[English] 日本語
 Yorodumi
Yorodumi- EMDB-4057: Structure of the yeast spliceosome immediately after branching. 3... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-4057 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of the yeast spliceosome immediately after branching. 3D class containing helicase module. | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationRNA exon ligation / snRNA metabolic process / snRNA modification / post-spliceosomal complex / U2-type post-mRNA release spliceosomal complex / cellular bud site selection / post-mRNA release spliceosomal complex / 3'-5' RNA helicase activity / generation of catalytic spliceosome for first transesterification step / cis assembly of pre-catalytic spliceosome ...RNA exon ligation / snRNA metabolic process / snRNA modification / post-spliceosomal complex / U2-type post-mRNA release spliceosomal complex / cellular bud site selection / post-mRNA release spliceosomal complex / 3'-5' RNA helicase activity / generation of catalytic spliceosome for first transesterification step / cis assembly of pre-catalytic spliceosome / splicing factor binding / U4/U6 snRNP / spliceosome conformational change to release U4 (or U4atac) and U1 (or U11) / 7-methylguanosine cap hypermethylation / Prp19 complex / snRNP binding / ATP-dependent activity, acting on RNA / pICln-Sm protein complex / small nuclear ribonucleoprotein complex / pre-mRNA binding / U2-type catalytic step 1 spliceosome / SMN-Sm protein complex / spliceosomal tri-snRNP complex / mRNA cis splicing, via spliceosome / U2-type spliceosomal complex / poly(U) RNA binding / commitment complex / U2-type prespliceosome assembly / U2-type catalytic step 2 spliceosome / U4 snRNP / U2 snRNP / U1 snRNP / U2-type prespliceosome / precatalytic spliceosome / Formation of TC-NER Pre-Incision Complex / generation of catalytic spliceosome for second transesterification step / spliceosomal complex assembly / Gap-filling DNA repair synthesis and ligation in TC-NER / DNA replication origin binding / mRNA 5'-splice site recognition / protein K63-linked ubiquitination / mRNA 3'-splice site recognition / Dual incision in TC-NER / DNA replication initiation / spliceosomal tri-snRNP complex assembly / U5 snRNA binding / U5 snRNP / U2 snRNA binding / U6 snRNA binding / spliceosomal snRNP assembly / positive regulation of cell cycle / pre-mRNA intronic binding / U1 snRNA binding / U4/U6 x U5 tri-snRNP complex / catalytic step 2 spliceosome / RNA splicing / positive regulation of RNA splicing / spliceosomal complex / RING-type E3 ubiquitin transferase / mRNA splicing, via spliceosome / ubiquitin-protein transferase activity / metallopeptidase activity / ubiquitin protein ligase activity / nucleic acid binding / RNA helicase activity / RNA helicase / DNA repair / chromatin binding / GTPase activity / mRNA binding / chromatin / GTP binding / ATP hydrolysis activity / mitochondrion / DNA binding / RNA binding / ATP binding / identical protein binding / nucleus / metal ion binding / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  | |||||||||
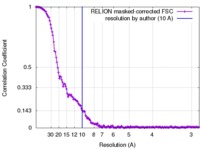
| Method | single particle reconstruction / cryo EM / Resolution: 10.0 Å | |||||||||
 Authors Authors | Galej WP / Wilkinson MF / Fica SM / Oubridge C / Newman AJ / Nagai K | |||||||||
