[English] 日本語
 Yorodumi
Yorodumi- EMDB-8160: Negative stain EM structure of Ebola GP in complex with mAb fab Q411 -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-8160 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
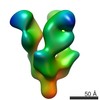
| Title | Negative stain EM structure of Ebola GP in complex with mAb fab Q411 | |||||||||
 Map data Map data | None | |||||||||
 Sample Sample |
| |||||||||
| Biological species |   | |||||||||
| Method | single particle reconstruction / negative staining / Resolution: 25.2 Å | |||||||||
 Authors Authors | Gui M / Xiang Y / Zhang Q / Zhang L | |||||||||
 Citation Citation |  Journal: Sci Rep / Year: 2016 Journal: Sci Rep / Year: 2016Title: Potent neutralizing monoclonal antibodies against Ebola virus infection. Authors: Qi Zhang / Miao Gui / Xuefeng Niu / Shihua He / Ruoke Wang / Yupeng Feng / Andrea Kroeker / Yanan Zuo / Hua Wang / Ying Wang / Jiade Li / Chufang Li / Yi Shi / Xuanling Shi / George F Gao / ...Authors: Qi Zhang / Miao Gui / Xuefeng Niu / Shihua He / Ruoke Wang / Yupeng Feng / Andrea Kroeker / Yanan Zuo / Hua Wang / Ying Wang / Jiade Li / Chufang Li / Yi Shi / Xuanling Shi / George F Gao / Ye Xiang / Xiangguo Qiu / Ling Chen / Linqi Zhang /   Abstract: Ebola virus infections cause a deadly hemorrhagic disease for which no vaccines or therapeutics has received regulatory approval. Here we show isolation of three (Q206, Q314 and Q411) neutralizing ...Ebola virus infections cause a deadly hemorrhagic disease for which no vaccines or therapeutics has received regulatory approval. Here we show isolation of three (Q206, Q314 and Q411) neutralizing monoclonal antibodies (mAbs) against the surface glycoprotein (GP) of Ebola virus identified in West Africa in 2014 through sequential immunization of Chinese rhesus macaques and antigen-specific single B cell sorting. These mAbs demonstrated potent neutralizing activities against both pseudo and live Ebola virus independent of complement. Biochemical, single particle EM, and mutagenesis analysis suggested Q206 and Q411 recognized novel epitopes in the head while Q314 targeted the glycan cap in the GP1 subunit. Q206 and Q411 appeared to influence GP binding to its receptor NPC1. Treatment with these mAbs provided partial but significant protection against disease in a mouse model of Ebola virus infection. These novel mAbs could serve as promising candidates for prophylactic and therapeutic interventions against Ebola virus infection. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_8160.map.gz emd_8160.map.gz | 20.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-8160-v30.xml emd-8160-v30.xml emd-8160.xml emd-8160.xml | 9 KB 9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_8160_fsc.xml emd_8160_fsc.xml | 6.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_8160.png emd_8160.png | 32.9 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-8160 http://ftp.pdbj.org/pub/emdb/structures/EMD-8160 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8160 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8160 | HTTPS FTP |
-Validation report
| Summary document |  emd_8160_validation.pdf.gz emd_8160_validation.pdf.gz | 363.9 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_8160_full_validation.pdf.gz emd_8160_full_validation.pdf.gz | 363.5 KB | Display | |
| Data in XML |  emd_8160_validation.xml.gz emd_8160_validation.xml.gz | 8.8 KB | Display | |
| Data in CIF |  emd_8160_validation.cif.gz emd_8160_validation.cif.gz | 11.6 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-8160 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-8160 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-8160 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-8160 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_8160.map.gz / Format: CCP4 / Size: 22.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_8160.map.gz / Format: CCP4 / Size: 22.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | None | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.68 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Ternary complex of Ebola virus glycoprotein and Fab Q411
| Entire | Name: Ternary complex of Ebola virus glycoprotein and Fab Q411 |
|---|---|
| Components |
|
-Supramolecule #1: Ternary complex of Ebola virus glycoprotein and Fab Q411
| Supramolecule | Name: Ternary complex of Ebola virus glycoprotein and Fab Q411 type: complex / ID: 1 / Parent: 0 |
|---|
-Supramolecule #2: Ebola virus glycoprotein
| Supramolecule | Name: Ebola virus glycoprotein / type: complex / ID: 2 / Parent: 1 |
|---|---|
| Source (natural) | Organism:  |
-Supramolecule #3: fab Q411
| Supramolecule | Name: fab Q411 / type: complex / ID: 3 / Parent: 1 |
|---|---|
| Source (natural) | Organism:  |
-Experimental details
-Structure determination
| Method | negative staining |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.02 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Staining | Type: NEGATIVE / Material: Uranyl Acetate |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F20 |
|---|---|
| Image recording | Film or detector model: GATAN ULTRASCAN 4000 (4k x 4k) / Average electron dose: 30.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Tecnai F20 / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller