[English] 日本語
 Yorodumi
Yorodumi- EMDB-5983: Structure of 2G12 (Fab)2 in Complex with Soluble and Fully Glycos... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-5983 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of 2G12 (Fab)2 in Complex with Soluble and Fully Glycosylated HIV-1 Env by Negative-Stain Single Particle Electron Microscopy | |||||||||
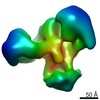
 Map data Map data | Reconstruction of HIV-1 Env SOSIP BG505.664 bound to soluble, two-domain CD4 (domains 1 and 2, sCD4) and 2G12 Fabs | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | HIV-1 / 2G12 / monoclonal antibodies / Envelope / CD4-bound Env | |||||||||
| Biological species |   Human immunodeficiency virus 1 / Human immunodeficiency virus 1 /  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / negative staining / Resolution: 26.0 Å | |||||||||
 Authors Authors | Murin CD / Julien JP / Sok D / Stanfield R / Khayat R / Cupo A / Moore JP / Burton DR / Wilson IA / Ward AB | |||||||||
 Citation Citation |  Journal: J Virol / Year: 2014 Journal: J Virol / Year: 2014Title: Structure of 2G12 Fab2 in complex with soluble and fully glycosylated HIV-1 Env by negative-stain single-particle electron microscopy. Authors: Charles D Murin / Jean-Philippe Julien / Devin Sok / Robyn L Stanfield / Reza Khayat / Albert Cupo / John P Moore / Dennis R Burton / Ian A Wilson / Andrew B Ward /  Abstract: The neutralizing anti-HIV-1 antibody 2G12 is of particular interest due to the sterilizing protection it provides from viral challenge in animal models. 2G12 is a unique, domain-exchanged antibody ...The neutralizing anti-HIV-1 antibody 2G12 is of particular interest due to the sterilizing protection it provides from viral challenge in animal models. 2G12 is a unique, domain-exchanged antibody that binds exclusively to conserved N-linked glycans that form the high-mannose patch on the gp120 outer domain centered on a glycan at position N332. Several glycans in and around the 2G12 epitope have been shown to interact with other potent, broadly neutralizing antibodies; therefore, this region constitutes a supersite of vulnerability on gp120. While crystal structures of 2G12 and 2G12 bound to high-mannose glycans have been solved, no structural information that describes the interaction of 2G12 with gp120 or the Env trimer is available. Here, we present a negative-stain single-particle electron microscopy reconstruction of 2G12 Fab2 in complex with a soluble, trimeric Env at ∼17-Å resolution that reveals the antibody's interaction with its native and fully glycosylated epitope. We also mapped relevant glycans in this epitope by fitting high-resolution crystal structures and by performing neutralization assays of glycan knockouts. In addition, a reconstruction at ∼26 Å of the ternary complex formed by 2G12 Fab2, soluble CD4, and Env indicates that 2G12 may block membrane fusion by induced steric hindrance upon primary receptor binding, thereby abrogating Env's interaction with coreceptor(s). These structures provide a basis for understanding 2G12 binding and neutralization, and our low-resolution model and glycan assignments provide a basis for higher-resolution studies to determine the molecular nature of the 2G12 epitope. IMPORTANCE: HIV-1 is a human virus that results in the deaths of millions of people around the world each year. While there are several effective therapeutics available to prolong life, a vaccine is ...IMPORTANCE: HIV-1 is a human virus that results in the deaths of millions of people around the world each year. While there are several effective therapeutics available to prolong life, a vaccine is the best long-term solution for curbing this global epidemic. Here, we present structural data that reveal the viral binding site of one of the first HIV-1-neutralizing antibodies isolated, 2G12, and provide a rationale for its effectiveness. These structures provide a basis for higher-resolution studies to determine the molecular nature of the 2G12 epitope, which will aid in vaccine design and antibody-based therapies. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_5983.map.gz emd_5983.map.gz | 2.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-5983-v30.xml emd-5983-v30.xml emd-5983.xml emd-5983.xml | 14 KB 14 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_5983.tif emd_5983.tif | 2.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-5983 http://ftp.pdbj.org/pub/emdb/structures/EMD-5983 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5983 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5983 | HTTPS FTP |
-Validation report
| Summary document |  emd_5983_validation.pdf.gz emd_5983_validation.pdf.gz | 78.4 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_5983_full_validation.pdf.gz emd_5983_full_validation.pdf.gz | 77.5 KB | Display | |
| Data in XML |  emd_5983_validation.xml.gz emd_5983_validation.xml.gz | 494 B | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5983 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5983 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5983 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5983 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_5983.map.gz / Format: CCP4 / Size: 3.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_5983.map.gz / Format: CCP4 / Size: 3.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Reconstruction of HIV-1 Env SOSIP BG505.664 bound to soluble, two-domain CD4 (domains 1 and 2, sCD4) and 2G12 Fabs | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 4.35 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Fab fragment of 2G12 monoclonal antibody and soluble, two-domain ...