 Citation Citation |  Journal: Nature / Year: 2016 Journal: Nature / Year: 2016Title: Cryo-EM structure of the spliceosome immediately after branching. Authors: Wojciech P Galej / Max E Wilkinson / Sebastian M Fica / Chris Oubridge / Andrew J Newman / Kiyoshi Nagai /  Abstract: Precursor mRNA (pre-mRNA) splicing proceeds by two consecutive transesterification reactions via a lariat-intron intermediate. Here we present the 3.8 Å cryo-electron microscopy structure of the ...Precursor mRNA (pre-mRNA) splicing proceeds by two consecutive transesterification reactions via a lariat-intron intermediate. Here we present the 3.8 Å cryo-electron microscopy structure of the spliceosome immediately after lariat formation. The 5'-splice site is cleaved but remains close to the catalytic Mg site in the U2/U6 small nuclear RNA (snRNA) triplex, and the 5'-phosphate of the intron nucleotide G(+1) is linked to the branch adenosine 2'OH. The 5'-exon is held between the Prp8 amino-terminal and linker domains, and base-pairs with U5 snRNA loop 1. Non-Watson-Crick interactions between the branch helix and 5'-splice site dock the branch adenosine into the active site, while intron nucleotides +3 to +6 base-pair with the U6 snRNA ACAGAGA sequence. Isy1 and the step-one factors Yju2 and Cwc25 stabilize docking of the branch helix. The intron downstream of the branch site emerges between the Prp8 reverse transcriptase and linker domains and extends towards the Prp16 helicase, suggesting a plausible mechanism of remodelling before exon ligation. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_4057.map.gz emd_4057.map.gz | 245.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-4057-v30.xml emd-4057-v30.xml emd-4057.xml emd-4057.xml | 57.6 KB 57.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_4057_fsc.xml emd_4057_fsc.xml | 14.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_4057.png emd_4057.png | 47 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-4057 http://ftp.pdbj.org/pub/emdb/structures/EMD-4057 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4057 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4057 | HTTPS FTP |
-Validation report
| Summary document |  emd_4057_validation.pdf.gz emd_4057_validation.pdf.gz | 279.5 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_4057_full_validation.pdf.gz emd_4057_full_validation.pdf.gz | 278.6 KB | Display | |
| Data in XML |  emd_4057_validation.xml.gz emd_4057_validation.xml.gz | 14.1 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-4057 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-4057 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-4057 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-4057 | HTTPS FTP |
-Related structure data
| Related structure data |  5lj5MC  4055C  4056C  4058C  4059C  5lj3C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | |
| EM raw data |  EMPIAR-10687 (Title: Yeast C, Ci, C*, and P complex spliceosomes / Data size: 8.9 TB EMPIAR-10687 (Title: Yeast C, Ci, C*, and P complex spliceosomes / Data size: 8.9 TBData #1: Unaligned movies of C-complex spliceosome with 3' splice site AG to AC mutation (Dataset 1) [micrographs - multiframe] Data #2: Unaligned movies of C and C*-complex spliceosomes with 3' splice site AG to AdG mutation (Dataset 2) [micrographs - multiframe] Data #3: Unaligned movies of C and C*-complex spliceosomes with 3' splice site AG to AdG mutation (Dataset 3) [micrographs - multiframe] Data #4: Aligned movies of C-complex spliceosomes with cold-sensitive prp16-302 mutation, purified with Cwc25 (Dataset 4) [micrographs - multiframe] Data #5: Unaligned movies of C-complex spliceosomes with cold-sensitive prp16-302 mutation, purified with Cwc25 and incubated with ATP and Mg (Dataset 5) [micrographs - multiframe] Data #6: Unaligned movies of C, C*, and P-complex spliceosomes with dominant-negative Prp22 mutation K512A, purified with Slu7 (Dataset 6) [micrographs - multiframe] Data #7: Unaligned movies of P-complex spliceosomes with dominant-negative Prp22 mutation K512A, treated with anti-3'exon RNaseH oligo, purified in presence of Mg (Dataset 9) [micrographs - single frame] Data #8: Selected C-complex particles after polishing [picked particles - single frame - processed] Data #9: Selected P-complex particles after polishing [picked particles - single frame - processed] Data #10: Various signal subtractions for C- and P-complex spliceosomes [picked particles - single frame - processed]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_4057.map.gz / Format: CCP4 / Size: 266.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_4057.map.gz / Format: CCP4 / Size: 266.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 1.43 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
