averages |
|---|---|
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Resolution.type: BY AUTHOR / Resolution: 22.0 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: SPARX / Number images used: 6798 |
| Initial angle assignment | Type: PROJECTION MATCHING / Software - Name: EMAN2 / Software - details: e2initialmodel |
| Final angle assignment | Type: PROJECTION MATCHING |
 Movie
Movie Controller
Controller