[English] 日本語
 Yorodumi
Yorodumi- EMDB-6429: Architecture of the complex formed by large and small Terminase s... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-6429 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Architecture of the complex formed by large and small Terminase subunits from Bacteriophage P22 | |||||||||
 Map data Map data | Asymmetric reconstruction of terminase holoenzyme from Bacteriophage P22 | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | viral genome-packaging motor / Terminase complex / Salmonella virus / P22 Bacteriophage | |||||||||
| Biological species |  Enterobacteria phage P22 (virus) Enterobacteria phage P22 (virus) | |||||||||
| Method | single particle reconstruction / negative staining / Resolution: 30.0 Å | |||||||||
 Authors Authors | McNulty R / Johnson JE | |||||||||
 Citation Citation |  Journal: J Mol Biol / Year: 2015 Journal: J Mol Biol / Year: 2015Title: Architecture of the Complex Formed by Large and Small Terminase Subunits from Bacteriophage P22. Authors: Reginald McNulty / Ravi Kumar Lokareddy / Ankoor Roy / Yang Yang / Gabriel C Lander / Albert J R Heck / John E Johnson / Gino Cingolani /   Abstract: Packaging of viral genomes inside empty procapsids is driven by a powerful ATP-hydrolyzing motor, formed in many double-stranded DNA viruses by a complex of a small terminase (S-terminase) subunit ...Packaging of viral genomes inside empty procapsids is driven by a powerful ATP-hydrolyzing motor, formed in many double-stranded DNA viruses by a complex of a small terminase (S-terminase) subunit and a large terminase (L-terminase) subunit, transiently docked at the portal vertex during genome packaging. Despite recent progress in elucidating the structure of individual terminase subunits and their domains, little is known about the architecture of an assembled terminase complex. Here, we describe a bacterial co-expression system that yields milligram quantities of the S-terminase:L-terminase complex of the Salmonella phage P22. In vivo assembled terminase complex was affinity-purified and stabilized by addition of non-hydrolyzable ATP, which binds specifically to the ATPase domain of L-terminase. Mapping studies revealed that the N-terminus of L-terminase ATPase domain (residues 1-58) contains a minimal S-terminase binding domain sufficient for stoichiometric association with residues 140-162 of S-terminase, the L-terminase binding domain. Hydrodynamic analysis by analytical ultracentrifugation sedimentation velocity and native mass spectrometry revealed that the purified terminase complex consists predominantly of one copy of the nonameric S-terminase bound to two equivalents of L-terminase (1S-terminase:2L-terminase). Direct visualization of this molecular assembly in negative-stained micrographs yielded a three-dimensional asymmetric reconstruction that resembles a "nutcracker" with two L-terminase protomers projecting from the C-termini of an S-terminase ring. This is the first direct visualization of a purified viral terminase complex analyzed in the absence of DNA and procapsid. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_6429.map.gz emd_6429.map.gz | 1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-6429-v30.xml emd-6429-v30.xml emd-6429.xml emd-6429.xml | 12.6 KB 12.6 KB | Display Display |  EMDB header EMDB header |
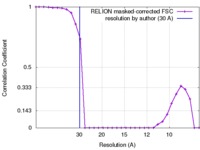
| FSC (resolution estimation) |  emd_6429_fsc.xml emd_6429_fsc.xml | 2.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_6429.png emd_6429.png | 38.9 KB | ||
| Masks |  emd_6429_msk_1.map emd_6429_msk_1.map | 1.4 MB |  Mask map Mask map | |
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-6429 http://ftp.pdbj.org/pub/emdb/structures/EMD-6429 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6429 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6429 | HTTPS FTP |
-Validation report
| Summary document |  emd_6429_validation.pdf.gz emd_6429_validation.pdf.gz | 77.5 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_6429_full_validation.pdf.gz emd_6429_full_validation.pdf.gz | 76.6 KB | Display | |
| Data in XML |  emd_6429_validation.xml.gz emd_6429_validation.xml.gz | 495 B | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6429 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6429 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6429 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6429 | HTTPS FTP |
-Related structure data
| Similar structure data |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_6429.map.gz / Format: CCP4 / Size: 1.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_6429.map.gz / Format: CCP4 / Size: 1.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Asymmetric reconstruction of terminase holoenzyme from Bacteriophage P22 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 4.1 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Segmentation: Mask of complex generated automatically with RELION
| Annotation | Mask of complex generated automatically with RELION | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| File |  emd_6429_msk_1.map emd_6429_msk_1.map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Bacteriophage P22 terminase holoenzyme
| Entire | Name: Bacteriophage P22 terminase holoenzyme |
|---|---|
| Components |
|
-Supramolecule #1000: Bacteriophage P22 terminase holoenzyme
| Supramolecule | Name: Bacteriophage P22 terminase holoenzyme / type: sample / ID: 1000 Oligomeric state: One nonamer of S-terminase binds to two L-terminase subunits Number unique components: 2 |
|---|---|
| Molecular weight | Experimental: 287 KDa / Theoretical: 287 KDa / Method: Native MS |
-Macromolecule #1: small terminase subunit
| Macromolecule | Name: small terminase subunit / type: protein_or_peptide / ID: 1 / Number of copies: 9 / Oligomeric state: 9 small subunits : 2 large subunits / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  Enterobacteria phage P22 (virus) / synonym: P22 Enterobacteria phage P22 (virus) / synonym: P22 |
| Recombinant expression | Organism:  |
-Macromolecule #2: large terminase subunit
| Macromolecule | Name: large terminase subunit / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Oligomeric state: 9 small subunits : 2 large subunits / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  Enterobacteria phage P22 (virus) / synonym: P22 Enterobacteria phage P22 (virus) / synonym: P22 |
| Recombinant expression | Organism:  |
-Experimental details
-Structure determination
| Method | negative staining |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.014 mg/mL |
|---|---|
| Buffer | pH: 8 Details: 20 mM Tris-HCl, 150 mM NaCl, 3 mm DTT, 5% glycerol, 1 mM MgCl2 |
| Staining | Type: NEGATIVE Details: Protein was adsorbed to the grid for 1 minute, blotted, and passed through four consecutive 40 microliter drops of 2% uranyl formate. |
| Grid | Details: 400 mesh copper grids charged with Gatan plasma cleaner |
| Vitrification | Cryogen name: NONE / Instrument: OTHER |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI 12 |
|---|---|
| Date | Mar 25, 2015 |
| Image recording | Category: CCD / Film or detector model: TVIPS TEMCAM-F416 (4k x 4k) / Number real images: 44 / Average electron dose: 20 e/Å2 / Details: Images were acquired using Leginon. |
| Electron beam | Acceleration voltage: 120 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 0.2 µm / Nominal defocus min: 0.1 µm / Nominal magnification: 52000 |
| Sample stage | Specimen holder model: SIDE ENTRY, EUCENTRIC |
 Movie
Movie Controller
Controller











 Z
Z Y
Y X
X