+Entire : Spliceosome immediately after branching
+Supramolecule #1: Spliceosome immediately after branching
+Macromolecule #1: U5 snRNA (small nuclear RNA)
+Macromolecule #2: Exon 1 (5' exon) of UBC4 pre-mRNA
+Macromolecule #3: Intron of UBC4 pre-mRNA
+Macromolecule #4: U2 snRNA (small nuclear RNA)
+Macromolecule #5: U6 snRNA (small nuclear RNA)
+Macromolecule #6: Pre-mRNA-splicing factor 8
+Macromolecule #7: Pre-mRNA-splicing helicase BRR2
+Macromolecule #8: Protein CWC16
+Macromolecule #9: Pre-mRNA-splicing factor CWC25
+Macromolecule #10: Pre-mRNA-splicing factor SNU114
+Macromolecule #11: Pre-mRNA-splicing factor ISY1
+Macromolecule #12: CWC22
+Macromolecule #13: Pre-mRNA-splicing factor PRP46
+Macromolecule #14: Pre-mRNA-processing protein 45
+Macromolecule #15: Pre-mRNA-splicing factor BUD31
+Macromolecule #16: Pre-mRNA-splicing factor CWC2
+Macromolecule #17: Pre-mRNA-splicing factor SLT11
+Macromolecule #18: Pre-mRNA-splicing factor CEF1
+Macromolecule #19: CWC15
+Macromolecule #20: Pre-mRNA-splicing factor CWC21
+Macromolecule #21: Pre-mRNA-splicing factor CLF1
+Macromolecule #22: Pre-mRNA-splicing factor SYF1
+Macromolecule #23: Small nuclear ribonucleoprotein-associated protein B
+Macromolecule #24: Small nuclear ribonucleoprotein Sm D3
+Macromolecule #25: Small nuclear ribonucleoprotein E
+Macromolecule #26: Small nuclear ribonucleoprotein F
+Macromolecule #27: Small nuclear ribonucleoprotein G
+Macromolecule #28: Small nuclear ribonucleoprotein Sm D1
+Macromolecule #29: Small nuclear ribonucleoprotein Sm D2
+Macromolecule #30: U2 small nuclear ribonucleoprotein A'
+Macromolecule #31: U2 small nuclear ribonucleoprotein B''
+Macromolecule #32: Pre-mRNA-splicing factor ATP-dependent RNA helicase PRP16
+Macromolecule #33: Pre-mRNA-processing factor 19
+Macromolecule #34: Pre-mRNA-splicing factor SNT309
+Macromolecule #35: unknown
+Macromolecule #36: MAGNESIUM ION
+Macromolecule #37: ZINC ION
+Macromolecule #38: GUANOSINE-5'-TRIPHOSPHATE
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.3 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.8 Component:
| ||||||||||||
| Grid | Model: Quantifoil R2/2 / Material: COPPER / Mesh: 400 / Support film - Material: CARBON / Support film - topology: CONTINUOUS / Support film - Film thickness: 6.0 nm / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Atmosphere: AIR | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK III Details: 3 microlitres sample were applied to the grid, left for 30 seconds and then blotted for 2.5-3.0 seconds before plunging.. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Quantum |
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Detector mode: SUPER-RESOLUTION / Digitization - Frames/image: 1-20 / Number real images: 2213 / Average exposure time: 0.8 sec. / Average electron dose: 2.0 e/Å2 Details: Total dose: 40 electrons/Angstrom^2 over 16 seconds. 20 movie frames collected at 1.25 frames per second. |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 35714 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 4.0 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 81000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Details | Used secondary structure restraints generated in ProSMART and LibG. |
|---|---|
| Refinement | Space: RECIPROCAL / Protocol: OTHER |
| Output model |  PDB-5lj5: |
 Movie
Movie Controller
Controller