| Entire | Name: Fab fragment of 2G12 monoclonal antibody and soluble, two-domain CD4 (domains 1 and 2, sCD4) bound to HIV-1 Env BG505.664 |
|---|---|
| Components |
|
-Supramolecule #1000: Fab fragment of 2G12 monoclonal antibody and soluble, two-domain ...
| Supramolecule | Name: Fab fragment of 2G12 monoclonal antibody and soluble, two-domain CD4 (domains 1 and 2, sCD4) bound to HIV-1 Env BG505.664 type: sample / ID: 1000 Oligomeric state: Env trimer bound to three 2G12 domain-swapped Fabs and three sCD4s Number unique components: 3 |
|---|---|
| Molecular weight | Experimental: 730 KDa / Theoretical: 729 KDa / Method: SEC-MALS |
-Macromolecule #1: HIV-1 Env
| Macromolecule | Name: HIV-1 Env / type: protein_or_peptide / ID: 1 / Name.synonym: SOSIP BG505.664 / Number of copies: 1 / Oligomeric state: Trimer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 / synonym: HIV-1 Human immunodeficiency virus 1 / synonym: HIV-1 |
| Molecular weight | Experimental: 360 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) / Recombinant cell: HEK 293F Homo sapiens (human) / Recombinant cell: HEK 293F |
-Macromolecule #2: soluble CD4
| Macromolecule | Name: soluble CD4 / type: protein_or_peptide / ID: 2 / Name.synonym: sCD4 / Number of copies: 1 / Oligomeric state: monomer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / synonym: Human Homo sapiens (human) / synonym: Human |
| Molecular weight | Theoretical: 23 KDa |
| Recombinant expression | Organism:  |
-Macromolecule #3: Human Monoclonal Antibody 2G12 IgG1 Fab Fragment
| Macromolecule | Name: Human Monoclonal Antibody 2G12 IgG1 Fab Fragment / type: protein_or_peptide / ID: 3 / Name.synonym: 2G12 Fab / Oligomeric state: Dimer of Fabs / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / synonym: Human Homo sapiens (human) / synonym: Human |
| Molecular weight | Experimental: 100 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) / Recombinant cell: HEK 293F / Recombinant plasmid: phCMV3 Homo sapiens (human) / Recombinant cell: HEK 293F / Recombinant plasmid: phCMV3 |
-Experimental details
-Structure determination
| Method | negative staining |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.03 mg/mL |
|---|---|
| Buffer | pH: 7.4 / Details: 150 mM NaCl, 10 mM Tris |
| Staining | Type: NEGATIVE Details: Grids with adsorbed protein were floated on 2% w/v uranyl formate for 30 seconds. |
| Grid | Details: 400 mesh copper grid with thin carbon support, Gatan plasma cleaned |
| Vitrification | Cryogen name: NONE / Instrument: OTHER |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI SPIRIT |
|---|---|
| Alignment procedure | Legacy - Astigmatism: Objective lens astigmatism was corrected at 100,000 times magnification. |
| Specialist optics | Energy filter - Name: FEI |
| Date | Jul 12, 2012 |
| Image recording | Category: CCD / Film or detector model: OTHER / Number real images: 149 / Average electron dose: 30 e/Å2 |
| Tilt angle min | 0 |
| Electron beam | Acceleration voltage: 120 kV / Electron source: TUNGSTEN HAIRPIN |
| Electron optics | Calibrated magnification: 52000 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 0.8 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 52000 |
| Sample stage | Specimen holder model: HOME BUILD |
| Experimental equipment |  Model: Tecnai Spirit / Image courtesy: FEI Company |
- Image processing
Image processing
| Details | The particles were selected using automatic selection program Leginon and processed using the Appion system. |
|---|---|
| Final reconstruction | Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 26.0 Å / Resolution method: OTHER / Software - Name: EMAN2 / Details: Final map was low pass filtered to 20 Angstrom. / Number images used: 5372 |
| Final two d classification | Number classes: 100 |
-Atomic model buiding 1
| Initial model | PDB ID: |
|---|---|
| Software | Name:  Chimera Chimera |
| Details | Three structures of HIV-1 gp120 bound to sCD4 were fit into the HIV-1 Env trimer portion of the map. |
| Refinement | Space: REAL / Protocol: RIGID BODY FIT |
-Atomic model buiding 2
| Initial model | PDB ID:  1op5 |
|---|---|
| Software | Name:  Chimera Chimera |
| Details | 2G12 Fab structures were fit by rigid body fitting using the UCSF Chimera volume fit option, simulating a map at an estimated resolution of 20 Angstrom. |
| Refinement | Space: REAL / Protocol: RIGID BODY FIT |
 Movie
Movie Controller
Controller